Abstract
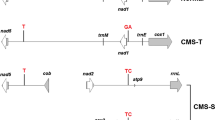
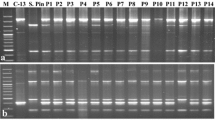
Restriction fragment length polymorphism (RFLP) of mitochondrial (mt) DNA provides a rapid and effective method to assess heterogeneity among male sterile cytoplasms. Six isonuclear A-lines (81 A1, with Tift 23A1, cytoplasm, ICMA 88001 (= 81Av) with Violaceum cytoplasm, 81A (=81A4) with monodli = violaceum cytoplasm, Pb 310A2 and Pb 311A2 with A2 cytoplasm from L 66A, and Pb 406A3 with A3 cytoplasm from L 67A), nine cytoplasmic male-sterility sources from Large-Seeded Genepool (LSGP 6, LSGP 14, LSGP 17, LSGP 22, LSGP 28, LSGP 36, LSGP 43, LSGP 55 and LSGP 66) and two each from Early Genepool (EGP 33 and EGP 15) and Population Varieties (PV 1 and PV 2) were characterized for variation in their mitochondrial genomes following Southern blot hybridizations using homologous (pearl millet 13.6 kb, 10.9 kb, 9.7 kb and 4.7 kb clones) and heterologous (maize atp6 and coxl clones) mitochondrial DNA (mtDNA) probes. Following cluster analysis based on similarity indices for the RFLP banding patterns observed, we identified seven cytoplasmic groups within LSGP. Two (LSGP 43 and LSGP 66) of these were quite distinct from each other as well as from other cytoplasms. This clearly indicates that besides serving as a source of diversity for agronomic and adaptation traits, broad-based gene pools can also provide diverse sources of cytoplasmic male sterility. These new CMS sources were also compared with standard CMS systems and cytoplasm-specific restriction fragments were identified.
Similar content being viewed by others
References
Ulistrop AJ, Ann Rev Phytopathology, 10 (1972) 37.
Pring OR & Lonsdale DM, Int Rev Cytol, 97 (1985) 1.
Ourbin RO & Uchytll TF, Biochem Genet, 15 (1977) 1143.
Burton GW & Athwal DS, Crop Sci, 7 (1967) 209.
Appadurai R, Raveendran TS & Nagarajan C, Indian J Agri Sci, 52 (1982) 832.
Aken’Ova ME, Euphytica, 34 (1985) 34.
Marchais L & Pernes J, Z. Pflanzenzucht, 95 (1985) 103.
Hanna WW, Crop Sci, 29 (1989) 1457.
Sujata V, Sivaramakrishnan S, Rai KN & Seetha K, Genome, 37 (1994) 482.
Rai KN & Hash CT, Crop Sci, 30 (1990) 889.
Virk DS & Brar JS, Theor Appl Genet, 87 (1993) 106.
Smith RL, Chowdhury MKU, Theor Appl Genet, 81 (1991) 793.
Smith RL, Chowdhury MKU & Pring DR, Plant Mol Biol, 9 (1987) 277.
Dewey RE, Levings CS III & Timothy DH, Plant Physiol, 79 (1980) 914.
Isaac PG, Jones VP & Leaver CJ, EMBOJ, 4 (1985) 1617.
Feinberg AP & Vogelstein B, Anal Biochem, 137 (1983) 266.
Nei M, Molecular evolutionary genetics. Columbia University Press, New York, USA (1987).
Rai KN, Plant Breeding, 114 (1995) 445.
Rai KN & Rao AS, International Crop Science Congress, New Delhi, India (1996) Abstract P5-0018.
Rajeshwari R, Sivaramakrishnan S, Smith RL & Subramanyam NC, Theor Appl Genet, 88 (1994) 441.
Rai KN, Virk DS & Harinarayana G, Cereals Program Annual Report. Cereals Program, International Crops Research Institute for the Semi-Arid Tropics, Patancheru, India (1991) P 79.
Chhabra AK, Ph.D Thesis, CCS Haryana Agricultural University, Hisar (1995).
Sivaramakrishnan S, Rajeshwari R, Subramanyam NC, Smith RL, Sujata V & Rai KN, Cereals Program Annual Report. Cereals Program, International Crops Research Institute for the Semi-Arid Tropics, Patancheru, India (1993) p 83.
Smith RL & Chowdhury MKU, Crop Sci, 29 (1989) 809.
Virk DS & Mangat BK, Millet News Lett, 6 (1987) 6.
Virk DS & Mangat BK, Millet News Lett, 7 (1988) 3.
Virk DS & Mangat BK, Millets News Lett, 8 (1989) 3.
Virk DS, Mangat BK & Gill KS, J Res Punjab Agric Uni, 27 (1990) 359.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Chhabra, A.K., Rai, K.N., Khairwal, I.S. et al. Mitochondrial DNA-RFLP Analysis Distinguishes New CMS Sources in Pearl Millet (Pennisetum glaucum (L.) R. Br.). J. Plant Biochem. Biotechnol. 7, 85–92 (1998). https://doi.org/10.1007/BF03263041
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF03263041