Abstract
A new biosensing method is presented to detect gene mutation by integrating the MutS protein from bacteria with a fiber optic particle plasmon resonance (FOPPR) sensing system. In this method, the MutS protein is conjugated with gold nanoparticles (AuNPs) deposited on an optical fiber core surface. The target double-stranded DNA containing an A and C mismatched base pair in a sample can be captured by the MutS protein, causing increased absorption of green light launching into the fiber and hence a decrease in transmitted light intensity through the fiber. As the signal change is enhanced through consecutive total internal reflections along the fiber, the limit of detection for an AC mismatch heteroduplex DNA can be as low as 0.49 nM. Because a microfluidic chip is used to contain the optical fiber, the narrow channel width allows an analysis time as short as 15 min. Furthermore, the label-free and real-time nature of the FOPPR sensing system enables determination of binding affinity and kinetics between MutS and single-base mismatched DNA. The method has been validated using a heterozygous PCR sample from a patient to determine the allelic fraction. The obtained allelic fraction of 0.474 reasonably agrees with the expected allelic fraction of 0.5. Therefore, the MutS-functionalized FOPPR sensor may potentially provide a convenient quantitative tool to detect single nucleotide polymorphisms in biological samples with a short analysis time at the point-of-care sites.
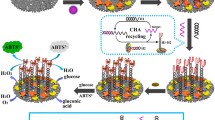
Graphical abstract






Similar content being viewed by others
Data availability
All data generated and analyses during this study are included in this published article and its supplementary material file.
Change history
04 May 2021
A Correction to this paper has been published: https://doi.org/10.1007/s00216-021-03353-0
References
Andrew AS, Gui J, Sanderson AC, Mason RA, Morlock EV, Schned AR, et al. Bladder cancer SNP panel predicts susceptibility and survival. Hum Genet. 2009;125:527–39. https://doi.org/10.1007/s00439-009-0645-6.
Yu L, Yin B, Qu K, Li J, Jin Q, Liu L, et al. Screening for susceptibility genes in hereditary non-polyposis colorectal cancer. Oncol Lett. 2018;15:9413–9. https://doi.org/10.3892/ol.2018.8504.
Fertrin KY, Costa FF. Genomic polymorphisms in sickle cell disease: implications for clinical diversity and treatment. Expert Rev Hematol. 2010;3:443–58. https://doi.org/10.1586/ehm.10.44.
Bare LA, Morrison AC, Rowland CM, Shiffman D, Luke MM, Iakoubova OA, et al. Five common gene variants identify elevated genetic risk for coronary heart disease. Gen Med. 2007;9:682–9. https://doi.org/10.1097/GIM.0b013e318156fb62.
Jonkers IH, Wijmenga C. Context-specific effects of genetic variants associated with autoimmune disease. Hum Mol Genet. 2017;26(R2):R185–92. https://doi.org/10.1093/hmg/ddx254.
Mardis ER. Next-generation DNA sequencing methods. Annu Rev Genom Hum Genet. 2008;9:387–402. https://doi.org/10.1146/annurev.genom.9.081307.164359.
Shendure J, Balasubramanian S, Church GM, Gilbert W, Rogers J, Schloss JA, et al. DNA sequencing at 40: past, present and future. Nature. 2017;550:345–53. https://doi.org/10.1038/nature24286.
Samura O. Update on noninvasive prenatal testing: a review based on current worldwide research. J Obstet Gynaecol Res. 2020;46:1246–54. https://doi.org/10.1111/jog.14268.
Zhang J, Li J, Saucier JB, Feng Y, Jiang Y, Sinson J, et al. Non-invasive prenatal sequencing for multiple Mendelian monogenic disorders using circulating cell-free fetal DNA. Nat Med. 2019;25:439–47. https://doi.org/10.1038/s41591-018-0334-x.
Fahy E, Nazarbaghi R, Zomorrodi M, Herrnstadt C, Parker WD, Davis RE, et al. Multiplex fluorescence-based primer extension method for quantitative mutation analysis of mitochondrial DNA and its diagnostic application for Alzheimer’s disease. Nucleic Acids Res. 1997;25:3102–9. https://doi.org/10.1093/nar/25.15.3102.
Chen YT, Hsu CL, Hou SY. Detection of single-nucleotide polymorphisms using gold nanoparticles and single-strand-specific nucleases. Anal Biochem. 2008;375:299–305. https://doi.org/10.1016/j.ab.2007.12.036.
Xu W, Xue X, Li T, Zeng H, Liu X. Ultrasensitive and selective colorimetric DNA detection by nicking endonuclease assisted nanoparticle amplification. Angew Chem Int Ed. 2009;48:6849–52. https://doi.org/10.1002/anie.200901772.
Deng M, Zhang D, Zhou Y, Zhou X. Highly effective colorimetric and visual detection of nucleic acids using an asymmetrically split peroxidase DNAzyme. J Am Chem Soc. 2008;130:13095–102. https://doi.org/10.1021/ja803507d.
Lapitan LDS Jr, Xu Y, Guo Y, Zhou D. Combining magnetic nanoparticle capture and poly-enzyme nanobead amplification for ultrasensitive detection and discrimination of DNA single nucleotide polymorphisms. Nanoscale. 2019;11:1195–204. https://doi.org/10.1039/C8NR07641C.
Gotoh M, Hasebe M, Ohira T, Hasegawa Y, Shinohara Y, Sota H, et al. Rapid method for detection of point mutations using mismatch binding protein (MutS) and an optical biosensor. Gen Anal Biomol Eng. 1997;14:47–50. https://doi.org/10.1016/S1050-3862(97)00009-0.
Ma X, Truong PL, Ahn NH, Sim SJ. Single gold nanoplasmonic sensor for clinical cancer diagnosis based on specific interaction between nucleic acids and protein. Biosens Bioelectron. 2015;67:59–65. https://doi.org/10.1016/j.bios.2014.06.038.
Lishanski A, Ostrander EA, Rine J. Mutation detection by mismatch binding protein, MutS, in amplified DNA: application to the cystic fibrosis gene. Proc Natl Acad Sci U S A. 1994;91:2674–8. https://doi.org/10.1073/pnas.91.7.2674.
Wagner R, Debble P, Radman M. Mutation detection using immobilized mismatch binding protein (MutS). Nucleic Acids Res. 1995;23:3944–8. https://doi.org/10.1093/nar/23.19.3944.
Su SS, Lahue RS, Au KG, Modrich P. Mispair specificity of methyl-directed DNA mismatch correction in vitro. J Biol Chem. 1988;263:6829–35. https://doi.org/10.1016/S0021-9258(18)68718-6.
Wang H, Yang Y, Schofield MJ, Du C, Fridman Y, Lee SD, et al. DNA bending and unbending by MutS govern mismatch recognition and specificity. Proc Natl Acad Sci U S A. 2003;100:14822–7. https://doi.org/10.1073/pnas.2433654100.
Stanislawska-Sachadyn A, Sachadyn P. MutS as a tool for mutation detection. Acta Biochim Polon. 2005;52:575–83. https://doi.org/10.18388/abp.2005_3417.
Brown J, Brown T, Fox KR. Affinity of mismatch-binding protein MutS for heteroduplexes containing different mismatches. Biochem J. 2001;354:627–33. https://doi.org/10.1042/0264-6021:3540627.
Behrensdorf HA, Pignot M, Windhab N, Kappel A. Rapid parallel mutation scanning of gene fragments using a microelectronic protein-DNA chip format. Nucleic Acids Res. 2002;30:e64. https://doi.org/10.1093/nar/gnf063.
Palecek E, Masarík M, Kizek R, Kuhlmeier D, Hassmann J, Schulein J. Sensitive electrochemical determination of unlabeled MutS protein and detection of point mutations in DNA. Anal Chem. 2004;76:5930–6. https://doi.org/10.1021/ac049474x.
Han A, Takarada T, Shibata T, Nakayama M, Maeda M. A MutS protein-immobilized Au electrode for detecting single-base mismatch of DNA. Anal Sci. 2006;22:663–6. https://doi.org/10.2116/analsci.22.663.
Chen H, Liu XJ, Liu YL, Jiang JH, Shen GL, Yu RQ. Electrochemical scanning of DNA point mutations via MutS protein-mediated mismatch recognition. Biosens Bioelectron. 2009;24:1955–61. https://doi.org/10.1016/j.bios.2008.09.029.
Li CZ, Long YT, Lee JS, Kraatz HB. Protein-DNA interaction: impedance study of MutS binding to a DNA mismatch. Chem Commun. 2004:574–5. https://doi.org/10.1039/B314642A.
Gong H, Zhong T, Gao L, Li X, Bi L, Kraatz HB. Unlabeled hairpin DNA probe for electrochemical detection of single-nucleotide mismatches based on MutS−DNA interactions. Anal Chem. 2009;81:8639–43. https://doi.org/10.1021/ac901371n.
Cho M, Lee S, Han SY, Park JY, Rahman MA, Shim YB, et al. Electrochemical detection of mismatched DNA using a MutS probe. Nucleic Acids Res. 2006;34:e75. https://doi.org/10.1093/nar/gkl364.
Kim S, Kim TG, Byon HR, Shin HJ, Ban C, Choi HC. Recognition of single mismatched DNA using MutS-immobilized carbon nanotube field effect transistor devices. J Phys Chem B. 2009;113:12164–8. https://doi.org/10.1021/jp9063559.
Su X, Robelek R, Wu Y, Wang G, Knoll W. Detection of point mutation and insertion mutations in DNA using a quartz crystal microbalance and MutS, a mismatch binding protein. Anal Chem. 2004;76:489–94. https://doi.org/10.1021/ac035175g.
Wilson PK, Jiang T, Minunni ME, Turner AFP, Mascini M. A novel optical biosensor format for the detection of clinically relevant TP53 mutations. Biosens Bioelectron. 2005;20:2310–3. https://doi.org/10.1016/j.bios.2004.11.020.
Ma X, Song S, Kim S, Kwon MS, Lee H, Park W, et al. Single gold-bridged nanoprobes for identification of single point DNA mutations. Nat Commun. 2019;10:836. https://doi.org/10.1038/s41467-019-08769-y.
Cheng SF, Chau LK. Colloidal gold-modified optical fiber for chemical and biochemical sensing. Anal Chem. 2003;75:16–21. https://doi.org/10.1021/ac020310v.
Chiang CY, Hsieh ML, Huang KW, Chau LK, Chang CM, Lyu SR. Fiber optic particle plasmon resonance sensor for detection of interleukin-1β in synovial fluids. Biosens Bioelectron. 2010;26:1036–42. https://doi.org/10.1016/j.bios.2010.08.047.
Wu CW, Chiang CY, Chen CH, Chiang CS, Wang CT, Chau LK. Self-referencing fiber optic particle plasmon resonance sensing system for real-time biological monitoring. Talanta. 2016;146:291–8. https://doi.org/10.1016/j.talanta.2015.08.047.
Wu WT, Chen CH, Chiang CY, Chau LK. Effect of surface coverage of gold nanoparticles on the refractive index sensitivity in fiber-optic nanoplasmonic sensing. Sensors. 2018;18:1759. https://doi.org/10.3390/s18061759.
Chau LK, Lin YF, Cheng SF, Lin TJ. Fiber-optic chemical and biochemical probes based on localized surface plasmon resonance. Sens Actuat B. 2006;113:100–5. https://doi.org/10.1016/j.snb.2005.02.034.
Origa R. β-Thalassemia. Genet Med. 2017;19:609–19. https://doi.org/10.1038/gim.2016.173.
Wang HC, Hsieh LL, Liu YC, Hsiao HH, Lin SK, Tsai WC, et al. The epidemiologic transition of thalassemia and associated hemoglobinopathies in southern Taiwan. Ann Hematol. 2017;96:183–8. https://doi.org/10.1007/s00277-016-2868-7.
Babic I, Andrew SE, Jirik FR. MutS interaction with mismatch and alkylated base containing DNA molecules detected by optical biosensor. Mutat Res. 1996;372:87–96. https://doi.org/10.1016/S0027-5107(96)00170-4.
Tseng YT, Li WY, Yu YW, Chiang CY, Liu SQ, Chau LK, et al. Fiber optic particle plasmon resonance biosensor for label-free detection of nucleic acids and its application to HLA-B27 mRNA detection in patients with ankylosing spondylitis. Sensors. 2020;20:3137. https://doi.org/10.3390/s20113137.
Chaudhari PP, Chau LK, Tseng YT, Huang CJ, Chen YL. A fiber optic nanoplasmonic biosensor for the sensitive detection of ampicillin and its analogs. Microchim Acta. 2020;187:396. https://doi.org/10.1007/s00604-020-04381-w.
Chang TC, Wu CC, Wang SC, Chau LK, Hsieh WH. Using a fiber optic particle plasmon resonance biosensor to determine kinetic constants of antigen-antibody binding reaction. Anal Chem. 2013;85:245–50. https://doi.org/10.1021/ac302590n.
Yu SN, Tsai TH, Chau LK. Optical biosensing system and method thereof. US Patent Appl Pub: US 2021/0011011 A1.
Hsu WT, Hsieh WH, Cheng SF, Jen CP, Wu CC, Li CH, et al. Integration of fiber optic-particle plasmon resonance biosensor with microfluidic chip. Anal Chim Acta. 2011;697:75–82. https://doi.org/10.1016/j.aca.2011.04.023.
Chiang CY, Huang TT, Wang CH, Huang CJ, Tsai TH, Yu SN, et al. Fiber optic nanogold-linked immunosorbent assay for rapid detection of procalcitonin at femtomolar concentration level. Biosens Bioelectron. 2020;151:111871. https://doi.org/10.1016/j.bios.2019.111871.
Sharma A, Doucette C, Biro FN, Hingorani MM. Slow conformational changes in MutS and DNA direct ordered transitions between mismatch search, recognition and signaling of DNA repair. J Mol Biol. 2013;425:4192–205. https://doi.org/10.1016/j.jmb.2013.08.011.
Funding
This work was supported by the Ministry of Science and Technology of Taiwan (Grants MOST 105-2113-M-194-009-MY3 and MOST 107-2119-M-194-001) and Center for Nano Bio-Detection from the Featured Research Areas College Development Plan of National Chung Cheng University.
Author information
Authors and Affiliations
Contributions
Conceptualization: Lai-Kwan Chau, Tze-Ta Huang. Methodology: Lai-Kwan Chau, Tze-Ta Huang. Resources: Pao-Lin Kuo. Investigation: Loan Thi Ngo, Wei-Kai Wang. Formal analysis: Loan Thi Ngo, Yen-Ta Tseng, Ting-Chou Chang. Writing-original draft: Loan Thi Ngo, Yen-Ta Tseng. Writing-review and editing: Lai-Kwan Chau, Tze-Ta Huang. Funding acquisition: Lai-Kwan Chau.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
This study was reviewed and approved by the Institutional Review Board (IRB) of National Cheng Kung University Hospital. The patients/participants provided their written informed consent to participate in this study.
Source of biological material
National Cheng Kung University Hospital.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: Unfortunately, during production the typesetter mistakenly uploaded the wrong file as the graphical abstract figure with black unreadable background color.
Supplementary information
ESM 1
(PDF 336 kb)
Rights and permissions
About this article
Cite this article
Ngo, L.T., Wang, WK., Tseng, YT. et al. MutS protein-based fiber optic particle plasmon resonance biosensor for detecting single nucleotide polymorphisms. Anal Bioanal Chem 413, 3329–3337 (2021). https://doi.org/10.1007/s00216-021-03271-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-021-03271-1