Abstract
Foxp3+ regulatory T cells (Tregs) co-expressing the Th17-lineage specification factor RORγt represent a unique Treg subpopulation that has been reported to be induced upon response to gut microbiota within the intestinal immune system. Hence, RORγt+ Tregs are considered to solely consist of peripherally induced Foxp3+ Tregs (pTregs), and the possibility that also thymus-derived Treg (tTregs) can upregulate RORγt expression and contribute to the pool of RORγt+ Tregs was largely ignored. Here, we expand our knowledge on the origin of RORγt+ Tregs by demonstrating that also tTregs can attain RORγt expression. In transgenic Foxp3∆CNS1-Cre reporter mice, a substantial fraction of CNS1-independent Tregs, predominantly consisting of tTregs, was found to co-express RORγt. In addition, genuine tTregs isolated from thymi of Foxp3hCD2RAGGFP reporter mice initiated RORγt expression both in vitro and in vivo, particularly under inflammatory conditions. In conclusion, our data demonstrate that tTregs can upregulate RORγt expression under inflammatory conditions and that hence RORγt+ Tregs can be regarded as a heterogeneous population consisting of both pTregs and tTregs.
Key messages
-
RORγt cannot be considered as a marker for pTregs.
-
CNS1-independent tTregs within the colon display RORγt expression.
-
RORγt can be induced in genuine tTregs, particularly under inflammatory conditions.
-
RORγt+ Tregs are a heterogeneous population consisting of both pTregs and tTregs.




Similar content being viewed by others
Abbreviations
- BAC:
-
Bacterial artificial chromosome
- mLN:
-
Mesenteric lymph node
- Treg:
-
Regulatory T cell
- pTreg:
-
Peripherally induced Treg
- tTreg:
-
Thymus-derived Treg
References
Sakaguchi S, Yamaguchi T, Nomura T, Ono M (2008) Regulatory T cells and immune tolerance. Cell 133:775–787
Hsieh CS, Lee HM, Lio CW (2012) Selection of regulatory T cells in the thymus. Nat Rev Immunol 12:157–167
Tanoue T, Atarashi K, Honda K (2016) Development and maintenance of intestinal regulatory T cells. Nat Rev Immunol 16:295–309
Huehn J, Beyer M (2015) Epigenetic and transcriptional control of Foxp3 regulatory T cells. Semin Immunol 27:10–18
Lochner M, Peduto L, Cherrier M, Sawa S, Langa F, Varona R, Riethmacher D, Si-Tahar M, Di Santo JP, Eberl G (2008) In vivo equilibrium of proinflammatory IL-17+ and regulatory IL-10+ Foxp3+ RORgamma t+ T cells. J Exp Med 205:1381–1393
Zhou L, Lopes JE, Chong MM, Ivanov II, Min R, Victora GD, Shen Y, Du J, Rubtsov YP, Rudensky AY et al (2008) TGF-beta-induced Foxp3 inhibits T(H)17 cell differentiation by antagonizing RORgammat function. Nature 453:236–240
Sefik E, Geva-Zatorsky N, Oh S, Konnikova L, Zemmour D, McGuire AM, Burzyn D, Ortiz-Lopez A, Lobera M, Yang J et al (2015) MUCOSAL IMMUNOLOGY. Individual intestinal symbionts induce a distinct population of RORgamma+ regulatory T cells. Science 349:993–997
Ohnmacht C, Park JH, Cording S, Wing JB, Atarashi K, Obata Y, Gaboriau-Routhiau V, Marques R, Dulauroy S, Fedoseeva M et al (2015) MUCOSAL IMMUNOLOGY. The microbiota regulates type 2 immunity through RORgammat+ T cells. Science 349:989–993
Yang BH, Hagemann S, Mamareli P, Lauer U, Hoffmann U, Beckstette M, Fohse L, Prinz I, Pezoldt J, Suerbaum S et al (2016) Foxp3+ T cells expressing RORgammat represent a stable regulatory T-cell effector lineage with enhanced suppressive capacity during intestinal inflammation. Mucosal Immunol 9:444–457
Tartar DM, Van Morlan AM, Wan X, Guloglu FB, Jain R, Haymaker CL, Ellis JS, Hoeman CM, Cascio JA, Dhakal M et al (2010) FoxP3+RORgammat+ T helper intermediates display suppressive function against autoimmune diabetes. J Immunol 184:3377–3385
Kluger MA, Luig M, Wegscheid C, Goerke B, Paust HJ, Brix SR, Yan I, Mittrucker HW, Hagl B, Renner ED et al (2014) Stat3 programs Th17-specific regulatory T cells to control GN. J Am Soc Nephrol 25:1291–1302
Kluger MA, Meyer MC, Nosko A, Goerke B, Luig M, Wegscheid C, Tiegs G, Stahl RA, Panzer U, Steinmetz OM (2016) RORgammat+Foxp3+ cells are an independent bifunctional regulatory T cell lineage and mediate crescentic GN. J Am Soc Nephrol 27:454–465
Chellappa S, Hugenschmidt H, Hagness M, Line PD, Labori KJ, Wiedswang G, Tasken K, Aandahl EM (2016) Regulatory T cells that co-express RORgammat and FOXP3 are pro-inflammatory and immunosuppressive and expand in human pancreatic cancer. Oncoimmunology 5: e1102828. DOI
Kim B-S, Lu H, Ichiyama K, Chen X, Zhang Y-B, Mistry NA, Tanaka K, Lee Y-h, Nurieva R, Zhang L et al (2017) Generation of RORγt+ antigen-specific T regulatory 17 cells from Foxp3+ precursors in autoimmunity. Cell reports 21:195–207
Solomon BD, Hsieh CS (2016) Antigen-specific development of mucosal Foxp3+RORgammat+ T cells from regulatory T Cell precursors. J Immunol 197:3512–3519
Tone Y, Furuuchi K, Kojima Y, Tykocinski ML, Greene MI, Tone M (2008) Smad3 and NFAT cooperate to induce Foxp3 expression through its enhancer. Nat Immunol 9:194–202
Huehn J, Polansky JK, Hamann A (2009) Epigenetic control of FOXP3 expression: the key to a stable regulatory T-cell lineage? Nat Rev Immunol 9:83–89
Schlenner SM, Weigmann B, Ruan Q, Chen Y, von Boehmer H (2012) Smad3 binding to the foxp3 enhancer is dispensable for the development of regulatory T cells with the exception of the gut. J Exp Med 209:1529–1535
Zheng Y, Josefowicz S, Chaudhry A, Peng XP, Forbush K, Rudensky AY (2010) Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate. Nature 463:808–812
Kitagawa Y, Ohkura N, Kidani Y, Vandenbon A, Hirota K, Kawakami R, Yasuda K, Motooka D, Nakamura S, Kondo M et al (2017) Guidance of regulatory T cell development by Satb1-dependent super-enhancer establishment. Nat Immunol 18:173–183
Josefowicz SZ, Niec RE, Kim HY, Treuting P, Chinen T, Zheng Y, Umetsu DT, Rudensky AY (2012) Extrathymically generated regulatory T cells control mucosal TH2 inflammation. Nature 482:395–399
Zhang Z, Zhang W, Guo J, Gu Q, Zhu X, Zhou X (2017) Activation and functional specialization of regulatory T cells lead to the generation of Foxp3 instability. J Immunol 198:2612–2625
Thiault N, Darrigues J, Adoue V, Gros M, Binet B, Perals C, Leobon B, Fazilleau N, Joffre OP, Robey EA et al (2015) Peripheral regulatory T lymphocytes recirculating to the thymus suppress the development of their precursors. Nat Immunol 16:628–634
Ohkura N, Hamaguchi M, Morikawa H, Sugimura K, Tanaka A, Ito Y, Osaki M, Tanaka Y, Yamashita R, Nakano N et al (2012) T cell receptor stimulation-induced epigenetic changes and Foxp3 expression are independent and complementary events required for Treg cell development. Immunity 37:785–799
Toker A, Engelbert D, Garg G, Polansky JK, Floess S, Miyao T, Baron U, Duber S, Geffers R, Giehr P et al (2013) Active demethylation of the Foxp3 locus leads to the generation of stable regulatory T cells within the thymus. J Immunol 190:3180–3188
Wheaton JD, Yeh CH, Ciofani M (2017) Cutting edge: c-Maf Is required for regulatory T Cells to adopt RORgammat+ and follicular phenotypes. J Immunol. https://doi.org/10.4049/jimmunol.1701134
Samanta A, Li B, Song X, Bembas K, Zhang G, Katsumata M, Saouaf SJ, Wang Q, Hancock WW, Shen Y et al (2008) TGF-beta and IL-6 signals modulate chromatin binding and promoter occupancy by acetylated FOXP3. Proc Natl Acad Sci U S A 105:14023–14027
Lal G, Zhang N, van der Touw W, Ding Y, Ju W, Bottinger EP, Reid SP, Levy DE, Bromberg JS (2009) Epigenetic regulation of Foxp3 expression in regulatory T cells by DNA methylation. J Immunol 182:259–273
O’Connor RA, Floess S, Huehn J, Jones SA, Anderton SM (2012) Foxp3+ Treg cells in the inflamed CNS are insensitive to IL-6-driven IL-17 production. Eur J Immunol 42:1174–1179
Gao Y, Tang J, Chen W, Li Q, Nie J, Lin F, Wu Q, Chen Z, Gao Z, Fan H et al (2015) Inflammation negatively regulates FOXP3 and regulatory T-cell function via DBC1. Proc Natl Acad Sci U S A 112:E3246–E3254
Haribhai D, Lin W, Edwards B, Ziegelbauer J, Salzman NH, Carlson MR, Li SH, Simpson PM, Chatila TA, Williams CB (2009) A central role for induced regulatory T cells in tolerance induction in experimental colitis. J Immunol 182:3461–3468
Huang YJ, Haist V, Baumgartner W, Fohse L, Prinz I, Suerbaum S, Floess S, Huehn J (2014) Induced and thymus-derived Foxp3+ regulatory T cells share a common niche. Eur J Immunol 44:460–468
Acknowledgments
We thank all members of the animal facility, the flow cytometry facility, and the Department Experimental Immunology of the Helmholtz Centre for Infection Research (HZI), especially Dr. Lothar Gröbe, Yassin Elfaki, and Ahmed Elfiky for the technical assistance, and Dr. Shohei Hori (University of Tokyo, Japan) for providing Foxp3hCD2RAGGFP mice.
Funding
This work was supported by the German Research Foundation (SFB738-C7) and the Alexander von Humboldt Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All animal experiments were approved by the Lower Saxony Committee on the Ethics of Animal Experiments as well as the responsible state office (Lower Saxony State Office of Consumer Protection and Food Safety) under the permit numbers 33.9-42502-04-13-1170 and 33.9-42502-04-12/1012. Foxp3∆CNS1-CreRosa26YFP mice (C57BL/6 background) were bred in-house (Institute of Microbiology, Chinese Academy of Sciences). All experiments were conducted according to the guidelines of the Institute of Microbiology, Chinese Academy of Sciences Institutional Animal Care and Use Committee (permit no. PZIMCAS2011005).
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Supplementary Figure 1.
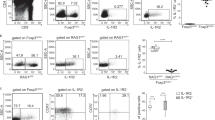
Genuine tTregs virtually lack RORγt expression and can be identified upon adoptive transfer into immunocompetent hosts. Genuine tTregs (CD25+Foxp3hCD2+RAGGFP+ CD4SP thymocytes) isolated from thymi of Foxp3hCD2RAGGFP reporter mice (CD45.1) were adoptively transferred into three-week-old immunocompetent recipient mice (CD45.2). PBS-injected mice served as controls. (a) Scatterplot summarizes RORγt expression among sorted CD25+Foxp3hCD2+RAGGFP+ CD4SP thymocytes before transfer. Data pooled from nine independent experiments are shown as mean ± SD. (b) Representative dotplot confirms lack of false-positive CD45.1+ cells among CD4+ cells from mLN of PBS-injected control mice four weeks post transfer. Number indicates frequency of CD45.1+ cells. (c) Scatterplot summarizes frequencies of CD45.1+ cells among CD4+ cells in mLN of recipient mice four weeks after adoptive transfer of genuine tTregs. (d) Scatterplot summarizes absolute number of CD45.1+CD4+ cells recovered from mLN of recipient mice four weeks after adoptive transfer of genuine tTregs. Each dot represents one mouse. Data pooled from three independent experiments are shown as mean ± SD (n = 6-9). (EPS 671 kb)
Supplementary Figure 2.
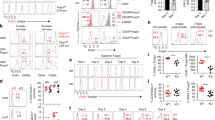
Impact of IL-6 on phenotype, survival and proliferation of in vitro cultured tTregs. Genuine tTregs (CD25+Foxp3hCD2+RAGGFP+ CD4SP thymocytes) isolated from thymi of Foxp3hCD2RAGGFP reporter mice were cultured for four days with or without IL-6, followed by flow cytometric analysis. (a) Scatterplots summarize mean fluorescence intensity (MFI) of RORγt (left) and Foxp3 expression (right) in indicated groups normalized to medium control. Each dot represents data from one independent experiment (n=5). (b) Representative dotplots determine LIVE/DEAD (LD) fixable Blue expression (left), and scatterplot (right) summarizes frequencies of LD+ cells among RORγt+ or RORγt- tTregs at the end of the culture. Each dot represents one independent experiment. Data are shown as mean ± SD. (c) Representative histograms depicts CellTrace Violet (CTV) signal of RORγt+ or RORγt- tTregs at the end of the culture (left), and scatterplot summarizes MFI of CTV signal among RORγt+ or RORγt- tTregs at the end of the culture. Unstimulated naïve T cells served as control. Each dot represents one independent experiment. Data are shown as mean ± SD. (EPS 735 kb)
Rights and permissions
About this article
Cite this article
Yang, J., Zou, M., Pezoldt, J. et al. Thymus-derived Foxp3+ regulatory T cells upregulate RORγt expression under inflammatory conditions. J Mol Med 96, 1387–1394 (2018). https://doi.org/10.1007/s00109-018-1706-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-018-1706-x