Abstract
Key message
Ogura CMS fertility-restored materials, with 18 chromosomes, normal seed setting, stable fertility and closer genetic background to the parent Chinese kale, were successfully developed in B. oleracea via a triploid strategy for the first time.
Abstract
Ogura cytoplasmic male sterility (CMS) is the most widely used sterile type in seed production for commercial hybrids of Brassica oleracea vegetables. However, the natural Ogura CMS restorer line has not been found in B. oleracea crops. In this study, the triploid strategy was used with the aim to create euploid B. oleracea progenies with the Rfo gene. The allotriploid AAC hybrid YL2 was used as a male parent to backcross with Ogura CMS Chinese kale. After successive backcrosses, the BC2 Rfo-positive individual 16CMSF2-11 and its BC3 progenies, with 18 chromosomes, were developed, which were morphologically identical to the parent Chinese kale. Compared with F1 and BC1 plants, it showed stable fertility performance, and regular meiosis behavior and could produce seeds normally under natural pollination. The genomic composition analysis of Rfo-positive progenies by using molecular markers showed that more than 87% of the C-genome components of BC3 Rfo-progenies recovered to the parent Chinese kale, while most or all of the An-genome segments were lost in 16CMSF2-11 and its progenies. The results suggested that the genetic background of Rfo-positive individuals was closer to that of the parent Chinese kale along with backcrossing. Hereof, the Ogura CMS fertility-restored materials of Chinese kale were successfully created via triploid strategy for the first time, providing a bridge for utilizing the Ogura CMS B. oleracea germplasm in the future. Moreover, our study indicates that the triploid strategy is effective for transferring genes from B. napus into B. oleracea.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Brassica oleracea is a typical cross-pollination species, and its F1 hybrids exhibit strong heterosis, such as the high yield, better quality, increased uniformity, wide adaptability and resistance to biotic stresses in single cross-hybrids of cabbage, cauliflower and broccoli (Fang et al. 1983; Pearson 1983; Kucera et al. 2006; Singh et al. 2009, 2019; Dey et al. 2014). Thus, development of F1 hybrids is one of the most important objectives for B. oleracea vegetables. Due to the size and structure of B. oleracea flowers, it is not cost-effective to produce commercial hybrid seeds by manual emasculation and pollination. Instead, the male-sterile (MS) breeding system is a widely used, effective method in the production of commercial hybrids of B. oleracea vegetables (Fang et al. 2004; Yamagishi and Bhat 2014).
Genic male sterility (GMS) and cytoplasmic male sterility (CMS) are two main types of male sterility systems used in hybrid seed production (Kumar et al. 2000; Chen and Liu 2014). Most natural GMS mutants are difficult to utilize in the hybrid seed production due to their recessive traits (Nieuwhof 1961; Dickson 1970; Fang et al. 2001). In CMS system, the naturally occurring CMS resources are absent in B. oleracea, and most of the CMS types used in B. oleracea were transferred from radish, Brassica napus and other related cruciferous species through distant hybridization (Thompson 1972; Pearson 1972; Yarrow et al. 1990; Bannerot et al. 1974; Shu et al. 2016). Ogura CMS is a spontaneous mutational CMS type discovered in a natural population of radish, which is stable in sterility and easy to transfer (Ogura 1968). Owing to its excellent sterile characteristics, through efforts over several generations, Ogura CMS has been successfully transferred to B. oleracea by distant hybridization and protoplast fusion (Bannerot et al. 1974; Walters et al. 1992; Dey et al. 2011). Currently, Ogura CMS is the most widely used CMS type in hybrid seed production for B. oleracea vegetables (Wang et al. 2012; Dey et al. 2013; Singh et al. 2019).
However, due to all the Ogura CMS germplasm could not be self-pollinated, the excellent Ogura CMS germplasm cannot be utilized. For instance, clubroot disease is now becoming an increasingly severe disease, and resistant germplasm resources are absent in B. oleracea vegetables. A few clubroot-resistant varieties of cabbage are available on the market; for example, Tuoni and XG336 show high resistance to clubroot disease. However, they cannot be reutilized due to their Ogura CMS cytoplasm (Zhang et al. 2016; Ning et al. 2018). The development of B. oleracea Ogura CMS restorer lines is of great importance for the innovation and utilization of Ogura CMS germplasm. However, to date, a natural Ogura CMS restorer line has not been found in B. oleracea crops.
Previously, to obtain Ogura CMS fertility-restored lines in B. oleracea, the restorer-fertility gene (Rfo) for Ogura CMS was successfully transferred from rapeseed into Chinese kale by interspecific hybridization combined with embryo rescue. Fertility-restored interspecific hybrids (YL2, ACC) have been obtained (Yu et al. 2016). Due to the poor fertility of allotriploid ACC, colchicine doubling was performed to improve the fertility of the interspecific hybrids. The interspecific hexaploid hybrids (YL2-3, AACCCC) were produced and used as the male parent for backcrossing with Ogura CMS Chinese kale to develop BC generations (Yu et al. 2017). However, fertility-restored BC progenies with different genetic backgrounds still exhibited polyploid compositions, abnormal meiotic behaviors and showed a low seed setting rate (less than one seed per pod) under natural pollination (Yu et al. 2018).
In the present study, to speed up the process of creating Ogura CMS restorer materials in B. oleracea, the allotriploid AAC hybrid YL2 was used as the male parent to backcross with Ogura CMS Chinese kale, with the expectation of isolating euploid C gametes derived from the Rfo-positive progenies without chromosome doubling. With marker-assisted selection (MAS), morphology, fertility, cytological observations and genetic component analysis, Ogura CMS fertility-restored materials of Chinese kale (2n = 18) were developed.
Materials and methods
Plant materials
The F1 interspecific allotriploid fertility-restored individual (code: YL2, ACC), with an average pollen viability of 36.5%, was developed by distant hybridization between Chinese kale and rapeseed in a previous study (Yu et al. 2016). In this study, the F1 triploid individual YL2 was used as the male parent to produce BC1 plants. A high-generation Ogura CMS Chinese kale (code: 15Y102) was chosen as the recurrent female parent to develop BC generations. All of the plants used in the present study were grown in a greenhouse in autumn under normal management.
Development of BC progenies and screening of Rfo-positive individuals
The F1 fertility-restored individual YL2 was backcrossed with Chinese kale 15Y102 by repeated hand pollination during the bud period. Due to the interspecific reproductive barrier and low seed setting rate, embryo rescue was performed after hand pollination in the BC1 generations. Immature pods at 10 days after pollination were removed from plants and cultured in vitro, and then, mature embryos were excised from these pods after culturing for 10–15 days. The details of embryo rescue were described by Yu et al. (2016).
The Rfo-positive individuals in the BC1 and BC2 generations, showing relatively good fertility performance, were chosen as pollen donors to backcross with the parent 15Y102 for developing the next generations by hand pollination (without embryo rescue). The backcross procedure is shown in Fig. 1.
During the seed harvest period, the number of pollinated pods and seeds harvested from each cross-combination were investigated. The seed setting rate (seeds per pod) was calculated. After growing to seedlings, the individuals harboring the Rfo gene were screened for the Rfo-specific marker (BnRFO-AS2F/BnRFO-NEW-R) (Yu et al. 2016). All Rfo-positive individuals were propagated in plant regeneration medium (MS medium with 1 mg l−1 6-BA and 0.1 mg l−1 NAA) to ensure pollen supply. The rooted seedlings were transferred into soil pots and cultivated in a greenhouse.
Morphological traits and fertility performance investigation
The morphological characteristics, including the plant type, color of basal leaves, shape of basal leaves, margin of basal leaves, leaf surface of basal leaves, wax on leaf surfaces, flower color and inflorescence type of the Rfo-positive individuals, were investigated according to the standards described in “Descriptors and data standards for Chinese kale” (Sun and Li 2008).
At anthesis, the fertility performance of the Rfo-positive individuals was assessed by measuring pollen viability using the acetocarmine dyeing method. The pollen grains were collected from three newly opened flowers and stained with 1% acetocarmine. More than 300 pollen grains were observed in each replication. Pollen viability was observed once every 10 days. The percentage of viable pollen, which was stained deep pink and had a plump shape, was calculated from three replications.
Ploidy identification and cytological analysis
The ploidy of Rfo-positive progenies was identified through flow cytometry (FCM, BD FACSCalibur™, BD Biosciences, San Jose, CA, USA). The nuclear DNA content was estimated according to the method of Dolezel et al. (2007) with some modifications. The specific procedure was as follows: (1) a small amount (approximately 200 mg) of plant leaf tissue was placed in a petri dish; (2) 1–1.5 ml of ice-cold Galbraith’s buffer for nucleus isolation was added, and the tissue was cut immediately in the buffer with a sharp scalpel; (3) the homogenate was mixed and then filtered through a 37-mm nylon mesh into a 1.5-ml sample tube (approximately 0.5 ml of filtrate); (4) the supernatant was carefully removed, and PI stock solution (50 mg ml−1 with 50 mg ml−1 RNase) was added; (5) the sample was incubated on ice (15–30 min) with occasional shaking before FCM analysis. Then, the Chinese kale parental DNA content (2C) was measured as per the reference. The G1 peak was positioned on the abscissa (200 channels) by adjusting the gain settings of the instrument. The coefficient of variation (CV) of all samples was below 5%.
For meiotic analysis, young floral buds were collected and treated with 8-hydroxyquinoline for 3–4 h at room temperature. Then, the samples were rinsed with distilled water and fixed in Carnoy’s solution [alcohol:acetic acid, 3: 1 (vol.)] for 24 h at room temperature and stored in 70% ethanol at 4 °C (Li et al. 2014). Anthers were separated and hydrolyzed by an enzyme mix (0.2% pectinase and 0.2% cellulase) for 4–5 h at 37 °C and then squashed and stained with PI solution (50 μg ml−1). Chromosome pairing at diakinesis and chromosome segregation at anaphase were observed and analyzed for 50 pollen mother cells (PMCs) in each BC Rfo-positive plant.
Indel marker design for genomic component analysis and polymorphism detection
To identify the background markers between the parents, resequencing of the parents at 30 × coverage over the whole genome (Chinese kale 15Y102 and B. napus 15Y403) was performed. This work was completed at the Beijing Genomics Institute (BGI) (Shenzhen, China). The resequencing data of the two parents were mapped to the TO1000 reference genome of B. oleracea (Co genome, http://plants.ensembl.org/Brassica_oleracea) using BWA (http://biobwa.sourceforge.net/index.shtml, Li and Durbin 2009) and then sorted by SAMtools (http://samtools.sourceforge.net). The 3–8 bp insertion-deletion mutations (Indels) between the parents were called using the Genome Analysis Toolkit (GATK), with the filter criteria of a QUAL value larger than 30 and a minimum depth of 10. Then, 150-bp flanking sequences on both sides of these Indels were extracted and searched against the B. napus reference genome (http://www.genoscope.cns.fr/brassicanapus/). Only unique and perfectly matched sequences located on the Cn genome were retained to avoid multiple loci on the B. napus reference genome. Primers were designed along chromosomes with intervals of 20 kb and with amplicon lengths varying from 100 to 200 bp, primer lengths ranging from 18 to 23 bp, GC contents of 40–50% and Tm values of 52–56 °C.
Because the An genome was specific to B. napus, specific An-markers covering the whole An genome were developed as follows: First, the An-genome sequences acquired from the B. napus reference genome (http://www.genoscope.cns.fr/brassicanapus/) were segmented into 500-bp fragments and marked with location information; then, these 500-bp sequences were aligned to the Co reference genome (TO1000) using BWA-MEM. A length of 6,432,500 bp of the An-genome-specific sequences, which could not be aligned to the Co reference genome (TO1000) genome, was retained and used to design primers by Primer3. The design principles were the same as those mentioned above.
In this study, two or three pairs of primers were selected at each interval of 5 Mb in the whole Co and An reference genomes, respectively. Finally, a total of 300 pairs of Co-genome primers and 156 pairs of An-genome primers were selected to detect polymorphisms between the parents (Supplementary Table 1).
DNA extraction and genetic background marker detection
Genomic DNA was extracted from fresh leaves using a modified cetyltrimethylammonium bromide (CTAB) protocol (Murray and Thompson 1980). The concentration of DNA was estimated using a spectrophotometer (BioDrop, UK) and adjusted to 40–50 ng/μL.
Polymerase chain reaction (PCR) experiments were performed in 10 µL reaction mixture containing 2 µL DNA template, 1 µL 10 × PCR buffer (Mg2+ included), 0.8 µL dNTPs (2.5 mM each), 0.4 µL forward and reverse primers (10 µM), 0.1 µL Taq DNA polymerase (5 U/µL), and 5.3 µL double-distilled H2O. PCR was performed using the following program: 94 °C for 5 min; 35 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 45 s; and 72 °C for 10 min. The amplicons were separated by 8% (w/v) polyacrylamide gel electrophoresis (160 V for 1.5 h) and visualized with silver nitrate staining (Bassam et al. 1991).
Results
Development of BC progenies and Rfo-positive screening
Successive backcrossing and MAS of Rfo were performed in the BC1–BC3 generations (Fig. 1, Table 1). The seed set of Rfo-positive individuals was investigated and compared among BC1–BC3 Rfo-positive individuals. Most of the pollinated pods with pollen of YL2 failed to elongate and expand, and had no seeds inside (Fig. 2a, e, d). Only 14 seeds survived without embryo rescue, with an average of 0.004 seeds per pod (14/3173) (Table 1). When using embryo rescue, 342 mature embryos were excised from 234 pods (1.47 embryos per pod on average). Many embryos excised from pods were collapsed, malformed or shriveled. Only three seedlings were obtained (Table 1). Finally, a total of eight BC1 seedlings were obtained through hand pollination and embryo rescue. Upon screening with the Rfo-specific marker, only one BC1 individual (code: 15CMSF-Y1), harboring the Rfo gene, was identified (Table 1).
The seed setting performance of recurrent parent 15Y102 when using YL2 (F1), 15CMSF-Y1 (BC1), and 16CMSF2-11 as pollen donors, respectively. a–c Inflorescences morphology of 15Y102 in 30 DAPs (days after pollination) after pollinated with YL2 (a), 15CMSF-Y1 (b), and 16CMSF2-11 (c), respectively. d Comparison of the pod morphology of 15Y102 in 40 DAPs after pollinated with YL2, 15CMSF-Y1 and 16CMSF2-11 (from left to right). e–g Comparison of seed growth in pods of 15Y102 in 50 DAPs after pollinated with YL2 (e), 15CMSF-Y1 (f) and 16CMSF2-11 (g), respectively
Then the individual 15CMSF-Y1, as the male parent, was backcrossed with the Ogura CMS Chinese kale 15Y102 to produce the BC2 progenies. A total of 6094 flower buds were pollinated, and 217 seeds were harvested from 2022 pods (Table 1). Similar to YL2, most of the harvested pods pollinated with the pollen of 15CMSF-Y1 had no seeds (Fig. 2b, d, f). Compared with the F1 individual YL2, the average seed setting rate of 15CMSF-Y1 was 0.11 seeds per pod, showing significant (P < 0.01) improvement. These seeds were sown, and 167 of the seeds germinated and grew into seedlings. Upon screening with the Rfo-specific marker, two Rfo-positive individuals, named 16CMSF2-11 and 16CMSF2-58, were detected.
The individual 16CMSF2-11, showing a better fertility performance in the flowering stage, was selected as a pollen donor for development of the BC3 generation. Unlike the F1 individual (YL2) and BC1 individual (15CMSF-Y1), the pollinated pods of 16CMSF2-11 expanded obviously with 6–10 plump seeds per pod. Relatively, few seeds were aborted during pod development (Fig. 2c, d, g). The seed setting rate of 16CMSF2-11 exhibited a normal level (average of 5.7 seeds per pod), which was significantly higher than those of F1 (YL2) and BC1 (15CMSF-Y1) (P < 0.01), and had no significant difference from that of the maintainer line of 15Y102 (P < 0.05). In the BC3 generation, a total of 18,533 seeds were harvested, and 10,000 seeds were randomly selected to sow for MAS screening (Table 1). A total of 8736 BC3 progenies were grown. Upon screening with the Rfo-specific marker, eight out of the 8736 BC3 progenies were positive (code: 17CMSF1-1 ~ 17CMSF1-8).
Morphological characterization and fertility performance of Rfo-positive individuals
Morphological characteristics, including leaf, bud, inflorescence, and flower, of Rfo-positive BC progenies were investigated. Unlike the interspecific F1 hybrid YL2, which has an intermediate morphology between Chinese kale and rapeseed (Yu et al. 2016), the BC1 individual 15CMSF-Y1 was morphologically similar to the parent Chinese kale 15Y102 (Fig. 3a). For example, the plant type (semi-erect), plant height (52.45–64.32), leaf shape (elliptical), leaf color (gray green with wax deposition) and flower color (nearly white) were all similar to those of Chinese kale (Fig. 3a, d), but there remained some differences from Chinese kale, such as thinner plant type and abnormal inflorescence (Fig. 3a). Compared with the BC1 individual 15CMSF-Y1, the morphology-related traits, including the plant type, leaves shape and flower color (entirely white) of two BC2 and eight BC3 Rfo-positive individuals were identical to those of the parent Chinese kale (Fig. 3b, c). The inflorescences of 10 BC2 and BC3 Rfo-positive individuals grew normally and had no abnormal or dead buds during plant growth (Fig. 3). Moreover, the BC2 and BC3 Rfo-positive individuals grew more vigorously than the parent Chinese kale 15Y102, and their flowering stage occurred was 10 to 15 days earlier than that of 15Y102.
Morphological characterization (a–c) and fertility performance (d–k) of the BC1 hybrid 15CMSF-Y1, BC2 hybrid 16CMSF2-11 and BC3 hybrid 17CMSF1-1, respectively. Plant morphology of 15CMSF-Y1 (a), 16CMSF2-11 (b) and 17CMSF1-1 (c). d, e Pollen performance of 15CMSF-Y1 during different flowering periods; f–h Pollen performance of 16CMSF2-11 (f), 16CMSF2-58 (g) and 17CMSF1-1 (h); i–k Pollen viability of 15CMSF-Y1 (i), 16CMSF2-11 (j) and 17CMSF1-1 (k)
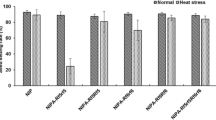
At anthesis, one BC1 (15CMSF-Y1), two BC2 (16CMSF2-11 and 16CMSF2-58) and eight BC3 Rfo-positive individuals were fertility-restored (Fig. 3d–k). For the BC1 Rfo-positive individual 15CMSF-Y1, the fertility performance was unstable during the whole flowering period (Fig. 3d, e). Its pollen viability varied throughout the flowering period, ranging from 15% to 65%, with the mean pollen viability of 45.5% (Fig. 3i). At the late flowering stage (after the 30th day of flowering), most flowers of 15CMSF-Y1 could not produce pollen grains (Fig. 3e). Compared with the BC1 individual 15CMSF-Y1, the BC2 individual 16CMSF2-11 showed a better and more stable fertility performance (Fig. 3f). The pollen viability of 16CMSF2-11 was always above 65%, with an average pollen viability of 76.2% (Fig. 3j), showing significant improvement compared with that of 15CMSF-Y1. Furthermore, all eight BC3 individuals showed stable fertility performance throughout the flowering period (Fig. 3h), and their mean pollen viability was all above 85% (Fig. 3k).
Ploidy identification and meiotic behaviors of Rfo-positive individuals
To determine the ploidy level, the DNA content of Rfo-positive individuals was estimated using flow cytometry. The G1 peak positions of 15CMSF-Y1 (BC1), 16CMSF2-11 (BC2) and 16CMSF2-58 (BC2) were located in 244 channels (Fig. 4a), 195 (Fig. 4b) and 205 channels, respectively, which suggested that their ploidy was close to that of the parent Chinese kale. The ploidy levels of all BC3 Rfo-positive plants, whose G1 peak positions located in 200 channels (Fig. 4c), were consistent with the ploidy of Chinese kale, suggesting that all eight BC3 Rfo-positive plants had returned to normal diploid level.
Ploidy identification (a–c) and cytogenetic characterization (d–l) of the BC1 hybrid 15CMSF-Y1, BC2 hybrid 16CMSF2-11 and BC3 hybrid 17CMSF1-1. Ploidy identification (relative nuclear DNA content) of 15CMSF-Y1 (a), 16CMSF2-11 (b) and 17CMSF1-1 (c) by FCM. d, e The PMC of 15CMSF-Y1 at diakinesis with abnormal chromosome configuration (d, e); multivalents (yellow arrows) and univalents (white arrows) were frequently observed. f One PMC of 15CMSF-Y1 with segregation of 9:13. g, h One PMC of 16CMSF2-11 at diakinesis with chromosome configuration 9II (g); multivalents (yellow arrows) and univalents were observed (h). i One PMC of 16CMSF2-11 with segregation of 9:9. j One PMC of 17CMSF1-1 at diakinesis with chromosome configuration 9II. k One PMC of 17CMSF1-1 equally segregated at anaphase I with chromosome segregation of 9: 9. l One PMC of 17CMSF1-1 equally segregated at anaphase II formed tetrad. Scale bars = 10 µm (color figure online)
The cytological analysis revealed that multivalents and univalents were observed in all PMCs of the BC1 Rfo-positive individual 15CMSF-Y1 at diakinesis, with the number of bivalents ranging from 6 to 10 (Fig. 4d, e). Moreover, all 50 PMCs produced unequal chromosome separation at anaphase I, with high frequency of the 10:12 and 9:13 chromosome distribution patterns (Fig. 4f). The chromosome number of most PMCs in 15CMSF-Y1 was 22 at anaphase I (Fig. 4f), yet some PMCs showed less than 22 chromosomes, which suggested that some chromosomes may be lost during the meiosis process. With regard to the two BC2 Rfo-positive individuals, the chromosome number of 16CMSF2-11 was 18 (Fig. 4i), while that of 16CMSF2-58 was 19. Abnormal chromosome behavior, including multivalents, univalents, and abnormal chromosome pairings could still be found in most PMCs of 16CMSF2-11 at diakinesis (Fig. 4h). However, the normal chromosome configuration of 9II was found in 6% of the PMCs of 16CMSF2-11 (3/50, Fig. 4g). In addition, 24% (12/50) of the PMCs showed a chromosome distribution pattern of 9:9 at anaphase I (Fig. 4i), indicating the possibility of producing euploid progenies with 18 chromosomes.
All eight BC3 individuals displayed a similar meiotic behavior at diakinesis, and the chromosomes pairing become more normal (Fig. 5j). All eight BC3 individuals had 18 chromosomes identical to 16CMSF2-11. The most frequently occurring configuration of 18 chromosomes, 9II, was observed in more than 55% of the analyzed PMCs (Fig. 5j), and the proportions were 64% (32/50), 70% (35/50), 72% (36/50), 56% (28/50), 84% (42/50), 60% (30/50), 60% (30/50) and 76% (38/50) for 17CMSF1-1 to 17CMSF1-8, respectively.
Analysis of the genomic composition of the BC1 hybrid 15CMSF-Y1, BC2 hybrids 16Q2-11, and BC3 hybrids 17CMSF1-1–17CMSF1-8 by the Co-genome primers. The marker name and marker locations are listed to the left of each chromosome. Red: loci of parent Chinese kale 15Y102 types. Yellow: loci of heterozygous types
Genomic composition analysis of Rfo-positive individuals
A total of 98 pairs of Co-genome polymorphic primers between parents were used to analyze the genomic components of the BC1–BC3 Rfo-positive individuals (Supplementary Fig. S1). In 15CMSF-Y1, 45 out of 98 pairs of Co-genome primers amplified identical single band patterns as those of the parent Chinese kale 15Y102 (indicated with red in Fig. 5), while the remaining 53 primers (indicated with yellow in Fig. 5) showed heterozygous bands. The results showed that 72.96% of the 15CMSF-Y1 genomic components of Co-genome had recovered to that of the parent 15Y102. For the two BC2 Rfo-positive individuals (16CMSF2-11 and 16CMSF2-58), 68 and 66 pairs of Co-genome primers amplified identical bands as the parent 15Y102, which suggested that 83.67% and 84.69% of the Co-genome components, respectively, had been recovered to those in the parent 15Y102 (Fig. 5). The genomic components of the Co genome in the eight BC3 Rfo-positive individuals were closer to those of the parent Chinese kale, and more than 87% (87.24–91.32%) of the Co-genome components were the same as those in the parent Chinese kale (Fig. 5). The individuals 17CMSF1-1 and 17CMSF1-5 contained the highest proportion of Co-genome components, while 17CMSF1-6 had the lowest recovery proportion. Genomic composition analysis of eight BC3 Rfo-positive individuals indicated that chromosome C09 had the highest proportion of heterozygosity compared with the other eight chromosomes (Fig. 5).
For the An genome, only 60 pairs of primers could amplify single clear bands in the parent rapeseed 15Y403 and F1 individual YL2 but not in Chinese kale 15Y102, and these primers were used to analyze the genomic component of the An-genome in the BC1–BC3 Rfo-positive individuals (Fig. 6). Among the 60 pairs of An-genome primers, 26 pairs were absent in the BC1 Rfo-positive individual 15CMSF-Y1 (indicated with blue in Fig. 6), which indicated that these loci were lost in 15CMSF-Y1. For the two BC2 Rfo-positive individuals, only 2 pairs and 5 pairs of primers were present in 16CMSF2-11 and 16CMSF2-58 (Fig. 6), respectively. In the BC3 generation, these two loci were lost in 17CMSF1-2, 17CMSF1-5 and 17CMSF1-6, which suggested that the An-genome was completely eliminated in these three BC3 Rfo-positive individuals (Fig. 6), whereas the other five BC3 Rfo-positive individuals retained a small number of An-genome fragments derived from the parent B. napus (Fig. 6).
Analysis of the genomic composition of the BC1 hybrid 15CMSF-Y1, BC2 hybrid 16Q2-11 and BC3 hybrids 17CMSF1-1–17CMSF1-8 by the An-genome specific primers. The marker name and marker locations are listed to the left of each chromosome. Red: presence of parent B. napus 15Y403 fragments. Blue: absence of parent B. napus 15Y403 fragments
Discussion
Ogura CMS is the most widely exploited cytoplasm in the F1 hybrid production of Brassica vegetables throughout the world (Sakai and Imamura 1990; Dey et al. 2011; Yamagishi and Bhat 2014). It plays an important role in exploiting Brassica vegetable heterosis and improving agricultural yield (Fang et al. 2004; Dey et al. 2017). Furthermore, the Ogura CMS system provided excellent protection for F1 commercial hybrids at the technical level because all of the offspring exhibited MS, especially in B. oleracea vegetables that lacked the natural Ogura CMS restorer line. However, on the other hand, the creation of restorer line might challenge the protection of varieties with this cytoplasm, because some breeders only used Ogura CMS to protect the varieties instead of the plant variety protection (PVP) system. The concept of essentially derived varieties (EDVs), developed in UPOV (1991), specified that the EDV may qualify for protection if authorization is obtained from the breeder of the initial variety (IV), aiming to provide better protection to the breeder of the IV. However, the concept is not equivalent to denying the rights of breeders of EDVs, and it encourages all forms of plant breeding, thereby also increasing the motivation for germplasm innovation (Jördens 2005). Taking knowledge from some crops with natural restoration materials, such as rice and peppers (Shifriss and Frankel 1971; Kumar et al. 2007; Chen and Liu 2014; Kim and Zhang 2018), breeders should rely mainly on PVP to protect their varieties. For the protected Ogura CMS varieties, we would like to encourage that the breeder to use the fertility-recovered Ogura CMS germplasm to develop IV, or EDVs with the authorization of the breeder of the IV.
In this work, the Ogura CMS fertility-restored materials in B. oleracea were successfully created for the first time. The Ogura CMS fertility-restored materials could provide a bridge for germplasm reutilization of Ogura CMS. For example, clubroot-resistant cabbage germplasms are very scarce in B. oleracea, and a few Ogura clubroot-resistant varieties of cabbage are available on the market (Zhang et al. 2016). Recently, we successfully transferred a widely used clubroot resistance gene from Ogura clubroot-resistant materials to our inbred line with a normal cytoplasm by bridge material 16CMSF2-11, which lays a solid foundation for improving clubroot resistance in varieties of B. oleracea (Ren et al. 2020). Thus, the development of the fertility-restored Ogura CMS material provides a bridge for the utilization of the Ogura CMS materials in B. oleracea.
Distant hybridization was widely used as an important tool for increasing genetic variation, gene introgression and cultivated crop improvement in plants (Kalloo 1992; Liu et al. 2014). Alien chromosome lines (substitution, addition and translocation), obtaining in the process of distant hybridization, play an important role in transferring a beneficial trait from wild plants into crops, especially in wheat (Qi et al. 2007; Yang et al. 2009; Jiang et al. 1994; Mei et al. 2015). Previously, chromosomal doubling was performed, and the amphidiploid individual (AACCCC) was obtained (Yu et al. 2016, 2017, 2018). Amphidiploids are usually used as a bridge for the development of alien gene introgression or alien chromosome lines (Liu et al. 2014). However, the genomic introgression of rapeseed was difficult to eliminate by continuous backcrossing, and the diploid progenies were difficult to obtain, possibly because of homoeologous chromosome pairing between B. napus and B. oleracea (Song et al. 1995; Xiong et al. 2011; Xiong & Pires 2011). As an alternative strategy, in our study, the Ogura CMS restorer gene (Rfo) was successfully introduced into B. oleracea from B. napus via an allotriploid strategy. The results indicated that pollen viability (45.5%) was very poor in the BC1 hybrids, which resulted in a low seed setting rate (11.1%), and very few BC2 seeds (217) were produced (Table 1). This result is also consistent with a previous study on interspecific crossing between B. oleracea and B. napus (Chiang et al. 1977; Ayotte et al. 1987; Quazi 1988; Inomata 2002; Ripley and Beversdorf 2003; Li et al. 2014). Due to reproductive isolation between parents, the cross-compatibility barrier, complex character separation and poor fertility of hybrid offspring are usually observed in the process of distant hybridization (Comai 2000; Kumar et al. 2011). Cytological analysis results also suggested that abnormal chromosomal behaviors occurred very frequently in the BC1 generations (Fig. 4). As backcrossing continued, there was a dramatic increase in fertility (76.2%) and seed set (6–10 seeds per pod) in the BC2 individual 16CMSF2-11 and its progenies (Fig. 3). Furthermore, their chromosome behaviors became more normal during meiosis (Fig. 4). Combined with the genomic composition results (Figs. 5, 6), we inferred that alien chromosome lines (substitution or translocation) containing the Rfo gene were successfully developed, and introgression of Cn components from B. napus to B. oleracea was shown to be available via the allotriploid strategy between the two genomes in our study. Thus, our study provides a reference strategy for transferring genes into B. oleracea from B. napus.
In the process of distant hybridization, it is necessary to confirm the genomic composition and chromosome recombination of the progenies. Cytological identification, in situ hybridization (ISH) and molecular markers are the most common methods used to analyze genomic composition and meiotic behavior in distant hybrids (Kerlan et al. 1993; Li et al. 1995; Snowdon et al. 1997; Snowdon 2007; Heneen et al. 2012; Wen et al. 2012; Younis et al. 2015; Yu et al. 2015). However, due to the small genome size and high homology between the A/C genome and A/C subgenome, cytological identification and ISH have many limitations in application (Xiong and Pires 2011). Molecular markers are also widely used to identify the genomic composition of Brassica interspecific hybrids (Navabi et al. 2011; Fredua-Agyeman et al. 2014). In this study, Indel markers, evenly distributed throughout the genome, were used to analyze the genomic composition of Rfo-positive individuals of the BC1–BC3 generations. With this approach, we found that except at Chr. C01, recombination occurred at the C chromosomes in 15CMSF-Y1, and the homozygous (Co/Co) blocks recovered nearly 50% of the C genome in the BC1 hybrid (Fig. 5). With successive backcrossing with Chinese kale, a large number of genomic components of the Co-genome had recovered to those in the parent 15Y102 (Fig. 5). Interestingly, the proportions of Co-genome blocks in BC2 (83.67% and 84.69%) and BC3 (87.24–91.32%) Rfo-positive individuals were lower than the expected value (87.5%, 93.75%). Two main possible reasons for this result can be proposed: (1) the introgression fragment linked with the Rfo gene, such as the genomic composition of chromosome C09, has a tendency to be heterozygous for the Cn/Co genome (Fig. 6), as observed in the process of developing a B. napus fertility-restored line (Delourme et al. 1998). (2) Conservation of more Cn genome regions from B. napus is necessary in Rfo-positive individuals for plant survival, similar to the study of Pele et al. (2017). Additionally, for the An genome, more than 50% of the loci in A chromosomes were lost in the BC1 hybrid 15CMSF-Y1 and most locus in some A chromosomes was completely eliminated in 16CMSF2-11 and its BC3 progenies (Fig. 6). Combined with the chromosome number of 15CMSF-Y1, we inferred that the alien chromosomes A03, A05 and A09 were additional chromosomes in the 15CMSF-Y1. Some chromosomes were partially eliminated in 15CMSF-Y1, most likely because of homoeologous exchanges between the A and C genomes (Ding et al. 2015; Yang et al. 2017; Pele et al. 2017; Li et al. 2018). Furthermore, the introgression of An fragments from An chromosomes was eliminated in BC3 hybrids, and even completely eliminated in some BC3 hybrids (Fig. 6). In summary, a small proportion of Cn genomic components from B. napus was present in Rfo-positive individuals, and the genomic composition of Rfo-positive individuals was complex, as shown by molecular markers, which probably explains why the transmission rate of Rfo was abnormal. Therefore, further backcross is needed to eliminate the B. napus chromosome fragments under the condition of maintaining the Rfo gene in future.
References
Ayotte R, Harney PM, Machado VS (1987) The transfer of triazine resistance from Brassica napus L. to B. oleracea L. I. Production of F1 hybrids through embryo rescue. Euphytica 36:615–624
Bannerot H, Boulidard L, Couderon Y, Temple J (1974) Transfer of cytoplasmic male sterility from Raphanus sativus to Brassica oleracea. In: Wills AB, North C (eds) Proc Eucarpia Meet Cruciferae. Scottish Hortic Res Inst, Invergavrie, pp 52–54
Bassam BJ, Caetano-Anollés G, Gresshoff PM (1991) Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal Biochem 196:80–83
Chen L, Liu YG (2014) Male sterility and fertility restoration in crops. Annu Rev Plant Biol 65:579–606
Chiang MS, Chiang BY, Grant WF (1977) Transfer of resistance to race 2 of plasmodiophora Brassica from Brassica napus to cabbage (B. oleracea var. capitata). I. Interspecific hybridization between B. napus and B. oleracea var. capitata. Euphytica 26:319–336
Comai L (2000) Genetic and epigenetic interactions in allopolyploid plants Plant Mol. Biol 43:387–399
Delourme R, Foisset N, Horvais R, Barret P, Champagne G, Cheung WY, Landry BS, Renard M (1998) Characterisation of the radish introgression carrying the Rfo restorer gene for the Ogu-INRA cytoplasmic male sterility in rapeseed (Brassica napus L.). Theor Appl Genet 97:129–134
Dey SS, Sharma SR, Bhatia R, Kumar PR, Parkash C (2011) Development and characterization of “Ogura” based improved CMS lines of cauliflower (Brassica oleracea var. botrytis L). Indian J Genet Plant Breed 71:37–42
Dey SS, Bhatia R, Sharma SR, Parkash C, Sureja AK (2013) Effects of chloroplast substituted Ogura male sterile cytoplasm on the performance of cauliflower (Brassica oleracea var. botrytis L.) F1 hybrids. Sci Hortic 157:45–51
Dey SS, Singh N, Bhatia R, Parkash C, Chandel C (2014) Genetic combining ability and heterosis for important vitamins and antioxidant pigments in cauliflower (Brassica oleracea var. botrytis L.). Euphytica 195:169–181
Dey SS, Bhatia R, Bhardwaj I, Mishra V, Sharma K, Parkash C, Kumar S, Sharma VK, Kumar R (2017) Molecular-agronomic characterization and genetic study reveals usefulness of refined Ogura cytoplasm based CMS lines in hybrid breeding of cauliflower (Brassica oleracea var. botrytis L.). Sci Hortic 224:27–36
Dickson MH (1970) A temperature sensitive male sterile gene in broccoli, Brassica oleracea L. var. italica. J Am Soc Hortic Sci 95:13–14
Ding Y, Mei J, Liu Y, Wang L, Li Y, Wan H, Li J, Qian W (2015) Transfer of Sclerotinia stem rot resistance from wild Brassica oleracea into B. rapa. Mol Breed 35:225
Dolezel J, Greilhuber J, Suda J (2007) Estimation of nuclear DNA content in plants using flow cytometry. Nat Protoc 2:2233–2244
Fang Z, Sun P, Liu Y (1983) Some problems of the utilization of heterosis in cabbage and the selection of self-incompatiblity. Sci Agric Sin 16:51–62
Fang Z, Sun P, Liu Y, Yang L, Wang X, Zhuang M (2001) Investigation of different types of male sterility and application of dominant male sterility in cabbage. China Veg 1:6–10
Fang Z, Liu Y, Yang L, Wang X, Zhuang M, Zhang Y, Sun P (2004) Breeding and seed production technique of dominant genic male sterile (DGMS) Line and cytoplasmic male sterile (CMS) line in cabbage. Sci Agric Sin 37:717–723
Fredua-Agyeman R, Coriton O, Huteau V, Parkin IAP, Chèvre AM, Rahman H (2014) Molecular cytogenetic identification of B genome chromosomes linked to blackleg disease resistance in Brassica napus × B. carinata interspecific hybrids. Theor Appl Genet 127:1305–1317
Heneen WK, Geleta M, Brismar K, Xiong ZY, Pires JC, Hasterok R, Stoute AI, Scott RJ, King GJ, Kurup S (2012) Seed colour loci, homoeology and linkage groups of the C genome chromosomes revealed in Brassica rapa–B. oleracea monosomic alien addition lines. Ann Bot 109:1227–1242
Inomata N (2002) A cytogenetic study of the progenies of hybrids between Brassica napus and Brassica oleracea, Brassica bourgeaui, Brassica cretica and Brassica montana. Plant Breed 121(2):174–176
Jiang J, Friebe B, Gill BS (1994) Recent advances in alien gene transfer in wheat. Euphytica 73:199–212
Jördens R (2005) 2005) Progress of plant variety protection based on the International convention for the protection of new varieties of plants (UPOV convention. World Patent Inf 27(3):232–243
Kalloo G (1992) Utilization of wild species. In: Kalloo G, Chowdhury JB (eds) Distant hybridization of crop plants. Springer, Berlin, pp 149–167
Kerlan MC, Chevre AM, Eber F (1993) Interspecific hybrids between a transgenic rapeseed (Brassica napus) and related species: cytogenetical characterization and detection of the transgene. Genome 36:1099–1106
Kim YJ, Zhang D (2018) Molecular control of male fertility for crop hybrid breeding. Trends Plant Sci 23(1):53–65
Kucera V, Chytilova V, Vyvadilova M, Klima M (2006) Hybrid breeding of cauliflower using self-incompatibility and cytoplasmic male sterility. Hort Sci 33:148–152
Kumar S, Banerjee MK, Kalloo G (2000) Male sterility: mechanisms and current status on identification, characterization and utilization in vegetables. Veg Sci 27(1):1–24
Kumar S, Singh V, Singh M, Rai S, Kumar S, Rai S, Rai M (2007) Genetics and distribution of fertility restoration associated RAPD markers in inbreds of pepper (Capsicum annuum L.). Sci Hortic 111:197–202
Kumar S, Imtiaz M, Gupta S, Pratap A (2011) Distant hybridization and alien gene introgression. In: Pratap A, Kumar J (eds) Biology and breeding of food legumes, pp 81–110
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760
Li Z, Liu HL, Luo P (1995) Production and cytogenetics of intergeneric hybrids between Brassica napus and Orychophragmus violaceus. Theor Appl Genet 91:131–136
Li Q, Zhou Q, Mei J, Zhang Y, Li J, Li Z, Ge X, Xiong Z, Huang Y, Qian W (2014) Improvement of Brassica napus via interspecific hybridization between B. napus and B. oleracea. Mol Breed 34:1955–1963
Li Q, Chen Y, Yue F, Qian W, Song H (2018) Microspore culture reveals high fitness of B. napus-like gametes in an interspecific hybrid between Brassica napus and B. oleracea. PLoS ONE 13(3):e0193548
Liu D, Zhang H, Zhang L, Yuan Z, Hao M, Zheng Y (2014) Distant hybridization: a tool for interspecific manipulation of chromosomes. In: Alien gene transfer in crop plants, Springer, New York, NY, vol 1, pp 25–42
Mei J, Liu Y, Wei D, Wittkop B, Ding Y et al (2015) Transfer of sclerotinia resistance from wild relative of Brassica oleracea into Brassica napus using a hexaploidy step. Theor Appl Genet 128:639–644
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4326
Navabi ZK, Stead KE, Pires JC, Xiong ZY, Sharpe AG, Parkin IAP, Rahman MH, Good AG (2011) Analysis of B-genome chromosome introgression in interspecific hybrids of Brassica napus × B. carinata. Genetics 187:659–673
Nieuwhof M (1961) Male sterility in some cole crops. Euphytica 10:351–356
Ning Y, Wang Y, Fang Z, Zhuang M, Zhang Y, Lv H, Liu Y, Li Z, Yang L (2018) Identification and characterization of resistance for Plasmodiophora brassicae race 4 in cabbage (Brassica oleracea var. capitata). Austral Plant Pathol 47(5):531–541
Ogura H (1968) Studies on the new male-sterility in Japanese radish, with special reference to the utilization of this sterility towards practical raising of hybrid seed. Mem Fac Agric Kagoshima Univ 6:39–78
Pearson OH (1972) Cytoplasmically inherited male sterility characters and flavor components from the species cross Brassica nigra (L.) Koch × B. oleracea L. Am Soc Hort Sci J 97:397–402
Pearson OH (1983) Heterosis in vegetable crops//Heterosis. Springer, Berlin, pp 138–188
Pele A, Trotoux G, Eber F, Lode M, Gilet M, Deniot G, Falentin C, Negre S, Morice J, Rousseau-Gueutin M, Chèvre AM (2017) The poor lonesome a subgenome of Brassica napus var. Darmor (AACC) may not survive without its mate. New Phytol 213:1886–1897
Qi L, Friebe B, Zhang P, Gill BS (2007) Homoeologous recombination, chromosome engineering and crop improvement. Chromosome Res 15:3–19
Quazi MH (1988) Interspecific hybrids between Brassica napus L. and B. oleracea L. developed by embryo culture. Theor Appl Genet 75:309–318
Ripley VL, Beversdorf WD (2003) Development of self-incompatible Brassica napus: (I) introgression of S-alleles from Brassica oleracea through interspecific hybridization. Plant Breed 122:1–5
Sakai T, Imamura J (1990) Intergeneric transfer of cytoplasmic male sterility between Raphanus sativus (cms line) and Brassica napus through cytoplast-protoplast fusion. Theor Appl Genet 80(3):421–427
Shifriss C, Frankel R (1971) New sources of cytoplasmic male sterility in cultivated peppers. J Heredity 62:254–256
Shu J, Liu Y, Li Z, Zhang LL, Fang ZY, Yang LM, Zhuang M, Zhang YY, Lv HH (2016) Detection of the diversity of cytoplasmic male sterility sources in broccoli (Brassica oleracea var. italica) using mitochondrial markers. Front Plant Sci 7:927
Singh BK, Sharma SR, Singh B (2009) Heterosis for mineral elements in single cross-hybrids of cabbage (Brassica oleracea var. capitata L.). Sci Hortic 122(1):32–36
Singh S, Dey S S, Bhatia R, Kumar R, Sharma K, Behera T K. (2019) Heterosis and combining ability in cytoplasmic male sterile and doubled haploid based Brassica oleracea progenies and prediction of heterosis using microsatellites. PloS ONE 14(8)
Snowdon RJ (2007) Cytogenetics and genome analysis in Brassica crops. Chromosome Res 15:85–95
Snowdon RJ, Kohler W, Friedt W, Kohler A (1997) Genomic in situ hybridization in Brassica amphidiploids and interspecific hybrids. Theor Appl Genet 95:1320–1324
Song K, Lu P, Tang K, Osborn TC (1995) Rapid genome change in synthetic polyploids of Brassica and its implications for polyploid evolution. Proc Natl Acad Sci USA 92:7719–7723
Sun L, Li X (2008) Descriptors and data standards for Chinese kale (Brassica alboglabra Bailey). China Agriculture Press, Beijing
Thompson KF (1972) Cytoplasmic male sterility in oilseed rape. Heredity 29:253–257
UPOV (1991) International convention for the protection of new varieties of plants. http://www.upov.int/en/publications/conventions/1991/content.htm International Union for the Protection of New Varieties of Plants, Geneva
Walters TW, Mutschler MA, Earle ED (1992) Protoplast fusion-derived Ogura male sterile cauliflower with cold tolerance. Plant Cell Rep 10:624–628
Wang QB, Zhang YY, Fang ZY, Liu YM, Yang LM, Zhuang M (2012) Chloroplast and mitochondrial SSR help to distinguish allo-cytoplasmic male sterile types in cabbage (Brassica oleracea L. var. capitata). Mol Breed 30:709–716
Wen J, Zhu LX, Qi LP, Ke HM, Yi B, Shen JX, Tu JX, Ma CZ, Fu TD (2012) Characterization of interploid hybrids from crosses between Brassica juncea and B. oleracea and the production of yellow-seeded B. napus. Theor Appl Genet 125:19–32
Xiong Z, Pires JC (2011) Karyotype and identification of all homoeologous chromosomes of allopolyploid Brassica napus and its diploid progenitors. Genetics 187:37–49
Xiong Z, Gaeta RT, Pires JC (2011) Homoeologous shuffling and chromosome compensation maintain genome balance in resynthesized allopolyploid Brassica napus. Proc Natl Acad Sci USA 108:7908–7913
Yamagishi H, Bhat SR (2014) Cytoplasmic male sterility in Brassicaceae crops. Breed Sci 64:38–47
Yang WY, Liu DC, Li J, Zhang LQ, Wei HT, Hu XR, Zheng YL, He ZH, Zou YC (2009) Synthetic hexaploid wheat and its utilization for wheat genetic improvement in China. J Genet Genom 36:539–546
Yang Y, Wei X, Shi G, Wei F, Braynen J, Zhang J, Tian B, Cao G, Zhang X (2017) Molecular and cytological analyses of A and C genomes at meiosis in synthetic allotriploid Brassica hybrids (ACC) between B. napus (AACC) and B. oleracea (CC). J Plant Biol 60:181–188
Yarrow SA, Burnett LA, Wildeman RP, Kemble RJ (1990) The transfer of ‘Polima’ cytoplasmic male sterility from oilseed rape (Brassica napus) to broccoli (B. oleracea) by protoplast fusion. Plant Cell Rep 9:185–188
Younis A, Ramzan F, Hwang YJ, Lim KB (2015) FISH and GISH: molecular cytogenetic tools and their applications in ornamental plants. Plant Cell Rep 34:1477–1488
Yu HL, Fang ZY, Liu YM, Yang LM, Zhuang M, Lv HH, Li ZS, Zhang YY (2015) Genetic background screen of inter-specific hybrids between SSR marker assisted Chinese kale and rapeseed. China Veg 8:14–21
Yu HL, Fang ZY, Liu YM, Yang LM, Zhuang M, Lv HH, Li ZS, Han FQ, Liu XP, Zhang YY (2016) Development of a novel allele-specific Rfo marker and creation of Ogura CMS fertility-restored interspecific hybrids in Brassica oleracea. Theor Appl Genet 129:1625–1637
Yu HL, Li ZY, Yang LM, Liu YM, Yang LM, Zhuang M, Zhang LG, Lv HH, Li ZS, Han FQ, Liu XP, Fang ZY, Zhang YY (2017) Morphological and molecular characterization of the second backcross progenies of Ogu-CMS Chinese kale and rapeseed. Euphytica 213:55
Yu HL, Li ZY, Yang LM, Liu YM, Zhuang M, Lv HH, Li ZS, Fang ZY, Zhang YY (2018) Development of fertility-restored BC3 progenies in Ogura CMS Chinese kale and analysis on gene transmission rate of Rfo and genetic background. Sci Agric Sin 51:1746–1757
Zhang XL, Liu YM, Fang ZY, Yang LM, Zhuang M, Zhang YY, Li ZS, Lv HH (2016) Identification of germplasm resistant to clubroot (Plasmodiophora brassicae Woronin) in Broccoli (Brassica oleracea L. var. italica Plenck) and its relatives. J Plant Genet Resour 17:1106–1115
Acknowledgements
This work was supported by grants from the National Key Research and Development Program of China (2016YFD0101702), the National Natural Science Foundation of China (31572141, 31872948). The work was performed in the Key Laboratory of Biology and Genetic Improvement of Horticultural Crops, Ministry of Agriculture, Beijing 100081, China.
Author information
Authors and Affiliations
Contributions
YH and ZY conceived and designed the research. YH, LZ and RW conducted experiments and wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Carlos F. Quiros.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
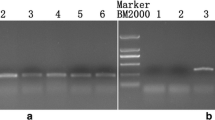
122_2020_3635_MOESM1_ESM.jpg
Supplementary Fig. S1. Screening results for the Co-genome primers (a) and An-genome primers (b) between male parent 15Y403 and female parent 15Y102. The black box indicates that those primers show polymorphismbetween the parents 15Y102 and 15Y403 or amplify single clear bands in 15Y403 (JPEG 48 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yu, Hl., Li, Zy., Ren, Wj. et al. Creation of fertility-restored materials for Ogura CMS in Brassica oleracea by introducing Rfo gene from Brassica napus via an allotriploid strategy. Theor Appl Genet 133, 2825–2837 (2020). https://doi.org/10.1007/s00122-020-03635-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-020-03635-8