Abstract
Improvements in mass spectrometry technology to include instrument duty cycle, resolution, and sensitivity suggest mass spectrometry as a highly competitive alternative to conventional microbiological proteomic techniques. Targeted mass spectral analysis, sans prior empirical measurements, has begun to solely use the enormous amount of available genomic information for assay development. An in silico tryptic digestion of a suspected antibiotic-resistant enzyme using only its genomic information for assay development was achieved. Both MRM and full-scan MS2 independent data acquisitions were obtained for an antibiotic-resistance microbe not previously measured using mass spectrometry. In addition, computation methods to determine highest responding peptides in positive ion mode liquid chromatography-mass spectrometry (LC-MS) were evaluated. Employment of the relative retention time (iRT) concept using a homemade peptide standard set revealed facile method transfer between two fundamental different mass spectral platforms: an ultra-high-pressure liquid chromatography triple quadrupole-mass spectrometer (UHPLC-MS) and nano-liquid chromatography parallel reaction monitoring (nano-LC-PRM) hybrid quadrupole orbitrap Q-exactive mass spectrometer supporting easy dissemination and rapid method implementation between laboratories.

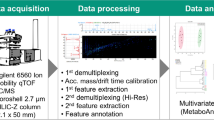
Graphical Abstract



Similar content being viewed by others
References
Peng J, Elias JE, Thoreen CC, Licklider LJ, Gygi SP. Evaluation of multidimensional chromatography coupled with tandem mass spectrometry (LC/LC− MS/MS) for large-scale protein analysis: the yeast proteome. J Proteome Res. 2003;2(1):43–50.
Gillet LC, Navarro P, Tate S, Röst H, Selevsek N, Reiter L, et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteomics. 2012;11(6):O111 016717.
Kelleher NL. Peer reviewed: top-down proteomics. Anal Chem. 2004;76(11):196 A–203 A. https://doi.org/10.1021/ac0415657.
Eidhammer I, Flikka K, Martens L, Mikalsen SO. Computational methods for mass spectrometry proteomics. Chichester; Hoboken: John Wiley & Sons; 2007.
Lane NM, Gregorich ZR, Ge Y. Top-down proteomics. In: Agnetti G, Lindsey M, Foster D, editors. Manual of cardiovascular proteomics. Cham: Springer; 2016.
Zhang Y, Fonslow BR, Shan B, Baek M-C, Yates JR III. Protein analysis by shotgun/bottom-up proteomics. Chem Rev. 2013;113(4):2343–94.
Bogdanov B, Smith RD. Proteomics by FTICR mass spectrometry: top down and bottom up. Mass Spectrom Rev. 2005;24(2):168–200.
Brock A, Horn DM, Peters EC, Shaw CM, Ericson C, Phung QT, et al. An automated matrix-assisted laser desorption/ionization quadrupole Fourier transform ion cyclotron resonance mass spectrometer for “bottom-up” proteomics. Anal Chem. 2003;75(14):3419–28.
Kalló G, Chatterjee A, Tóth M, Rajnavölgyi É, Csutak A, Tőzsér J, et al. Relative quantification of human β-defensins by a proteomics approach based on selected reaction monitoring. Rapid Commun Mass Spectrom. 2015;29(18):1623–31.
Angel TE, Aryal UK, Hengel SM, Baker ES, Kelly RT, Robinson EW, et al. Mass spectrometry-based proteomics: existing capabilities and future directions. Chem Soc Rev. 2012;41(10):3912–28.
Chen G, Pramanik BN. Application of LC/MS to proteomics studies: current status and future prospects. Drug Discov Today. 2009;14(9–10):465–71.
Metzker ML. Sequencing technologies—the next generation. Nat Rev Genet. 2010;11(1):31.
Deutsch EW, Lam H, Aebersold R. PeptideAtlas: a resource for target selection for emerging targeted proteomics workflows. EMBO Rep. 2008;9(5):429–34.
Lange V, Picotti P, Domon B, Aebersold R. Selected reaction monitoring for quantitative proteomics: a tutorial. Mol Syst Biol. 2008;4(1):222.
Charretier Y, Dauwalder O, Franceschi C, Degout-Charmette E, Zambardi G, Cecchini T, et al. Rapid bacterial identification, resistance, virulence and type profiling using selected reaction monitoring mass spectrometry. Sci Rep. 2015;5:13944.
Han X, Aslanian A, Yates JR III. Mass spectrometry for proteomics. Curr Opin Chem Biol. 2008;12(5):483–90.
Vandermarliere E, Mueller M, Martens L. Getting intimate with trypsin, the leading protease in proteomics. Mass Spectrom Rev. 2013;32(6):453–65.
Craig R, Cortens J, Fenyo D, Beavis RC. Using annotated peptide mass spectrum libraries for protein identification. J Proteome Res. 2006;5(8):1843–9.
Craig R, Cortens JP, Beavis RC. Open source system for analyzing, validating, and storing protein identification data. J Proteome Res. 2004;3(6):1234–42.
Kusebauch U, Deutsch EW, Campbell DS, Sun Z, Farrah T, Moritz RL. Using PeptideAtlas, SRMAtlas, and PASSEL: comprehensive resources for discovery and targeted proteomics. Curr Protoc Bioinformatics. 2014;46:13.25. 1–8.
Anderson L, Hunter CL. Quantitative mass spectrometric multiple reaction monitoring assays for major plasma proteins. Mol Cell Proteomics. 2006;5(4):573–88.
MacLean B, Tomazela DM, Shulman N, Chambers M, Finney GL, Frewen B, et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics. 2010;26(7):966–8.
Liu H, Sadygov RG, Yates JR. A model for random sampling and estimation of relative protein abundance in shotgun proteomics. Anal Chem. 2004;76(14):4193–201.
Fusaro VA, Mani D, Mesirov JP, Carr SA. Prediction of high-responding peptides for targeted protein assays by mass spectrometry. Nat Biotechnol. 2009;27(2):190.
Gallien S, Peterman S, Kiyonami R, Souady J, Duriez E, Schoen A, et al. Highly multiplexed targeted proteomics using precise control of peptide retention time. Proteomics. 2012;12(8):1122–33.
Escher C, Reiter L, MacLean B, Ossola R, Herzog F, Chilton J, et al. Using iRT, a normalized retention time for more targeted measurement of peptides. Proteomics. 2012;12(8):1111–21.
Krokhin OV, Craig R, Spicer V, Ens W, Standing KG, Beavis RC, et al. An improved model for prediction of retention times of tryptic peptides in ion pair reversed-phase HPLC its application to protein peptide mapping by off-line HPLC-MALDI MS. Mol Cell Proteomics. 2004;3(9):908–19.
Krokhin OV, Ying S, Cortens JP, Ghosh D, Spicer V, Ens W, et al. Use of peptide retention time prediction for protein identification by off-line reversed-phase HPLC− MALDI MS/MS. Anal Chem. 2006;78(17):6265–9.
Calculator S-SR. Algorithm for peptide retention prediction in ion-pair RP-HPLC: application to 300-and 100-Å pore size C18 sorbents Krokhin, Oleg V. Anal Chem. 2006;78(22):7785–95.
Chen C-Y, Nace GW, Solow B, Fratamico P. Complete nucleotide sequences of 84.5-and 3.2-kb plasmids in the multi-antibiotic resistant Salmonella enterica serovar typhimurium U302 strain G8430. Plasmid. 2007;57(1):29–43.
Perez JJ, Chen C-Y. Detection of acetyltransferase modification of kanamycin, anaminoglycoside antibiotic, in bacteria using ultrahigh-performance liquid chromatography tandem mass spectrometry. Rapid Commun Mass Spectrom. 2018;32:1549–1556. https://doi.org/10.1002/rcm.8160.
Perez JJ, Chen C-Y. Rapid detection and quantification of aminoglycoside phosphorylationproducts using direct-infusion high-resolution and ultra-high-performance liquid chromatography/mass spectrometry. Rapid Commun Mass Spectrom. 2018;32:1822–1828. https://doi.org/10.1002/rcm.8241.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Published in the topical collection Young Investigators in (Bio-)Analytical Chemistry with guest editors Erin Baker, Kerstin Leopold, Francesco Ricci, and Wei Wang.
Electronic supplementary material
ESM 1
(PDF 698 kb)
Rights and permissions
About this article
Cite this article
Perez, J.J., Chen, CY. Implementation of normalized retention time (iRT) for bottom-up proteomic analysis of the aminoglycoside phosphotransferase enzyme facilitating method distribution. Anal Bioanal Chem 411, 4701–4708 (2019). https://doi.org/10.1007/s00216-018-1377-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-018-1377-z