Abstract
The membrane is the major protective barrier separating the cell from the environment and is thus important for bacteria to survive environmental stress. This study investigates changes in membrane lipid compositions and membrane physiology of meat spoiling bacteria in response to high CO2 (30%) and O2 (70%) concentrations, as commonly used for modified atmosphere packaging of meat. Therefore, the fatty acid profile as well as membrane fluidity, permeability and cell surface were determined and correlated to the genomic settings of five meat spoiling bacteria Brochothrix (B.) thermosphacta, Carnobacterium (C.) divergens, C. maltaromaticum, Leuconostoc (L.) gelidum subsp. gelidum and L. gelidum subsp. gasicomitatum cultivated under different gas atmospheres. We identified different genomic potentials for fatty acid adaptations, which were in accordance with actual measured changes in the fatty acid composition for each species in response to CO2 and/or O2, e.g., an increase in saturated, iso and cyclopropane fatty acids. Even though fatty acid changes were species-specific, the general physiological responses were similar, comprising a decreased membrane permeability and fluidity. Thus, we concluded that meat spoiling bacteria facilitate a change in membrane fatty acids upon exposure to O2 and CO2, what leads to alteration of membrane fluidity and permeability. The observed adaptations might contribute to the resistance of meat spoilers against detrimental effects of the gases O2 and CO2 and thus help to explain their ability to grow under different modified atmospheres. Furthermore, this study provides fundamental knowledge regarding the impact of fatty acid changes on important membrane properties of bacteria.
Similar content being viewed by others
Introduction
Fresh meats are packaged under an artificial atmosphere to decelerate bacterially induced spoilage. This technique, called modified atmosphere packaging (MAP), is mainly based on the protective action of the gases oxygen and carbon dioxide, which generally suppress bacterial growth and thus prolong the shelf life of meat [1,2,3,4].
The harmful effect of oxygen and carbon dioxide on bacterial cells can be attributed to the fact that these molecules are rather non-polar and small, enabling them to easily penetrate the cell membrane by simple diffusion [5]. The mechanistic way of action of carbon dioxide within the cell has not been examined in detail yet, but it is supposed to decrease the intracellular pH and thus lead to essential loss of function of enzymes [6,7,8,9]. Furthermore, a mass action effect of carbon dioxide has been supposed, as specific enzymes such as the iso-citrate and malate dehydrogenase as well as decarboxylases, e.g. the phosphoenolpyruvate decarboxylase were inhibited due to high concentrations of carbon dioxide [6, 10, 11]. This can result in energetically decoupling, a phenomenon called futile cycle. Contrary, high oxygen concentrations are known to induce production of reactive oxygen species (ROS) within the cell, resulting in DNA, protein and lipid damage [12,13,14]. Thus, it can be hypothesized that survival of bacteria under these conditions is linked to their ability to adjust the biophysical properties of their membrane to enhance impermeability of these gases.
Membrane fluidity and permeability are two main properties of the membrane, which can be regulated by the cell. Both properties are mainly depending on the content of distinct fatty acids in the cell membrane [15]. Fatty acids differ according to the carbon chain length (long and short chain fatty acids), the interconnection of carbon atoms (saturated and unsaturated fatty acids (SFA and UFA)), the steric conformation of double bonds (cis and trans-UFAs) and the location of methyl groups (iso and anteiso-branched fatty acids) [15]. By modifying the structures of existing fatty acids as well as regulating biosynthesis of new fatty acids, bacteria are capable to alter physiological membrane properties resulting in regulation of transport processes, protein–protein interactions and passive permeability of hydrophobic molecules [15, 16]. Nevertheless, little is known regarding changes in the fatty acid profile providing lower permeability of the gases O2 and CO2 as used upon MAP.
In our previous full proteomic study targeting to investigate the effect of MAP on the metabolism of meat spoilage bacteria [9], we had also found certain proteins of e.g. biosynthesis of distinct fatty acids to be upregulated. Driven by this indication, the present study aimed to investigate if gases applied in modified atmosphere packaging influence bacterial fatty acid composition and to study the impact of these changes on physiological properties of the membrane.
Materials and methods
Bacterial strains and cultivation
Five major meat spoilage bacteria were selected in this study. One representative strain of each species was chosen from the TMW strain collection, according to its dominant abundance on high-oxygen packaged meat. L. gelidum subsp. gelidum TMW2.1618 and L. gelidum subsp. gasicomitatum TMW2.1619 where isolated from beef steaks by Hilgarth et al. [17], C. divergens TMW2.1577 and C. maltaromaticum TMW2.1581 from skinless chicken breast by Höll et al. [18] and B. thermosphacta TMW2.2101 from minced beef by Hilgarth et al. [19].
Comparative genome analysis
To evaluate bacteria’s potential to change their fatty acid profile in response to carbon dioxide and high oxygen exposure, a genomic prediction of all genes needed for biosynthesis, conversion and degradation of fatty acids was performed. Genomes of the five species haven been published in a previous study [20] and are available on NCBI with the accession numbers: CP017196 (L. gelidum subsp. gelidum TMW2.1618), CP017197 (L. gelidum subsp. gasicomitatum TMW2.1619), RSDV000000001 (C. divergens TMW2.1577), CP016844 (C. maltaromaticum TMW2.1581) and RSDU000000001 (B. thermosphacta TMW2.2101). All genes listed were additionally blasted manually to ensure correct annotation.
Experimental setup
Bacterial cultivation was performed in gas tight glass bottles filled with 0.4 L of a specifically developed meat simulation medium (MSM). The composition and species specific adaptation of the media is described in detail in a previous study by Kolbeck et al. [21]. Briefly, main components of MSM comprise meat extract, glycerol, tween80 and heminchlorid. Inoculation of bacteria in MSM was performed with a defined optical density (OD600 nm = 0.01). During cultivation time (48 h), MSM was aerated constantly with one of the following atmospheres, atmosphere1: 21%_O2/0%_CO2/79%_N2 (air), atmosphere2: 21%_O2/30%_CO2/49%_N2, atmosphere3: 70%_O2/30%_CO2/0%_N2. Atmosphere 3 represents the composition of commonly used high-oxygen MAP, while atmosphere 1 (air) acted as a control and atmosphere 2 was selected to study the effect of 30% CO2 as employed in MAP while retaining the oxygen concentration of air. By performing a pairwise comparison of the amount of each fatty acid identified in atmosphere 1 vs. atmosphere 2, we were able to uncover the effect of 30% CO2 on the fatty acid profile. The effect of 70% O2 was uncovered by comparing atmosphere 2 and atmosphere 3. Biological replicates were performed for each strain and gas mixture in three independent culture bottles. MSM was stirred constantly and temperature was set to 25 °C ± 2 °C. Bacteria were harvested at exponential growth phase for fatty acid analysis and physiological cell membrane assays.
Measurement of fatty acids
Harvested cells were frozen at -80 °C and freeze dried. The dry weight of each sample was determined afterwards. To detect losses during sample preparation, glyceryl trinonanoate was added as an internal standard. Lipid extraction was performed as described by Santivarangkna et al. [22]. Methylation of fatty acids was done by dissolving the extracted fat in 200 µl tetrahydrofuran (THF) and 400 µl of 0.5 mol l−1 sodium methylate (in Methanol). Samples were incubated for 15 min at 80 °C. Afterwards, 200 µl water and 200 µl hexane was added. Finally, the hexane layer was transferred into a vial for analysis.
Samples were determined using a gas chromatograph coupled to a mass spectrometer (GCMS-TQ8040, Shimadzu) with a split of 10 and injection volume of 1 µl. Sample separation was performed on a Zebron ZB-5MSplus column (30 m × 0.25 mm × 0.25 µm) from Phenomenex. Helium was chosen as a carrier gas. The following temperature gradient was run: 35 °C for 2 min, a rise of 10 °C min−1 until 140 °C, 140 °C for 10 min, a rise of 2 °C min−1 until 240 °C and finally 240 °C for 10.5 min. Data analyzation was performed with the software LabSolution—GCMSsolution (Version 4.30, Shimadzu). For fatty acid identification, the bacterial acid methyl ester standard (BAME Mix, Sigma-Aldrich, St. Louis, Missouri) as well as a mixture of fatty acid methyl esters (Supelco 37 FAME mix, Supelco, Bellefonte, PA, USA) was used. Furthermore, peak identification was confirmed by running a mass spectra search against the Nist11 database. Quantification was done by correlating the peak area of each fatty acid to the peak area of the used internal standard (C9:0) and its known concentration. Afterwards, total amounts of fatty acids were calculated per gram of cell mass and finally expressed in percentage of total fatty acids per single cell. All further calculations are based on this values which were calculated for all species in replicates under the three gas atmospheres.
Measurement of membrane fluidity
Membrane fluidity of cells was determined using the fluorescence dye Laurdan (6-Dodecanoyl-2-Dimethylaminonaphthalene). Sample were prepared as described by Behr et al. [23]. A LS 50B luminescence spectrometer (Perkin-Elmer, Rodgau-Jügesheim, Germany) was used for measuring of dyed samples (excitation wavelength 360 nm, emission wavelengths 380 to 550 nm, 5 nm steps). For comparing the membrane fluidity of a strain grown at different gas atmospheres, the general polarization (GP), defined as
was calculated for each replicate, where I440 is the relative fluorescence at 440 nm and I490 is the relative fluorescence at 490 nm [24]. To compare calculated GP values between different samples, the general polarization was calculated per cell. Therefore, GP values where divided by the viable cell count (CFU) of each sample, which was determined after fluorescence measurement.
Measurement of membrane permeability
A fluorescence assay using the dye propidium iodide (PI; Sigma-Aldrich) was performed to detect changes in membrane permeability [25]. Samples were prepared as described by Klotz et al. [26]. In detail, cells suspensions were adjusted to an OD600 = 1.0, washed with PBS, mixed with PI to a final concentration of 2.9 µM, incubated for 10 min at room temperature and washed in PBS. Fluorescence was determined at an excitation wavelength of 544 nm and an emission wavelength of 620 nm with a FLUOstar Omega microplate reader (BMG Labtech, Ortenberg, Germany). Afterwards, CFU was determined for each sample, and determined fluorescence intensity values where calculated per cell to ensure proper comparisons between different samples.
Measurement of cell surface
Cell sizes were determined of 50 cells from exponential growth phase per replicate by an Axiostar plus microscope (Zeiss, Jena, Germany) according to Sun, Liu [27]. The surface of the spheric-shaped Leuconostoc gelidum bacteria were calculated based on the following mode (shape code 1-H):
with a representing the diameter of the sphere. The surface of the rod-shaped bacteria B. thermosphacta, C. divergens and C. maltaromaticum were calculated based on the models (shape code 5-H):
with a representing the length and b the wide of the rod-shaped bacteria.
Statistical data and growth curve analysis
Data of fatty acid analysis and physiological assays (fluidity, permeability) were performed in three independent replicates and checked for statistical significance by conducting a variance and T-test (double side confidence interval 95%). Significant differences between variances or means where defined as p < 0.05. Microscopy analyses were performed in 50 replicates per strain and gas. Statistically significant differences of microscopy data where analyzed like described above. Growth curves were analyzed using the open source software RStudio ver. 3.3.0 (RStudio, Inc., Boston, MA, United States) running the CRAN package grofit ver. 1.1.1–1 with default settings. This package calculates the lag-Phase, maximal optical density (ODmax) and maximal growth rate (µmax) for each replicate by applying a fitting model to the data. Significant differences of these parameters between the three replicates and gas mixtures were calculated for each strain by performing an one-way analysis of variances (ANOVA). A post hoc Tukey test was connected to the ANOVA, assigning significant differences between means by applying a confidence interval of 95% (p = 0.05).
Results
Comparative genomic analysis for prediction of fatty acid metabolism
Bacterial genomes were analyzed focusing on the presence and absence of all genes needed to establish a membrane associated pathway using the database KEGG and own literature research. All bacteria in this study were able to predictively perform fatty acid de-novo biosynthesis and fatty acid membrane incorporation (Table 1). Regarding conversion of existing fatty acids, both L. gelidum subspecies and C. divergens TMW2.1577 are predictively able to build cyclopropane fatty acids as they exhibit the cyclopropane fatty acid synthase gene (Cfas). Direct fatty acid analysis also revealed the production of the corresponding fatty acid (C19:0 delta) for this species. Although, conjugated and unsaturated fatty acids could also be directly determined for all species (e.g., 18:2 conj.), no genes for unsaturation or conjugation of single steps leading to conversion of those fatty acids could be found in the five genomes. Genes facilitating degradation of short chain fatty acids were present in the genomes of all bacteria, whereas genes facilitating degradation of saturated medium and long chain fatty acid were only found in the genomes of both Carnobacterium species and B. thermosphacta TMW2.2101. Furthermore, both Carnobacterium spp. can predictively degrade polyunsaturated fatty acids. A detailed list of enzymes needed for membrane lipid metabolism is given in Table S1.
Revealing the effects of carbon dioxide and high oxygen by comparison of different gas compositions
The aim of this study was to uncover the effects of carbon dioxide and high oxygen on bacterial cell membrane properties. Therefore, bacteria were grown under atmosphere1: 21%_O2/0%_CO2/79%_N2 (air), atmosphere2: 21%_O2/30%_CO2/49%_N2 and atmosphere3: 70%_O2/30%_CO2/0%_N2. This experimental approach allowed uncovering the effects of carbon dioxide by comparing results obtained from atmosphere1 and atmosphere2 and the effects of high oxygen comparing results obtained from atmosphere2 and atmosphere3 (Table 2). Subsequently, all results obtained from different experiments where analyzed in this comparative manner and all differences stated are statistically significant.
Cell growth
Growth of bacteria in MSM under the different atmospheres was monitored over 48 h by measuring their optical density (Figure S1). Furthermore, growth analyses were performed, calculating the lag-Phase, division rate (µmax) and maximal OD for each replicate and atmosphere (Table S2).
Effect of CO2 (0% compared to 30%): A significant prolongation of the lag-phase could be determined for L. gelidum subsp. gelidum TMW2.1618, C. maltaromaticum TMW2.1581, C. divergens TMW2.1577 and B. thermosphacta TMW2.2101 but not for L. gelidum subsp. gasicomitatum TMW2.1619. The maximal OD was not affected by carbon dioxide for any species. We further detected an increase in the growth rate of C. maltaromaticum TMW2.1581 and L. gelidum subsp. gelidum TMW2.1618. The other species did not exhibit changes in their growth rate.
Effect of high O2 (21% compared to 70%): B. thermosphacta TMW2.2101 was the only species exhibiting a shortened lag phase due to high oxygen exposure. We further detected a decrease in maximal OD for L. gelidum subsp. gasicomitatum TMW2.1619 and an increase for C. maltaromaticum TMW2.1581. The other species were not affected in their maximal OD. Both L. gelidum subspecies exhibit a significant lower growth rate due to high oxygen exposure, whereas the growth rate of C. maltaromaticum TMW2.1581 was increased.
Changes of the fatty acid composition and predicted effect on membrane permeability and fluidity
The fatty acid composition of each species grown under the three atmospheres was analyzed and quantified in triplicates. The effects of carbon dioxide and high oxygen on single fatty acid regulation are listed in Table S3. An overview of the effects of carbon dioxide and high oxygen on fatty acids with similar properties e.g. chain length, branched methyl groups or double bounds is to be seen in Table 3. The predicted effect of those fatty acid changes on the membrane properties fluidity and permeability are summarized in Table 4 and marked by “FA”.
Effect of CO2 (0% compared to 30%): We determined an increase in iso- and short chain fatty acids and a decrease in long chain fatty acids for B. thermosphacta TMW2.2101, which is predicted to result in a decrease of the membrane fluidity. C. divergens TMW2.1577 and L. gelidum subsp. gelidum TMW2.1618 exhibit an increase in saturated and a decrease in unsaturated fatty acids, predictively leading to a less fluid and permeable cell membrane. Similar was also detected for L. gelidum subsp. gasicomitatum TMW2.1619, who exhibit a decrease in iso-, long chain and unsaturated fatty acids as well as an increase in short chain, saturated and total fatty acids, predictively resulting in a decreased membrane fluidity and permeability.
Effect of high O2 (21% compared to 70%): B. thermosphacta TMW2.2101 exhibits an increase in iso- and unbranched fatty acids and a decrease in anteiso- and branched fatty acids, predictively resulting in a reduced membrane fluidity. Contrary, C. divergens TMW2.1577 exhibits an increase in unsaturated, branched and total fatty acids and a decrease in saturated and unbranched fatty acids, leading to an increase in membrane fluidity and permeablity. We further determined an increase in total fatty acids for C. maltaromaticum TMW2.1581, which is predictively decreasing the membrane fluidity. L. gelidum subsp. gelidum TMW2.1618 increases its long chain and concomitantly decrease its short chain fatty acids under oxygen exposure, having no predicted effect on membrane fluidity or permeability. At least, we determined a slight decrease in iso-fatty acids for L. gelidum subsp. gasicomitatum TMW2.1619, presumably leading to an increase in membrane fluidity.
Measured membrane fluidity, permeability and cell surface
Physiological assays were performed for each species by measuring the effect of carbon dioxide and high oxygen on membrane fluidity (Fig. 1) and membrane permeability (Fig. 2). Furthermore, the cell surface was determined for all species cultivated under carbon dioxide and oxygen conditions (Fig. 3). Cell surfaces sizes where significantly different for most species and gases but absolute values do not exhibit strong differences (less than twofold regulation). Thus, determined surface values were only taken into account for interpretation of species significantly changing their total fatty acid content, e.g. an increase in total fatty acid content concomitant with an increase in cell surface does not affect cell membrane properties, whereas an increase in total fatty acids concomitant with an unchanged or even reduced cell surface implies a tighter and denser packaged cell membrane with reduced fluidity and permeability. Results from the physiological assays are also listed in Table 4 and marked by an “M”.
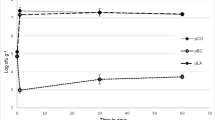
Measurement of the cell membrane fluidity of the five meat bacteria B. thermosphacta TMW2.2101, C. divergens TMW2.1577, C. maltaromaticum TMW2.1581, L. gelidum subsp. gelidum TMW2.1618 and L. gelidum subsp. gasicomitatum TMW2.1619 under the following atmospheres: (white square) atmosphere1: 21%_O2/0%_CO2/79%_N2 (air); (light grey square) atmosphere2: 21%_O2/30%_CO2/49%_N2; (dark grey square) atmosphere3: 70%_O2/30%_CO2/0%_N2. Bars represent mean values of three independent experiments, error bars represent standard deviations. * indicate significant differences between gas atmospheres identified by a two-sided T-test with p < 0.05
Measurement of the cell membrane permeability of the five meat bacteria B. thermosphacta TMW2.2101, C. divergens TMW2.1577, C. maltaromaticum TMW2.1581, L. gelidum subsp. gelidum TMW2.1618 and L. gelidum subsp. gasicomitatum TMW2.1619 under the following atmospheres: (white square) atmosphere1: 21%_O2/0%_CO2/79%_N2 (air); (light grey square) atmosphere2: 21%_O2/30%_CO2/49%_N2; (dark grey square) atmosphere3: 70%_O2/30%_CO2/0%_N2. Bars represent mean values of three independent experiments, error bars represent standard deviations. * indicate significant differences between gas atmospheres identified by a two-sided T-test with p < 0.05
Determination of cell surface of B. thermosphacta TMW2.2101, C. divergens TMW2.1577, C. maltaromaticum TMW2.1581, L. gelidum subsp. gelidum TMW2.1618 and L. gelidum subsp. gasicomitatum TMW2.1619 grown under the different atmospheres: (white square) atmosphere1: 21%_O2/0%_CO2/79%_N2 (air); (light grey square) atmosphere2: 21%_O2/30%_CO2/49%_N2; (dark grey square) atmosphere3: 70%_O2/30%_CO2/0%_N2. Boxes are based on 50 determined values. * indicate significant differences between gas atmospheres identified by a two-sided T-test with p < 0.05. Black diamonds represent outlier of the analysis
Effect of CO2 (0% compared to 30%): B. thermosphacta TMW2.2101 exhibits an increase in cell surface, but was not affected in its total fatty acid amount, thus having no effect on membrane fluidity or permeability. Similarly, C. divergens TMW2.1577 exhibited a decrease in cell surface without any regulation in total fatty acids. We measured a significant decrease in membrane fluidity for C. maltaromaticum TMW2.1581 due to CO2 exposition as well as an increase in cell surface which does not affect membrane properties, as no regulation of total fatty acids could be determined. Furthermore, we measured an increase in the cell surface as well as total fatty acid for L. gelidum subsp. gasicomitatum TMW2.1619, what in combination is predictively not affecting the membrane fluidity.
Effect of high O2 (21% compared to 70%): We determined a decrease in cell surface for B. thermosphacta TMW2.2101, which does not affect membrane properties, as no changes in total fatty acids could be detected. C. divergens TMW2.1577 exhibits an increase in cell surface and total fatty acids, both in combination also not affecting membrane properties. Contrary, we detected a significant decrease in cell surface and increase in total fatty acids for C. maltaromaticum TMW2.1581, predictively resulting in a decrease of membrane fluidity, which was further confirmed by a measured decrease in membrane fluidity due to high oxygen exposure for this species. Furthermore, C. maltaromaticum TMW2.1581 exhibits a measurable increase in membrane permeability under high oxygen exposure. L. gelidum subsp. gelidum TMW2.1618 exhibits an increase in cell surface with no change in total fatty acids, thus not affecting membrane properties, while L. gelidum subsp. gasicomitatum TMW2.1619 exhibits a measured increase in permeability under high oxygen conditions.
Figure 4 summarizes the major findings of this paper comprising the gas-depending shifts in the fatty acid profile and their effect on physiological membrane properties.
Graphical summary of the cell response to high oxygen (70%) or carbon dioxide (30%) exposure regarding fatty acid and physiological membrane changes. A shift in the fatty acid composition does influence the physiological membrane behaviour, e.g., fluidity and permeability. Arrows indicate if those membrane properties are increased or decreased in response to oxygen or carbon dioxide. Data shown are based on the overall reaction of all tested meat spoiling bacteria, which behave quite similar
Discussion
Protective MAP gases, i.e., oxygen and carbon dioxide, widely employed in the food industry, penetrate the cell membrane of the initial spoilage bacteria, induce oxidation of fatty acids and reduce protein functions, resulting in irremediable cell damage [12, 28]. In our previous full proteomic study, we have investigated the effect of different (modified) atmospheres on the metabolism of different meat spoilage bacteria. We had found a proteomic upregulation of several enzymes of the fatty acid synthesis II complex in response to carbon dioxide exposure for the species B. thermosphacta TMW2.2101 and C. maltaromaticum TMW2.1581, whereas latter also upregulated those genes under oxic conditions. Furthermore, all three subunits of the branched-chain alpha-keto acid dehydrogenase complex were upregulated under high oxic conditions for both Carnobacteria species. No metabolic regulations of fatty acid genes could be observed for both Leuconostoc species, which could be explained by posttranslational modifications of the corresponding proteins [9]. Thus, our previous results hinted that adaptation of the cell membrane fatty acid composition might contribute to enhance survival of spoilage bacteria on MAP meat by e.g. preventing penetration of detrimental gases into the cell. Consequently, this study investigates the potential of different meat spoiling bacteria to adapt their fatty acid profile and thus their membrane behavior to exposure to different MAP gases. Therefore, we analyzed the genomic potential to build, degrade and convert existing fatty acids and determined their actual fatty acid profile and their direct physiological changes in membrane properties such as fluidity and permeability under exposure to 30% CO2 or 70% O2.
Genomic potential for membrane fatty acid adaptation
Fatty acid degradation and interconversion are two important factors for rapidly changing of the existing fatty acid profile of bacteria. Indeed, we detected differences between those factors for the analyzed species. We predict the highest genomic potential for rapid fatty acid adaptation for the species C. divergens TMW2.1577 and C. maltaromaticum TMW2.1581 based on their extended ability to degrade different fatty acids and further produce cyclopropane fatty acids. Both L. gelidum subspecies are predictively also able to produce cyclopropane fatty acids but are disabled in saturated long and medium chain as well as polyunsaturated fatty acid degradation. B. thermosphacta TMW2.2101 is not able to produce cyclopropane fatty acids and degrade polyunsaturated fatty acids and thus exhibits the lowest potential to adapt its membrane to environmental stress. However, genomic analysis perse are not sufficient to predict membrane adaptation potentials of bacteria. Thus, further analysis for actual determination of fatty acid composition, membrane fluidity and permeability measurements were conducted.
Carbon dioxide (30%) induced membrane changes in meat spoilage bacteria
The restricted growth recorded for almost all species in this study in response to carbon dioxide exposure proves the high sensitivity of those bacteria to this gas. Even though all bacteria reached similar ODmax after 48 h, lag phases were significantly prolonged due to carbon dioxide, assuming a long metabolic adaptation phase comprising alteration of membrane fatty acid metabolism. Up to now, there are only few reports in the literature analyzing changes in membrane properties of bacteria due to the gas carbon dioxide, whereas there are no reports on the changes in fatty acid profiles, yet. Furthermore, there is only a single study investigating the effect of carbon dioxide on the interfacial tensions between water and oil phases (benzene) by Sears, Eisenberg [29]. Affiliated from their results on model systems, a decrease of ion permeability by CO2 for biological membranes was supposed [29]. Our study revealed an increase in SFAs and decrease in UFAs for C. divergens TMW2.1577 and both L. gelidum subspecies due to carbon dioxide. According to Zhang, Rock [15], SFAs lower, whereas cis-UFAs increase ion permeability of membranes. Thus, our results can confirm the assumption of Sears and Eisenberg, revealing a decrease of ion permeability of the cell membranes as a response to carbon dioxide exposure for these species.
In contrast to the few previous studies analyzing the direct effect of carbon dioxide on the cell membrane of bacteria, there are plenty of studies on membrane adaptations to acid stress in bacteria [21]. As one supposed way of action of carbon dioxide is the reduction of intracellular pH [6, 30, 31], membrane adaptation mechanisms of carbon dioxide and acid stress can be considered similar. In this way, an acid stress induced decrease of membrane permeability was also demonstrated for many Gram-positive bacteria, e.g. Listeria (Li.) monocytogenes [32] or Streptococcus mutans [33]. An association of acid stress and carbon dioxide is also demonstrated by our data, as the amount of lactobacillic acid (C19:0 delta), a cyclopropane fatty acid commonly found in lactic acid bacteria, was strongly upregulated due to carbon dioxide exposure for both L. gelidum subspecies and C. divergens TMW2.1577. Indeed, our genomic analysis revealed those species to be the only one encoding for the corresponding cyclopropane fatty acid synthase. According to the literature, an increase of cyclopropane fatty acids has been described to enhance acid resistance and further reduces cell membrane permeability [15, 34, 35]. Thus, these results further confirm the assumption of reduced membrane permeability to be one major adaptation mechanism of C. divergens TMW2.1577, L. gelidum subsp. gelidum TMW2.1618 and L. gelidum subsp. gasicomitatum TMW2.1619. Furthermore, as the corresponding enzyme for cyclization is missing in B. thermosphacta TMW2.2101 and C. maltaromaticum TMW2.1581, we assume these two species to lack an important regulatory mechanism, which presence would enable to adapt to acid as well as carbon dioxide induced stress.
Zhang and Rock have also demonstrated that a shift from anteiso to iso-fatty acids results in a reduction of membrane fluidity, as the methyl groups of anteiso-fatty acids are more distanced from the chain end and thus have a higher disruptive effect compared to iso-fatty acids. In our study, the named shift could be observed B. thermosphacta TMW2.2101 concluding a decrease in membrane fluidity for this species. Furthermore, according to Zhang, Rock [15], a shift from UFAs to SFAs also contributes to decreasing membrane fluidity [15] as demonstrated for C. divergens TMW2.1577 and both L. gelidum subspecies. We further directly measured a decreasing membrane fluidity for the species C. maltaromaticum TMW2.1581. Thus, all analyzed species seem to reduce their membrane fluidity in response to carbon dioxide by different changes within the membrane fatty acid profiles, concluding this mechanism to be a common bacterial response to carbon dioxide exposure.
Regarding cell surface sizes, we detected significant changes; however, the effect of carbon dioxide on cell size changes was not consistent for all species and absolute cell surface values were of comparable magnitude despite statistical differences. The elevated fatty acid content per cell determined for L. gelidum subsp. gasicomitatum TMW2.1619 could be, therefore, correlated to a bigger cell size and not to a tighter packaging of the fatty acids within the membrane. From this, we concluded fatty acid composition changes to be a more powerful tool of bacteria to respond to environmental stress than denser packaging of the fatty acids per se.
High oxygen (70%) induced membrane changes on meat bacteria
Molecular oxygen is used by aerobic microorganisms for respiratory growth. Nevertheless, high oxygen levels also result in RNA, DNA, protein and lipid damage [12] due to generation of reactive oxygen species (ROS). Depending on the genomic equipment, bacteria can greatly vary in their oxidative stress tolerance, as can be seen for the set of our species, comprising those who exhibit higher tolerance to high oxygen exposure (Carnobacteria species) and those with lower tolerance (Leuconostoc species). Our data further confirmed that oxygen also influences the fatty acid profile of the bacteria. We measured increased amounts of unsatured fatty acid for C. divergens TMW2.1577 in response to high oxygen exposure. Furthermore, the species C. maltaromaticum TMW2.1581 also exhibits an enhanced production of long-chain UFAs (conjugated linoleic acid (18:2 conj.), arachidonic (C20:4 ∆5,8,11,14) and eicosapentaenoic acid (C20:5 ∆5,8,11,14,17). Assuming that theses long-chain UFAs are generated by the activity of an oxygen-depending desaturation system [36,37,38], catabolic oxygen consumption can be seen as one adaptation mechanism of this species to high oxygen gas atmospheres [35, 39]. The increase in UFAs determined for C. maltaromaticum TMW2.1581 and C. divergens TMW2.1577, also assumed to increase membrane permeability of this species, which was confirmed by physiological assays. We further directly measured increased membrane permeability for the species L. gelidum subsp. gasicomitatum TMW2.1619. Thus, we concluded that high oxygen levels facilitate increased membrane permeability of meat spoiling bacteria, which could result in a reduction of the membrane barrier function.
We further determined a shift from anteiso- to iso-fatty acids for B. thermosphacta TMW2.2101 and a denser packaged cell membrane for C. maltaromaticum TMW2.1581. Latter was determined by an increase in total amounts of fatty acids concomitant with a decrease in cell volume. This is in accordance to our previously findings on an upregulation of several enzymes of the fatty acid synthesis II complex for C. maltaromaticum TMW2.1581 under high oxygen conditions [9]. Thus, B. thermosphacta TMW2.2101 and C. maltaromaticum TMW2.1581 seem to decrease their membrane fluidity in response to high oxygen stress. Contrary, C. divergens TMW2.1577 and L. gelidum subsp. gasicomitatum TMW2.1619 both increased their membrane fluidity by either increasing their amount of branched and unsaturated fatty acids or by reducing their amount of iso fatty acids. This is also in accordance to our previous proteomic study, where we observed a strong upregulation of the branched-chain alpha-keto acid dehydrogenase complex under oxic conditions for C. divergens TMW2.1577. Thus, adaptation of the membrane fluidity in response to exposure to high oxygen concentrations appears to be species-specific instead of a general mechanism.
This is the first study investigating potential bacterial adaptation of their cell membrane composition in response to exposure to gases applied in modified atmosphere packaging. Furthermore, this study gives insights into basic fatty acid regulatory mechanisms and their effect on changing important membrane properties of bacteria. In this manner, we first analyzed the genomic settings of our species to predict and validate their potential to adapt their fatty acid profile to environmental stress. We predicted different genomic potentials for each species based on presence or absence of genes involved in fatty acid degradation and cyclisation pathways. In accordance to this, we identified several species-specific modifications of fatty acid profiles due to high O2 and CO2 concentrations by actual physiological measurements. Interestingly, those fatty acid modification result in similar physiological membrane changes comprising reduced fluidity and permeability. It has been demonstrated that decreased membrane fluidity or permeability increases bacterial resistance to organic solvent [40] as well as antibiotics [41]. Thus, it appears that those changes facilitate also an enhanced resistance to the detrimental effects of gases such as CO2 or O2, by e.g. decreasing the diffusion into the cell or counteracting oxidative and acid stress as well as futile cycles. Supporting this hypothesis, we further identified an increased amount of lactobacillic acid in response to carbon dioxide exposure, predictively facilitating acid resistance. However, further studies should be conducted to confirm increased gas tolerance due to reduction of membrane fluidity and permeability of Gram-positive bacteria. Nevertheless, the present findings complement the previous demonstrated adaptive metabolic responses of meat spoiling bacteria to CO2 (30%) and high O2 (70%) within the full proteomic expression study.
References
Gill CO, Tan KH (1979) Effect of carbon dioxide on growth of Pseudomonas fluorescens. Appl Environ Microbiol 38(2):237–240
Ercolini D, Russo F, Torrieri E, Masi P, Villani F (2006) Changes in the spoilage-related microbiota of beef during refrigerated storage under different packaging conditions. Appl Environ Microbiol 72(7):4663–4671. https://doi.org/10.1128/aem.00468-06
Soldatou N, Nerantzaki A, Kontominas MG, Savvaidis IN (2009) Physicochemical and microbiological changes of “Souvlaki”: a Greek delicacy lamb meat product: evaluation of shelf-life using microbial, colour and lipid oxidation parameters. Food Chem 113(1):36–42. https://doi.org/10.1016/j.foodchem.2008.07.006
Esmer OK, Irkin R, Degirmencioglu N, Degirmencioglu A (2011) The effects of modified atmosphere gas composition on microbiological criteria, color and oxidation values of minced beef meat. Meat Sci 88(2):221–226. https://doi.org/10.1016/j.meatsci.2010.12.021
Yang NJ, Hinner MJ (2015) Getting across the cell membrane: an overview for small molecules, peptides, and proteins. Methods Mol Biol 1266:29–53. https://doi.org/10.1007/978-1-4939-2272-7_3
Daniels JA, Krishnamurthi R, Rizvi SSH (1985) A review of effects of carbon dioxide on microbial growth and food quality. J Food Prot 48(6):532–537. https://doi.org/10.4315/0362-028x-48.6.532
King A Jr, Nagel C (2006) Influence of carbon dioxide upon the metabolism of Pseudomonas aeruginosa. J Food Sci 40:362–366. https://doi.org/10.1111/j.1365-2621.1975.tb02202.x
Foster JW, Davis JB (1949) Carbon dioxide inhibition of anaerobic fumarate formation in the mold Rhizopus nigricans. Arch Biochem 21(1):135–142
Kolbeck S, Ludwig C, Meng C, Hilgarth M, Vogel RF (2020) Comparative proteomics of meat spoilage bacteria predicts drivers for their coexistence on modified atmosphere packaged meat. Front Microbiol 11 (209). doi:https://doi.org/10.3389/fmicb.2020.00209
King AD Jr, Nagel CW (1975) Influence of carbon dioxide upon the metabolism of Pseudomonas aeruginosa. J Food Sci 40(2):362–366. https://doi.org/10.1111/j.1365-2621.1975.tb02202.x
Jones RP, Greenfield PF (1982) Effect of carbon dioxide on yeast growth and fermentation. Enzyme Microb Technol 4(4):210–223. https://doi.org/10.1016/0141-0229(82)90034-5
Cabiscol E, Tamarit J, Ros J (2000) Oxidative stress in bacteria and protein damage by reactive oxygen species. Int Microbiol Off J Spanish Soc Microbiol 3(1):3–8
Condon S (1987) Responses of lactic acid bacteria to oxygen*. FEMS Microbiol Rev 3(3):269–280. https://doi.org/10.1111/j.1574-6968.1987.tb02465.x
Sies H (1993) Damage to plasmid DNA by singlet oxygen and its protection. Mutation Res/Gen Toxicol 299(3):183–191. https://doi.org/10.1016/0165-1218(93)90095-U
Zhang YM, Rock CO (2008) Membrane lipid homeostasis in bacteria. Nat Rev Microbiol 6(3):222–233. https://doi.org/10.1038/nrmicro1839
Cronan JE Jr, Gelmann EP (1975) Physical properties of membrane lipids: biological relevance and regulation. Bacteriol Rev 39(3):232–256
Hilgarth M, Behr J, Vogel RF (2018) Monitoring of spoilage-associated microbiota on modified atmosphere packaged beef and differentiation of psychrophilic and psychrotrophic strains. J Appl Microbiol 124(3):740–753. https://doi.org/10.1111/jam.13669
Höll L, Behr J, Vogel RF (2016) Identification and growth dynamics of meat spoilage microorganisms in modified atmosphere packaged poultry meat by MALDI-TOF MS. Food Microbiol 60:84–91. https://doi.org/10.1016/j.fm.2016.07.003
Hilgarth M, Lehner EM, Behr J, Vogel RF (2019) Diversity and anaerobic growth of Pseudomonas spp. isolated from modified atmosphere packaged minced beef. J Appl Microbiol 127 (1):159–174. doi:https://doi.org/10.1111/jam.14249
Kolbeck S, Reetz L, Hilgarth M, Vogel RF (2019) Quantitative oxygen consumption and respiratory activity of meat spoiling bacteria upon high oxygen modified atmosphere. Front Microbiol 10 (2398). doi:https://doi.org/10.3389/fmicb.2019.02398
Kolbeck S, Behr J, Vogel R, Ludwig C, Ehrmann M (2019) Acid stress response of Staphylococcus xylosus elicits changes in the proteome and cellular membrane. J Appl Microbiol. https://doi.org/10.1111/jam.14224
Santivarangkna C, Kulozik U, Kienberger H, Foerst P (2009) Changes in membrane fatty acids of Lactobacillus helveticus during vacuum drying with sorbitol. Lett Appl Microbiol 49(4):516–521. https://doi.org/10.1111/j.1472-765X.2009.02703.x
Behr J, Gänzle MG, Vogel RF (2006) Characterization of a highly hop-resistant <em>Lactobacillus brevis</em> Strain lacking hop transport. Appl Environ Microbiol 72(10):6483–6492. https://doi.org/10.1128/aem.00668-06
Parasassi T (1990) Phase fluctuation in phospholipid membranes revealed by Laurdan fluorescence. Biophys J 57:1179
Sträuber H, Müller S (2010) Viability states of bacteria: specific mechanisms of selected probes. Cytometry Part A 77A(7):623–634. https://doi.org/10.1002/cyto.a.20920
Klotz B, Mañas P, Mackey B (2010) The relationship between membrane damage, release of protein and loss of viability in Escherichia coli exposed to high hydrostatic pressure. Int J Food Microbiol 137:214–220. https://doi.org/10.1016/j.ijfoodmicro.2009.11.020
Sun J, Liu D (2003) Geometric models for calculating cell biovolume and surface area for phytoplankton. J Plank Res 25(11):1331–1346. https://doi.org/10.1093/plankt/fbg096
Aluise CD, Rose K, Boiani M, Reyzer ML, Manna JD, Tallman K, Porter NA, Marnett LJ (2013) Peptidyl-prolyl cis/trans-isomerase A1 (Pin1) is a target for modification by lipid electrophiles. Chem Res Toxicol 26(2):270–279. https://doi.org/10.1021/tx300449g
Sears DF, Eisenberg RM (1961) A model representing a physiological role of CO2 at the cell membrane. J Gen Physiol 44:869–887. https://doi.org/10.1085/jgp.44.5.869
Farber JM (1991) Microbiological aspects of modified-atmosphere packaging technology: a review. J Food Protect 54(1):58–70. https://doi.org/10.4315/0362-028X-54.1.58
Becker ZE (1936) A comparison between the action of carbonic acid and other acids upon the living cell. Protoplasma 25(1):161–175. https://doi.org/10.1007/BF01839067
Datta AR, Benjamin MM (1997) Factors controlling acid tolerance of Listeria monocytogenes: effects of nisin and other ionophores. Appl Environ Microbiol 63(10):4123–4126
Boyd DA, Cvitkovitch DG, Bleiweis AS, Kiriukhin MY, Debabov DV, Neuhaus FC, Hamilton IR (2000) Defects in D-alanyl-lipoteichoic acid synthesis in Streptococcus mutans results in acid sensitivity. J Bacteriol 182(21):6055–6065. https://doi.org/10.1128/jb.182.21.6055-6065.2000
Shabala L, Ross T (2008) Cyclopropane fatty acids improve Escherichia coli survival in acidified minimal media by reducing membrane permeability to H+ and enhanced ability to extrude H+. Res Microbiol 159:458–461. https://doi.org/10.1016/j.resmic.2008.04.011
Guerzoni ME, Lanciotti R, Cocconcelli PS (2001) Alteration in cellular fatty acid composition as a response to salt, acid, oxidative and thermal stresses in Lactobacillus helveticus. Microbiology 147(8):2255–2264. https://doi.org/10.1099/00221287-147-8-2255
Brown CM, Rose AH (1969) Fatty-acid composition of Candida utilis as affected by growth temperature and dissolved-oxygen tension. J Bacteriol 99(2):371–378
Bajpai P, Bajpai PK (1993) Eicosapentaenoic acid (EPA) production from microorganisms: a review. J Biotechnol 30(2):161–183. https://doi.org/10.1016/0168-1656(93)90111-Y
Erwin J, Bloch K (1964) Biosynthesis of unsaturated fatty acids in microorganisms. Science 143(3610):1006. https://doi.org/10.1126/science.143.3610.1006
Guerzoni ME, Ferruzzi M, Sinigaglia M, Criscuoli GC (1997) Increased cellular fatty acid desaturation as a possible key factor in thermotolerance in Saccharomyces cerevisiae. Can J Microbiol 43(6):569–576. https://doi.org/10.1139/m97-080
Dyrda G, Boniewska-Bernacka E, Man D, Barchiewicz K, Słota R (2019) The effect of organic solvents on selected microorganisms and model liposome membrane. Mol Biol Rep 46(3):3225–3232. https://doi.org/10.1007/s11033-019-04782-y
Nakae T (1995) Role of membrane permeability in determining antibiotic resistance in pseudomonas aeruginosa. Microbiol Immunol 39(4):221–229. https://doi.org/10.1111/j.1348-0421.1995.tb02193.x
Acknowledgements
This work was partially funded by the German Federal Ministry for Economic Affairs and Energy via the German Federation of Industrial Research Associations (AiF) and the Industry Association for Food Technology and Packaging (IVLV): Project Number IGF 19993N.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
SK designed the study, performed the experiments and data evaluation, and wrote the first draft of the manuscript. MH helped to draft the manuscript, data interpretation and supervised the work of SK. RV initiated the project and supervised the work of SK. HK and KK helped performing fatty acid acquisition and data analysis. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
No conflict of interest is declared by the authors.
Compliance with ethics requirements
This article does not contain any studies with human or animal subject.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kolbeck, S., Kienberger, H., Kleigrewe, K. et al. Effect of high levels of CO2 and O2 on membrane fatty acid profile and membrane physiology of meat spoilage bacteria. Eur Food Res Technol 247, 999–1011 (2021). https://doi.org/10.1007/s00217-020-03681-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00217-020-03681-y