Abstract
The comprehensive analysis of untargeted metabolomics data acquired using LC-MS is still a major challenge. Different data analysis tools have been developed in recent years such as XCMS (various forms (X) of chromatography mass spectrometry) and multivariate curve resolution alternating least squares (MCR-ALS)-based strategies. In this work, metabolites extracted from rice tissues cultivated in an environmental test chamber were subjected to untargeted full-scan LC-MS analysis, and the obtained data sets were analyzed using XCMS and MCR-ALS. These approaches were compared in the investigation of the effects of copper and cadmium exposure on rice tissue (roots and aerial parts) samples. Both methods give, as a result of their application, the whole set of resolved elution and spectra profiles of the extracted metabolites in control and metal-treated samples, as well as the values of their corresponding chromatographic peak areas. The effects caused by the two considered metals on rice samples were assessed by further chemometric analysis and statistical evaluation of these peak area values. Results showed that there was a statistically significant interaction between the considered factors (type of metal of treatment and tissue). Also, the discrimination of the samples according to both factors was possible. A tentative identification of the most discriminant metabolites (biomarkers) was assessed. It is finally concluded that both XCMS- and MCR-ALS-based strategies provided similar results in all the considered cases despite the completely different approaches used by these two methods in the chromatographic peak resolution and detection strategies. Finally, advantages and disadvantages of using these two methods are discussed.

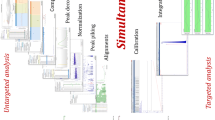
Summary of the workflow for untargeted metabolomics using the compared approaches





Similar content being viewed by others
References
Fukusaki E, Kobayashi A (2005) Plant metabolomics: potential for practical operation. J Biosci Bioeng 100(4):347–354
Xiao JF, Zhou B, Ressom HW (2012) Metabolite identification and quantitation in LC-MS/MS-based metabolomics. TrAC Trends Anal Chem 32:1–14
Roessner U, Luedemann A, Brust D, Fiehn O, Linke T, Willmitzer L, Fernie AR (2001) Metabolic profiling allows comprehensive phenotyping of genetically or environmentally modified plant systems. Plant Cell 13(1):11–29
Lommen A (2009) Metalign: Interface-driven, versatile metabolomics tool for hyphenated full-scan mass spectrometry data preprocessing. Anal Chem 81(8):3079–3086
Pluskal T, Castillo S, Villar-Briones A, Orešič M (2010) MZmine 2: Modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinformatics 11:395
Smith CA, Want EJ, O’Maille G, Abagyan R, Siuzdak G (2006) XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal Chem 78(3):779–787
Johnson CH, Ivanisevic J, Benton HP, Siuzdak G (2015) Bioinformatics: the next frontier of metabolomics. Anal Chem 87(1):147–156
Tautenhahn R, Patti GJ, Rinehart D, Siuzdak G (2012) XCMS online: A web-based platform to process untargeted metabolomic data. Anal Chem 84(11):5035–5039
Jaumot J, de Juan A, Tauler R (2015) MCR-ALS GUI 2.0: new features and applications. Chemometr Intell Lab 140:1–12
Farrés M, Piña B, Tauler R (2014) Chemometric evaluation of Saccharomyces cerevisiae metabolic profiles using LC-MS. Metabolomics 11:210–224
Cramer GR, Urano K, Delrot S, Pezzotti M, Shinozaki K (2011) Effects of abiotic stress on plants: a systems biology perspective. BMC Plant Biol 11:163
D’Alessandro A, Taamalli M, Gevi F, Timperio AM, Zolla L, Ghnaya T (2013) Cadmium stress responses in Brassica juncea: hints from proteomics and metabolomics. J Proteome Res 12(11):4979–4997
Villiers F, Ducruix C, Hugouvieux V, Jarno N, Ezan E, Garin J, Junot C, Bourguignon J (2011) Investigating the plant response to cadmium exposure by proteomic and metabolomic approaches. Proteomics 11(9):1650–1663
Järup L (2003) Hazards of heavy metal contamination. Brit Med Bull 68(1):167–182
Ahsan N, Nakamura T, Komatsu S (2012) Differential responses of microsomal proteins and metabolites in two contrasting cadmium (Cd)-accumulating soybean cultivars under Cd stress. Amino Acids 42(1):317–327
Antti H, Ebbels TMD, Keun HC, Bollard ME, Beckonert O, Lindon JC, Nicholson JK, Holmes E (2004) Statistical experimental design and partial least squares regression analysis of biofluid metabonomic NMR and clinical chemistry data for screening of adverse drug effects. Chemometr Intell Lab 73(1 SPEC. ISS):139–149
Johnson HE, Lloyd AJ, Mur LAJ, Smith AR, Causton DR (2007) The application of MANOVA to analyse Arabidopsis thaliana metabolomic data from factorially designed experiments. Metabolomics 3(4):517–530
Aina R, Labra M, Fumagalli P, Vannini C, Marsoni M, Cucchi U, Bracale M, Sgorbati S, Citterio S (2007) Thiol-peptide level and proteomic changes in response to cadmium toxicity in Oryza sativa L. roots. Environ Exp Bot 59(3):381–392
Roth U, Von Roepenack-Lahaye E, Clemens S (2006) Proteome changes in Arabidopsis thaliana roots upon exposure to Cd 2+. J Exp Bot 57(15):4003–4013
Huang SM, Toh W, Benke PI, Tan CS, Ong CN (2014). MetaboNexus: an interactive platform for integrated metabolomics analysis. Metabolomics 10:1084–1093
Prince JT, Marcotte EM (2006) Chromatographic alignment of ESI-LC-MS proteomics data sets by ordered bijective interpolated warping. Anal Chem 78(17):6140–6152
Tautenhahn R, Bottcher C, Neumann S (2008) Highly sensitive feature detection for high resolution LC/MS. BMC Bioinformatics 9:504
De Juan A, Jaumot J, Tauler R (2014) Multivariate Curve Resolution (MCR). Solving the mixture analysis problem. Anal Methods 6(14):4964–4976
de Juan A, Tauler R (2007) Factor analysis of hyphenated chromatographic data. Exploration, resolution and quantification of multicomponent systems. J Chromatogr A 1158(1–2):184–195
Ruckebusch C, Blanchet L (2013) Multivariate curve resolution: a review of advanced and tailored applications and challenges. Anal Chim Acta 765:28–36
Golub GH, Loan CFV (1996) Matrix computations, 3rd edn. Johns Hopkins University Press, Baltimore
Windig W, Guilment J (1991) Interactive self-modeling mixture analysis. Anal Chem 63(14):1425–1432
Tauler R, Maeder M, de Juan A (2010) Multiset data analysis: extended multivariate curve resolution. In: Brown SD, Tauler R, Walczak, B (ed) Comprehensive Chemometrics, vol 2. Elsevier B.V., Amsterdam, pp 473–505
Wold S, Esbensen K, Geladi P (1987) Principal component analysis. Chemometr Intell Lab 2(1–3):37–52
Barker M, Rayens W (2003) Partial least squares for discrimination. J Chemom 17(3):166–173
Jansen JJ, Hoefsloot HCJ, Van Der Greef J, Timmerman ME, Westerhuis JA, Smilde AK (2005) ASCA: analysis of multivariate data obtained from an experimental design. J Chemom 19(9):469–481
Kuhl C, Tautenhahn R, Böttcher C, Larson TR, Neumann S (2012) CAMERA: an integrated strategy for compound spectra extraction and annotation of liquid chromatography/mass spectrometry data sets. Anal Chem 84(1):283–289
Ortiz-Villanueva E, Jaumot J, Benavente F, Piña B, Sanz-Nebot V, Tauler R (2015) Combination of CE-MS and advanced chemometric methods for high-throughput metabolic profiling. Electrophoresis 36:2324–2335
Schmidtke LM, Blackman JW, Clark AC, Grant-Preece P (2013) Wine metabolomics: objective measures of sensory properties of semillon from GC-MS profiles. J Agric Food Chem 61(49):11957–11967
Horai H, Arita M, Kanaya S, Nihei Y, Ikeda T, Suwa K, Ojima Y, Tanaka K, Tanaka S, Aoshima K, Oda Y, Kakazu Y, Kusano M, Tohge T, Matsuda F, Sawada Y, Hirai MY, Nakanishi H, Ikeda K, Akimoto N, Maoka T, Takahashi H, Ara T, Sakurai N, Suzuki H, Shibata D, Neumann S, Iida T, Tanaka K, Funatsu K, Matsuura F, Soga T, Taguchi R, Saito K, Nishioka T (2010) MassBank: a public repository for sharing mass spectral data for life sciences. J Mass Spectrom 45(7):703–714
Tautenhahn R, Cho K, Uritboonthai W, Zhu Z, Patti GJ, Siuzdak G (2012) An accelerated workflow for untargeted metabolomics using the METLIN database. Nat Biotechnol 30(9):826–828
Wishart DS, Knox C, Guo AC, Eisner R, Young N, Gautam B, Hau DD, Psychogios N, Dong E, Bouatra S, Mandal R, Sinelnikov I, Xia J, Jia L, Cruz JA, Lim E, Sobsey CA, Shrivastava S, Huang P, Liu P, Fang L, Peng J, Fradette R, Cheng D, Tzur D, Clements M, Lewis A, de Souza A, Zuniga A, Dawe M, Xiong Y, Clive D, Greiner R, Nazyrova A, Shaykhutdinov R, Li L, Vogel HJ, Forsythei I (2009) HMDB: a knowledgebase for the human metabolome. Nucleic Acids Res 37(Suppl 1):D603–D610
Kanehisa M, Goto S, Sato Y, Furumichi M, Tanabe M (2012) KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res 40(D1):D109–D114
Sun X, Zhang J, Zhang H, Ni Y, Zhang Q, Chen J, Guan Y (2010) The responses of Arabidopsis thaliana to cadmium exposure explored via metabolite profiling. Chemosphere 78(7):840–845
Satterfield M, Brodbelt JS (2001) Structural characterization of flavonoid glycosides by collisionally activated dissociation of metal complexes. J Am Soc Mass Spectrom 12(5):537–549
Gyurcsik B, Nagy L (2000) Carbohydrates as ligands: coordination equilibria and structure of the metal complexes. Coordin Chem Rev 203(1):81–149
Acknowledgments
The research leading to these results has received funding from the European Research Council under the European Union’s Seventh Framework Programme (FP/2007-2013)/ERC Grant Agreement n. 320737. Also, recognition from the Catalan government (grant 2014 SGR 1106) is acknowledged. JJ acknowledges a CSIC JAE-Doc contract cofounded by the FSE, and AGR thanks CONICET for a fellowship.
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 382 kb)
Rights and permissions
About this article
Cite this article
Navarro-Reig, M., Jaumot, J., García-Reiriz, A. et al. Evaluation of changes induced in rice metabolome by Cd and Cu exposure using LC-MS with XCMS and MCR-ALS data analysis strategies. Anal Bioanal Chem 407, 8835–8847 (2015). https://doi.org/10.1007/s00216-015-9042-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-015-9042-2