Abstract
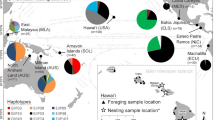
Although green turtles (Chelonia mydas Linnaeus) do not nest in Barbados, the easternmost island in the Caribbean archipelago, juveniles are regularly seen foraging in nearshore waters. To examine the stock composition of this foraging population, mitochondrial (mt) DNA control region sequences were analysed from 60 juvenile (31–70 cm curved carapace length) green turtles and compared with data published for key nesting populations in the Atlantic, as well as other feeding grounds (FGs) in the Caribbean. Eight distinct haplotypes were recognised among the 60 individual green turtles sampled around Barbados. Three of the haplotypes found have only previously been reported from western Caribbean nesting beaches, and two only from South Atlantic beaches. The nesting beach origin of one of the Barbados FG haplotypes is as yet unidentified. Stock mixture analysis based on Bayesian methods showed that the Barbados FG population is a genetically mixed stock consisting of approximately equal contributions from nesting beaches in Ascension Island (25.0%), Aves Island/Surinam (23.0%), Costa Rica (19.0%), and Florida (18.5%), with a lesser but significant contribution from Mexico (10.3%). Linear regression analysis indicated no significant effects of rookery population size or distance of the rookery from the FG on estimated contributions from the source rookeries to the Barbados FG. Our data suggest that the similar-sized green turtles sampled on the Barbados FG are a mixed stock of more diverse origins than any previously sampled feeding aggregations in the Caribbean region. The relatively large contribution from the Ascension Island rookery to the Barbados FG indicates that hatchlings from distant rookeries outside the Caribbean basin enter the North Atlantic gyre and become a significant part of the pool from which eastern Caribbean foraging populations are derived. These data support a life cycle model that incorporates a tendency of immatures to migrate from their initial foraging grounds at settlement towards suitable foraging grounds closer to their natal rookeries as they mature.


Similar content being viewed by others
References
Allard MW, Miyamoto MM, Bjorndal KA, Bolten AB, Bowen BW (1994) Support for natal homing in green turtles from mtDNA sequences. Copeia 1994:34–41
Bass AL, Witzell WN (2000) Demographic composition of immature green turtles (Chelonia mydas) from the east central Florida coast: evidence from mtDNA markers. Herpetologica 56:357–367
Bass AL, Lagueux CJ, Bowen BW (1998) Origin of green turtles, Chelonia mydas, at “sleeping rocks” off the northern coast of Nicaragua. Copeia 1998:1064–1069
Bjorndal KA, Bolten AB (1988) Growth rates of immature green turtles, Chelonia mydas, on feeding grounds in the southern Bahamas. Copeia 1988:555–564
Bolker B, Okuyama T, Bjorndal K, Bolten A (2003) Sea turtle stock estimation using genetic markers: accounting for sampling error of rare genotypes. Ecol Appl 13:763–775
Bowen BW, Karl SA (1997) Population genetics, phylogeography, and molecular evolution. In: Lutz P, Musick JA (eds) The biology of sea turtles. CRC Press, Boca Raton, Fla., pp 29–50
Bowen BW, Meylan AB, Ross JP, Limpus CJ, Balazs GH, Avise JC (1992) Global population structure and natural history of the green turtle (Chelonia mydas) in terms of matriarchal phylogeny. Evolution 46:865–881
Chapman RW (1996) A mixed stock analysis of the green turtle: the need for a null hypothesis. In: Bowen BW, Witzell WN (eds) Proceedings of the International Symposium on Sea Turtle Conservation Genetics. NOAA Tech Memo NMFS-SEFSC-396:137–146
Clewer AG, Scarisbrick DH (2001) Practical statistics and experimental design for plant and crop science. Wiley, Chichester
Dutton PH (1996) Methods for collection and preservation of samples for sea turtle genetic studies. In: Bowen BW, Witzell WN (eds) Proceedings of the International Symposium on Sea Turtle Conservation Genetics. NOAA Tech Memo NMFS-SEFSC-396:17–24
Encalada SE, Lahanas PN, Bjorndal KA, Bolten AB, Miyamoto MM, Bowen BW (1996) Phylogeography and population structure of the Atlantic and Mediterranean green turtle Chelonia mydas: a mitochondrial DNA control region sequence assessment. Mol Ecol 5:473–483
Lahanas PN, Miyamoto MM, Bjorndal KA, Bolten AB (1994) Molecular evolution and population genetics of greater Caribbean green turtles (Chelonia mydas) as inferred from mitochondrial DNA control region sequences. Genetica 94:57–67
Lahanas PN, Bjorndal KA, Bolten AB, Encalada SE, Miyamoto MM, Valverde RA, Bowen BW (1998) Genetic composition of a green turtle feeding ground population: evidence for multiple origins. Mar Biol 130:345–352
Norman JA, Moritz C, Limpus CJ (1994) Mitochondrial DNA control region polymorphisms: genetic markers for ecological studies of marine turtles. Mol Ecol 3:363–373
Pella M, Masuda J (2001) Bayesian methods for analysis of stock mixtures from genetic characters. NOAA Fish Bull 99:151–167
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, New York
Acknowledgements
We thank Stanton Thomas, Fred Watson, and Michael Armstrong for assistance in obtaining the tissue samples. Financial support from George Balazs, the National Marine Fisheries Service (NMFS), and the University of the West Indies to K. Luke are gratefully acknowledged. Thanks to A. Abreu-Grobois for stimulating interest in Atlantic current patterns. Genetic analysis was funded by the NMFS. Samples were obtained under CITES permit numbers 99US786600/9 and 00US844694/9. All work complies with current laws of Barbados and the United States.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P.W. Sammarco, Chauvin
Rights and permissions
About this article
Cite this article
Luke, K., Horrocks, J.A., LeRoux, R.A. et al. Origins of green turtle (Chelonia mydas) feeding aggregations around Barbados, West Indies. Marine Biology 144, 799–805 (2004). https://doi.org/10.1007/s00227-003-1241-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00227-003-1241-2