Abstract
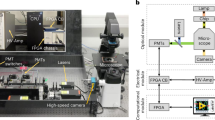
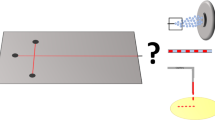
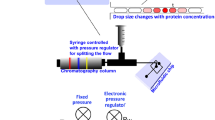
A new approach is presented for cell lysate identification which uses SERS-active silver nanoparticles and a droplet-based microfluidic chip. Eighty-nanoliter droplets are generated by injecting silver nanoparticles, KCl as aggregation agent, and cell lysate containing cell constituents, such as nucleic acids, carbohydrates, metabolites, and proteins into a continuous flow of mineral oil. This platform enables accurate mixing of small volumes inside the meandering channels of the quartz chip and allows acquisition of thousands of SERS spectra with 785 nm excitation at an integration time of 1 s. Preparation of three batches of three leukemia cell lines demonstrated the experimental reproducibility. The main advantage of a high number of reproducible spectra is to apply statistics for large sample populations with robust classification results. A support vector machine with leave-one-batch-out cross-validation classified SERS spectra with sensitivities, specificities, and accuracies better than 99% to differentiate Jurkat, THP-1, and MONO-MAC-6 leukemia cell lysates. This approach is compared with previous published reports about Raman spectroscopy for leukemia detection, and an outlook is given for transfer to single cells.

A quartz chip was designed for SERS at 785 nm excitation. Principal component analysis of SERS spectra clearly separates cell lysates using variations in band intensity ratios.





Similar content being viewed by others
References
Kong K, Kendall C, Stone N, Notingher I. Raman spectroscopy for medical diagnostics—from in-vitro biofluid assays to in-vivo cancer detection. Adv Drug Deliver Rev. 2015;89:121–34.
Krafft C, Schie IW, Meyer T, Schmitt M, Popp J. Developments in spontaneous and coherent Raman scattering microscopic imaging for biomedical applications. Chem Soc Rev. 2016;45(7):1819–49.
Krafft C, Schmitt M, Schie IW, Cialla-May D, Matthaeus C, Bocklitz T, et al. Label-free molecular imaging of biological cells and tissues by linear and non-linear Raman spectroscopic approaches. Angew Chem Int Ed. 2017;56(16):4392–430. https://doi.org/10.1002/anie.201607604.
Chan JW, Taylor DS, Lane SM, Zwerdling T, Tuscano J, Huser T. Nondestructive identification of individual leukemia cells by laser trapping Raman spectroscopy. Anal Chem. 2008;80(6):2180–7.
Lau AY, Lee LP, Chan JW. An integrated optofluidic platform for Raman-activated cell sorting. Lab Chip. 2008;8(7):1116–20.
Chan JW, Taylor DS, Thompson DL. The effect of cell fixation on the discrimination of normal and leukemia cells with laser tweezers Raman spectroscopy. Biopolymers. 2009;91(2):132–9.
Vanna R, Ronchi P, Lenferink A, Tresoldi C, Morasso C, Mehn D, et al. Label-free imaging and identification of typical cells of acute myeloid leukaemia and myelodysplastic syndrome by Raman microspectroscopy. Analyst. 2015;140(4):1054–64.
Happillon T, Untereiner V, Beljebbar A, Gobinet C, Daliphard S, Cornillet-Lefebvre P, et al. Diagnosis approach of chronic lymphocytic leukemia on unstained blood smears using Raman microspectroscopy and supervised classification. Analyst. 2015;140(13):4465–72.
Managò S, Valente C, Mirabelli P, Circolo D, Basile F, Corda D, et al. A reliable Raman-spectroscopy-based approach for diagnosis, classification and follow-up of B-cell acute lymphoblastic leukemia. Sci Rep. 2016;6:24821.
Jun BH, Noh MS, Kim J, Kim G, Kang H, Kim MS, et al. Multifunctional silver-embedded magnetic nanoparticles as SERS nanoprobes and their applications. Small. 2010;6(1):119–25.
Nguyen CT, Nguyen JT, Rutledge S, Zhang J, Wang C, Walker GC. Detection of chronic lymphocytic leukemia cell surface markers using surface enhanced Raman scattering gold nanoparticles. Cancer Lett. 2010;292(1):91–7.
MacLaughlin CM, Mullaithilaga N, Yang G, Ip SY, Wang C, Walker GC. Surface-enhanced Raman scattering dye-labeled Au nanoparticles for triplexed detection of leukemia and lymphoma cells and SERS flow cytometry. Langmuir. 2013;29(6):1908–19.
Khetani A, Momenpour A, Alarcon EI, Anis H. Hollow core photonic crystal fiber for monitoring leukemia cells using surface enhanced Raman scattering (SERS). Biomed Opt Express. 2015;6(11):4599–609.
Perozziello G, Candeloro P, De Grazia A, Esposito F, Allione M, Coluccio ML, et al. Microfluidic device for continuous single cells analysis via Raman spectroscopy enhanced by integrated plasmonic nanodimers. Opt Express. 2016;24(2):A180–90.
Hassoun M, Schie IW, Tolstik T, Stanca S, Krafft C, Popp J. Surface enhanced Raman spectroscopy of cell lysates mixed with silver nanoparticles for tumor classification. Beilstein J Nanotechnol. 2017;8:1183–90.
Walter A, März A, Schumacher W, Rösch P, Popp J. Towards a fast, high specific and reliable discrimination of bacteria on strain level by means of SERS in a microfluidic device. Lab Chip. 2011;11(6):1013–21.
Dochow S, Beleites C, Henkel T, Mayer G, Albert J, Clement J, et al. Quartz microfluidic chip for tumour cell identification by Raman spectroscopy in combination with optical traps. Anal Bioanal Chem. 2013;405(8):2743–6. https://doi.org/10.1007/s00216-013-6726-3.
Strehle KR, Cialla D, Rösch P, Henkel T, Köhler M, Popp J. A reproducible surface-enhanced Raman spectroscopy approach. Online SERS measurements in a segmented microfluidic system. Anal Chem. 2007;79(4):1542–7.
Zhang X, Yin H, Cooper JM, Haswell SJ. Characterization of cellular chemical dynamics using combined microfluidic and Raman techniques. Anal Bioanal Chem. 2008;390(3):833–40.
Zhang JY, Do J, Premasiri WR, Ziegler LD, Klapperich CM. Rapid point-of-care concentration of bacteria in a disposable microfluidic device using meniscus dragging effect. Lab Chip. 2010;10(23):3265–70.
Leopold N, Lendl B. A new method for fast preparation of highly surface-enhanced Raman scattering (SERS) active silver colloids at room temperature by reduction of silver nitrate with hydroxylamine hydrochloride. J Phys Chem B. 2003;107(24):5723–7.
März A, Ackermann KR, Malsch D, Bocklitz T, Henkel T, Popp J. Towards a quantitative SERS approach–online monitoring of analytes in a microfluidic system with isotope-edited internal standards. J Biophotonics. 2009;2(4):232–42.
CoreTeam R. R: a language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2012.
Beleites C, Sergo V (2015) hyperSpec: a package to handle hyperspectral data sets in R. R package version 0.98-20150304. http://hyperSpec.r-forge.r-project.org
Liland KH, Mevik B-H. Baseline: baseline correction of spectra. R Package Version. 2015;1:2–1. https://CRAN.R-project.org/package=baseline
Moskovits M. Surface-enhanced spectroscopy. Rev Mod Phys. 1985;57(3):783.
Freitag I, Matthäus C, Csaki A, Clement JH, Cialla-May D, Weber K, et al. Differentiation of MCF-7 tumor cells from leukocytes and fibroblast cells using epithelial cell adhesion molecule targeted multicore surface-enhanced Raman spectroscopy labels. J Biomed Opt. 2015;20(5):055002-055002.
Nima ZA, Mahmood M, Xu Y, Mustafa T, Watanabe F, Nedosekin DA, et al. Circulating tumor cell identification by functionalized silver-gold nanorods with multicolor, super-enhanced SERS and photothermal resonances. Sci Rep. 2014;4:4752.
Chan JW, Taylor DS, Zwerdling T, Lane SM, Ihara K, Huser T. Micro-Raman spectroscopy detects individual neoplastic and normal hematopoietic cells. Biophys J. 2006;90(2):648–56.
Freitag I, Beleites C, Dochow S, Clement J, Krafft C, Popp J. Recognition of tumor cells by immune SERS markers in a microfluidic chip at continuous flow. Analyst. 2016;141:5986–9.
Acknowledgements
The PhD work of M. Hassoun is supported by a grant from the German Academic Exchange Service (DAAD). Funding of the research project "EXASENS" (13N13856) by the Federal Ministry of Education and Research, Germany, is acknowledged. The authors gratefully thank Dr. Izabella Jahn for introducing in microfluidic experiments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This article does not contain any studies with human or animal subjects.
Conflict of interest
The authors state that there are no conflicts of interest.
Additional information
Published in the topical collection celebrating ABCs 16th Anniversary.
Rights and permissions
About this article
Cite this article
Hassoun, M., Rüger, J., Kirchberger-Tolstik, T. et al. A droplet-based microfluidic chip as a platform for leukemia cell lysate identification using surface-enhanced Raman scattering. Anal Bioanal Chem 410, 999–1006 (2018). https://doi.org/10.1007/s00216-017-0609-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-017-0609-y