Abstract
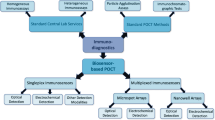
Immunodiagnostic tests performed at the point of care (POC) today usually employ antibodies for biorecognition and are read out either visually or with specialized equipment. Availability of alternative biorecognition elements with promising features as well as smartphone-based approaches for signal readout, however, challenge the described established configuration in terms of analytical performance and practicability. Assessing these developments’ clinical relevance and their impact on POC immunodiagnostics is demanding. The first part of this review will therefore give an overview on suitable diagnostic biosensors based on alternative recognition elements (such as nucleic acid-based aptamers or engineered binding proteins) and exemplify advantages and drawbacks of these biomolecules on the base of selected assays. The second part of the review then focuses on smartphone-connected diagnostics and discusses the indispensable considerations required for successful future clinical POCT implementation. Together, the joint depiction of two of the most innovative and exciting developments in the field will enable the reader to cast a glance into the distant future of POC immunodiagnostics.

Similar content being viewed by others
Abbreviations
- Ab:
-
Antibody
- CBP:
-
Calcium-binding protein
- CLIP:
-
Combinatorial library of improved peptide
- DARPins:
-
Designed ankyrin repeat proteins
- IVD:
-
In vitro diagnostics
- LFA:
-
Lateral flow immunoassays
- LFD:
-
Lateral flow device
- MW:
-
Molecular weight
- MIP:
-
Molecular imprinted polymer
- SAW:
-
Surface acoustic wave
- SELEX:
-
Systematic evolution of ligands by exponential enrichment
- SPR:
-
Surface plasmon resonance
References
Ko Ferrigno P. Non-antibody protein-based biosensors. Essays Biochem. 2016;60(1):19–25. https://doi.org/10.1042/EBC20150003.
Bradbury A, Plückthun A. Reproducibility: standardize antibodies used in research. Nature. 2015;518(7537):27–9. https://doi.org/10.1038/518027a.
Luppa PB, Bietenbeck A, Beaudoin C, Giannetti A. Clinically relevant analytical techniques, organizational concepts for application and future perspectives of point-of-care testing. Biotech Adv. 2016;34(3):139–60. https://doi.org/10.1016/j.biotechadv.2016.01.003.
Vashist SK, Luppa PB, Yeo LY, Ozcan A, Luong JHT. Emerging technologies for next-generation point-of-care testing. Trends Biotechnol. 2015;33(11):692–705. https://doi.org/10.1016/j.tibtech.2015.09.001.
Chen A, Yang S. Replacing antibodies with aptamers in lateral flow immunoassay. Biosens Bioelectron. 2015;71:230–42. https://doi.org/10.1016/j.bios.2015.04.041.
Li Z, Chen GY. Current conjugation methods for immunosensors. Nanomaterials (Basel). 2018;8(5). https://doi.org/10.3390/nano8050278.
Ellington AD, Szostak JW. In vitro selection of RNA molecules that bind specific ligands. Nature. 1990;346(6287):818–22. https://doi.org/10.1038/346818a0.
Iliuk AB, Hu L, Tao WA. Aptamer in bioanalytical applications. Anal Chem. 2011;83(12):4440–52. https://doi.org/10.1021/ac201057w.
Stenken JA, Poschenrieder AJ. Bioanalytical chemistry of cytokines - a review. Anal Chim ASepcta. 2015;853:95–115. https://doi.org/10.1016/j.aca.2014.10.009.
Gopinath SC, Lakshmipriya T, Chen Y, Phang WM, Hashim U. Aptamer-based 'point-of-care testing'. Biotech Adv. 2016;34(3):198–208. https://doi.org/10.1016/j.biotechadv.2016.02.003.
Nery AA, Wrenger C, Ulrich H. Recognition of biomarkers and cell-specific molecular signatures: aptamers as capture agents. J Sep Sci. 2009;32(10):1523–30. https://doi.org/10.1002/jssc.200800695.
Fang X, Tan W. Aptamers generated from cell-SELEX for molecular medicine: a chemical biology approach. Acc Chem Res. 2010;43(1):48–57. https://doi.org/10.1021/ar900101s.
Guthrie JW, Hamula CL, Zhang H, Le XC. Assays for cytokines using aptamers. Methods. 2006;38(4):324–30. https://doi.org/10.1016/j.ymeth.2006.01.001.
Nezlin R. Use of aptamers in immunoassays. Mol Immunol. 2016;70:149–54. https://doi.org/10.1016/j.molimm.2015.12.009.
Seo HB, Gu MB. Aptamer-based sandwich-type biosensors. J Biol Eng. 2017;11:11. https://doi.org/10.1186/s13036-017-0054-7.
Zainol Abidin AS, Rahim RA, Md Arshad MK, Fatin Nabilah MF, Voon CH, Tang TH, et al. Current and potential developments of cortisol aptasensing towards point-of-care diagnostics (POCT). Sensors (Basel). 2017;17(5). https://doi.org/10.3390/s17051180.
Sanghavi BJ, Moore JA, Chavez JL, Hagen JA, Kelley-Loughnane N, Chou CF, et al. Aptamer-functionalized nanoparticles for surface immobilization-free electro-chemical detection of cortisol in a microfluidic device. Biosens Bioelectron. 2016;78:244–52. https://doi.org/10.1016/j.bios.2015.11.044.
Bruno JG. Application of DNA aptamers and quantum dots to lateral flow test strips for detection of foodborne pathogens with improved sensitivity versus colloidal gold. Pathogens. 2014;3(2):341–55. https://doi.org/10.3390/pathogens3020341.
Wang L, Wu A, Wei G. Graphene-based aptasensors: from molecule-interface interactions to sensor design and biomedical diagnostics. Analyst. 2018;143(7):1526–43. https://doi.org/10.1039/c8an00081f.
Sze JYY, Ivanov AP, Cass AEG, Edel JB. Single molecule multiplexed nanopore protein screening in human serum using aptamer modified DNA carriers. Nat Commun. 2017;8(1):1552. https://doi.org/10.1038/s41467-017-01584-3.
Binz HK, Amstutz P, Pluckthun A. Engineering novel binding proteins from nonimmunoglobulin domains. Nat Biotechnol. 2005;23(10):1257–68. https://doi.org/10.1038/nbt1127.
Martin HL, Bedford R, Heseltine SJ, Tang AA, Haza KZ, Rao A, et al. Non-immunoglobulin scaffold proteins: precision tools for studying protein-protein interactions in cancer. New Biotechnol. 2018;45:28–35. https://doi.org/10.1016/j.nbt.2018.02.008.
Reverdatto S, Burz DS, Shekhtman A. Peptide aptamers: development and applications. Curr Top Med Chem. 2015;15(12):1082–101.
Colas P, Cohen B, Jessen T, Grishina I, McCoy J, Brent R. Genetic selection of peptide aptamers that recognize and inhibit cyclin-dependent kinase 2. Nature. 1996;380(6574):548–50. https://doi.org/10.1038/380548a0.
Reverdatto S, Rai V, Xue J, Burz DS, Schmidt AM, Shekhtman A. Combinatorial library of improved peptide aptamers, clips to inhibit rage signal transduction in mammalian cells. PLoS One. 2013;8(6):e65180. https://doi.org/10.1371/journal.pone.0065180.
Koide A, Koide S. Monobodies: antibody mimics based on the scaffold of the fibronectin type iii domain. Methods Mol Biol. 2007;352:95–109. https://doi.org/10.1385/1-59745-187-8:95.
Skerra A. Alternative binding proteins: anticalins - harnessing the structural plasticity of the lipocalin ligand pocket to engineer novel binding activities. FEBS J. 2008;275(11):2677–83. https://doi.org/10.1111/j.1742-4658.2008.06439.x.
Richter A, Skerra A. Anticalins directed against vascular endothelial growth factor receptor 3 (VEGFR-3) with picomolar affinities show potential for medical therapy and in vivo imaging. Biol Chem. 2017;398(1):39–55. https://doi.org/10.1515/hsz-2016-0195.
Amstutz P, Koch H, Binz HK, Deuber SA, Plückthun A. Rapid selection of specific MAP kinase-binders from designed ankyrin repeat protein libraries. Protein Eng Des Sel. 2006;19(5):219–29. https://doi.org/10.1093/protein/gzl004.
Parizek P, Kummer L, Rube P, Prinz A, Herberg FW, Plückthun A. Designed ankyrin repeat proteins (DARPins) as novel isoform-specific intracellular inhibitors of c-Jun N-terminal kinases. ACS Chem Biol. 2012;7(8):1356–66. https://doi.org/10.1021/cb3001167.
Plückthun A. Designed ankyrin repeat proteins (DARPins): binding proteins for research, diagnostics, and therapy. Annu Rev Pharmacol Toxicol. 2015;55:489–511. https://doi.org/10.1146/annurev-pharmtox-010611-134654.
Nord K, Nilsson J, Nilsson B, Uhlen M, Nygren PA. A combinatorial library of an alpha-helical bacterial receptor domain. Protein Eng. 1995;8(6):601–8.
Löfblom J, Feldwisch J, Tolmachev V, Carlsson J, Stahl S, Frejd FY. Affibody molecules: engineered proteins for therapeutic, diagnostic and biotechnological applications. FEBS Lett. 2010;584(12):2670–80. https://doi.org/10.1016/j.febslet.2010.04.014.
Saylan Y, Akgonullu S, Yavuz H, Unal S, Denizli A. Molecularly imprinted polymer based sensors for medical applications. Sensors (Basel). 2019;19(6). https://doi.org/10.3390/s19061279.
Morales MA, Halpern JM. Guide to selecting a biorecognition element for biosensors. Bioconjug Chem. 2018;29(10):3231–9. https://doi.org/10.1021/acs.bioconjchem.8b00592.
Harmansa S, Affolter M. Protein binders and their applications in developmental biology. Development. 2018;145(2). https://doi.org/10.1242/dev.148874.
Yu X, Yang YP, Dikici E, Deo SK, Daunert S. Beyond antibodies as binding partners: the role of antibody mimetics in bioanalysis. Ann Rev Anal Chem. 2017;10(1):293–320. https://doi.org/10.1146/annurev-anchem-061516-045205.
Cerwall P, Lundvall A, Jonsson P, Carson S, Svenningsson R, Lindberg P, Öhman K, Sandin T, Rangel L, Karapantelakis A, Elmgren S, Wieweg L, Halen M, Edstam J, Queirós R, Muller F, Englund L (2018) Ericsson mobility report.
Reusche R, Buchanan PJ, Kozlow JH, Vercler CJ. A systematic review of smartphone applications for plastic surgery providers: target audience, uses, and cost. Ann Plas Surg. 2016;77(1):6–12. https://doi.org/10.1097/sap.0000000000000792.
Jahanshir A, Karimialavijeh E, Sheikh H, Vahedi M, Momeni M. Smartphones and medical applications in the emergency department daily practice. Emergency. 2017;5(1):e14.
Valle J, Godby T, Paul DP, 3rd, Smith H, Coustasse A (2017) Use of smartphones for clinical and medical education. Health Care Manag 36 (3):293–300. doi:https://doi.org/10.1097/hcm.0000000000000176.
Vashist SK, Luong JHT. Chapter 16 - smartphone-based immunoassays. In: Vashist SK, Luong JHT, editors. Handbook of immunoassay technologies: Academic Press; 2018. p. 433–53. https://doi.org/10.1016/B978-0-12-811762-0.00016-5.
Xu X, Akay A, Wei H, Wang S, Pingguan-Murphy B, Erlandsson BE, et al. Advances in smartphone-based point-of-care diagnostics. Proc IEEE. 2015;103(2):236–47. https://doi.org/10.1109/JPROC.2014.2378776.
Yang K, Peretz-Soroka H, Liu Y, Lin F. Novel developments in mobile sensing based on the integration of microfluidic devices and smartphones. Lab Chip. 2016;16(6):943–58. https://doi.org/10.1039/c5lc01524c.
Zarei M. Advances in point-of-care technologies for molecular diagnostics. Biosens Bioelectron. 2017;98:494–506. https://doi.org/10.1016/j.bios.2017.07.024.
Xu D, Huang X, Guo J, Ma X. Automatic smartphone-based microfluidic biosensor system at the point of care. Biosens Bioelectron. 2018;110:78–88. https://doi.org/10.1016/j.bios.2018.03.018.
Zhang D, Liu Q. Biosensors and bioelectronics on smartphone for portable biochemical detection. Biosens Bioelectron. 2016;75:273–84. https://doi.org/10.1016/j.bios.2015.08.037.
Romeo A, Leung TS, Sanchez S. Smart biosensors for multiplexed and fully integrated point-of-care diagnostics. Lab Chip. 2016;16(11):1957–61. https://doi.org/10.1039/c6lc90046a.
Huang X, Xu D, Chen J, Liu J, Li Y, Song J, et al. Smartphone-based analytical biosensors. Analyst. 2018;143(22):5339–51. https://doi.org/10.1039/c8an01269e.
Park JN, Paek SH, Kim DH, Seo SM, Lim GS, Kang JH, et al. Conformation-sensitive antibody-based point-of-care immunosensor for serum ca(2+) using two-dimensional sequential binding reactions. Biosens Bioelectron. 2016;85:611–7. https://doi.org/10.1016/j.bios.2016.05.061.
Lee S, O'Dell D, Hohenstein J, Colt S, Mehta S, Erickson D. Nutriphone: a mobile platform for low-cost point-of-care quantification of vitamin B12 concentrations. Sci Rep. 2016;6:28237. https://doi.org/10.1038/srep28237.
Srinivasan B, O'Dell D, Finkelstein JL, Lee S, Erickson D, Mehta S. Ironphone: mobile device-coupled point-of-care diagnostics for assessment of iron status by quantification of serum ferritin. Biosens Bioelectron. 2018;99:115–21. https://doi.org/10.1016/j.bios.2017.07.038.
Paterson AS, Raja B, Mandadi V, Townsend B, Lee M, Buell A, et al. A low-cost smartphone-based platform for highly sensitive point-of-care testing with persistent luminescent phosphors. Lab Chip. 2017;17(6):1051–9. https://doi.org/10.1039/c6lc01167e.
Fan Y, Liu J, Wang Y, Luo J, Xu H, Xu S, et al. A wireless point-of-care testing system for the detection of neuron-specific enolase with microfluidic paper-based analytical devices. Biosens Bioelectron. 2017;95:60–6. https://doi.org/10.1016/j.bios.2017.04.003.
Vashist SK, Schneider EM, Venkatesh AG, Luong JHT. Emerging human fetuin A assays for biomedical diagnostics. Trends Biotechnol. 2017;35(5):407–21. https://doi.org/10.1016/j.tibtech.2016.12.006.
Vashist SK, van Oordt T, Schneider EM, Zengerle R, von Stetten F, Luong JHT. A smartphone-based colorimetric reader for bioanalytical applications using the screen-based bottom illumination provided by gadgets. Biosens Bioelectron. 2015;67:248–55. https://doi.org/10.1016/j.bios.2014.08.027.
Liu Y, Liu Q, Chen S, Cheng F, Wang H, Peng W. Surface plasmon resonance biosensor based on smart phone platforms. Sci Rep. 2015;5:12864. https://doi.org/10.1038/srep12864.
Battaglia TM, Masson JF, Sierks MR, Beaudoin SP, Rogers J, Foster KN, et al. Quantification of cytokines involved in wound healing using surface plasmon resonance. Anal Chem. 2005;77(21):7016–23. https://doi.org/10.1021/ac050568w.
Masson JF, Battaglia TM, Khairallah P, Beaudoin S, Booksh KS. Quantitative measurement of cardiac markers in undiluted serum. Anal Chem. 2007;79(2):612–9. https://doi.org/10.1021/ac061089f.
Ross GMS, Bremer M, Nielen MWF. Consumer-friendly food allergen detection: moving towards smartphone-based immunoassays. Anal Bioanal Chem. 2018. https://doi.org/10.1007/s00216-018-0989-7.
Bedin F, Boulet L, Voilin E, Theillet G, Rubens A, Rozand C. Paper-based point-of-care testing for cost-effective diagnosis of acute Flavivirus infections. J Med Virol. 2017;89(9):1520–7. https://doi.org/10.1002/jmv.24806.
Kaushik A, Yndart A, Kumar S, Jayant RD, Vashist A, Brown AN, et al. A sensitive electrochemical immunosensor for label-free detection of Zika-virus protein. Sci Rep. 2018;8(1):9700. https://doi.org/10.1038/s41598-018-28035-3.
Turbé V, Gray ER, Lawson VE, Nastouli E, Brookes JC, Weiss RA, et al. Towards an ultra-rapid smartphone- connected test for infectious diseases. Sci Rep. 2017;7(1):11971. https://doi.org/10.1038/s41598-017-11887-6.
Jiang H, Wu D, Song L, Yuan Q, Ge S, Min X, et al. A smartphone-based genotyping method for hepatitis B virus at point-of-care settings. SLAS Technol. 2017;22(2):122–9. https://doi.org/10.1177/2211068216680163.
Berg B, Cortazar B, Tseng D, Ozkan H, Feng S, Wei Q, et al. Cellphone-based hand-held microplate reader for point-of-care testing of enzyme-linked immunosorbent assays. ACS Nano. 2015;9(8):7857–66. https://doi.org/10.1021/acsnano.5b03203.
Laksanasopin T, Guo TW, Nayak S, Sridhara AA, Xie S, Olowookere OO, et al. A smartphone dongle for diagnosis of infectious diseases at the point of care. Sci Transl Med. 2015;7(273):273re271. https://doi.org/10.1126/scitranslmed.aaa0056.
Herbst de Cortina S, Bristow CC, Humphries R, Vargas SK, Konda KA, Caceres CF, et al. Laboratory evaluation of a smartphone-based electronic reader of rapid dual point-of-care tests for antibodies to human immunodeficiency virus and Treponema pallidum infections. Sex Transm Dis. 2017;44(7):412–6. https://doi.org/10.1097/olq.0000000000000628.
Yeo SJ, Choi K, Cuc BT, Hong NN, Bao DT, Ngoc NM, et al. Smartphone-based fluorescent diagnostic system for highly pathogenic H5N1 viruses. Theranostics. 2016;6(2):231–42. https://doi.org/10.7150/thno.14023.
Cho S, Park TS, Nahapetian TG, Yoon JY. Smartphone-based, sensitive micropad detection of urinary tract infection and gonorrhea. Biosens Bioelectron. 2015;74:601–11. https://doi.org/10.1016/j.bios.2015.07.014.
Lin B, Yu Y, Cao Y, Guo M, Zhu D, Dai J, et al. Point-of-care testing for streptomycin based on aptamer recognizing and digital image colorimetry by smartphone. Biosens Bioelectron. 2018;15(100):482–9.
Ganguli A, Ornob A, Yu H, Damhorst GL, Chen W, Sun F, et al. Hands-free smartphone-based diagnostics for simultaneous detection of Zika, Chikungunya, and Dengue at point-of-care. Biomed Microdevices. 2017;19(4):73. https://doi.org/10.1007/s10544-017-0209-9.
Kotz D, Fu K, Gunter C, Rubin A. Security for mobile and cloud frontiers in healthcare. Commun ACM. 2015;58(8):21–3. https://doi.org/10.1145/2790830.
Kolmar H, Skerra A. Alternative binding proteins get mature: rivalling antibodies. FEBS J. 2008;275(11):2667. https://doi.org/10.1111/j.1742-4658.2008.06437.x.
Bruno JG. Predicting the uncertain future of aptamer-based diagnostics and therapeutics. Molecules. 2015;20(4):6866–87. https://doi.org/10.3390/molecules20046866.
Funding
This work was financially supported in part by the European Commission (NANODEM, #318372) and the German Bundesministerium für Bildung und Forschung (Q-Flow, #13N13867; and KAREL, #13GW0154D).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Published in the topical collection New Developments in Biosensors with guest editors Francesco Baldini and Maria Minunni.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Thaler, M., Luppa, P.B. Highly sensitive immunodiagnostics at the point of care employing alternative recognition elements and smartphones: hype, trend, or revolution?. Anal Bioanal Chem 411, 7623–7635 (2019). https://doi.org/10.1007/s00216-019-01974-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-01974-0