Abstract
Infection with hepatitis C virus (HCV) is a major cause of chronic liver disease. Hepatic fibrosis may develop in subjects with chronic HCV infection, culminating in cirrhosis and an increased risk of hepatocellular carcinoma. The rate of development of fibrosis varies substantially between individuals; while it is influenced by a number of demographic and environmental factors, these account for only a small proportion of the variability.
There are no clinical markers or tests that predict the rate of fibrosis progression in an individual subject. Thus, there has been increasing interest in the influence of host genetic factors on the rate of disease progression, and whether a genetic signature can be developed to reliably identify individuals at risk of severe disease. Numerous case-control, candidate gene, allele-association studies have examined the relationship between host single nucleotide polymorphisms or other genetic mutations and fibrosis in patients with chronic HCV infection. However, these studies have generally been irreproducible and disappointing. As seen with genetic studies for other diseases, small study cohorts and poor study design have contributed to limited meaningful findings. The successful determination of genetic signatures for fibrosis progression in chronic HCV will require multicenter collaborations using genome-wide association studies, with large, phenotypically well-defined sample sets. While these studies will require a significant financial commitment, a successful outcome offers the potential for personalized therapy and better patient management.
Similar content being viewed by others
Infection with hepatitis C virus (HCV) is a major cause of chronic liver disease, with an estimated 170 million people infected worldwide or ~3% of the world population.[1] Currently there is no prophylactic vaccine. HCV is transmitted mainly through contact with blood or blood products. Acute infection is usually asymptomatic or shows nonspecific mild symptoms, but most patients (70–80%) with HCV infection fail to clear the virus from their body and go on to develop chronic hepatitis.[2] The most significant consequence of chronic infection is the slow evolution of hepatic fibrosis over many years, culminating in cirrhosis and an increased risk of developing hepatocellular carcinoma.[2,3] The rate of development of fibrosis varies substantially among patients and is influenced by a number of demographic and environmental factors. However, these factors account for only a small proportion of the variability[4] and there are no clinical markers or tests that predict the rate of fibrosis progression in an individual subject. Thus, there has been increasing interest in the influence of host genetic factors on the rate of disease progression, and whether a genetic signature can be developed to reliably identify subjects at risk of severe disease.
1. The Virus and Antiviral Therapy
Discovered in 1989,[5] the HCV is a positive-strand RNA virus that belongs to the Flaviviridae family. The 9.6-kilobase HCV genome comprises untranslated regions that flank an uninterrupted open-reading frame encoding a single polyprotein of ~3000 amino acids that is processed into structural and nonstructural subunits by host and viral proteases.[6] HCV is highly heterogeneous and has been classified into six major genotypes (HCV 1–6) that can be further divided into numerous subtypes.[7] The six HCV genotypes show marked differences in geographic distribution. Genotypes 1–3 have a worldwide distribution, while genotype 4 is most common in the Middle East and North Africa; genotypes 5 and 6 are rarely seen outside of South Africa and Southeast Asia, respectively.[1,8]
Although the different genotypes are not considered to differ dramatically in their virulence or pathogenicity, viral genotype does influence responsiveness to antiviral therapy. The principal goal of therapy for chronic HCV infection is permanent viral eradication, termed a sustained virologic response (SVR). However, treatment of HCV is suboptimal and many patients do not tolerate or respond to therapy. Currently, the best available treatment is the combination of pegylated interferon (Peg-IFN) and ribavirin. Combination therapy is expensive and may have debilitating adverse effects, including fatigue, influenza-like illness, neuropsychiatric symptoms, gastrointestinal disturbances, thyroid dysfunction, dermatologic effects, and hematologic abnormalities.[9–11] Adverse effects impact compliance as they may require dose reductions in up to 40% of patients and drug discontinuation in up to 14% of patients.[10]
HCV genotype is the single most important predictor of treatment response. Peg-IFN/ribavirin therapy results in viral eradication in approximately 40–50% of individuals with genotype 1 or 4 infections and 70–80% of those with non-1/4 type infections.[12,13] Other factors that contribute to nonresponse to treatment include advanced fibrosis, high viral load, older age, obesity,[12–16] and possibly host genetic factors.[17]
2. Factors Associated with Increased Rate of Fibrosis Progression
Percutaneous liver biopsy is the gold standard for assessing disease activity and severity in patients with chronic HCV infection. Liver disease is characterized by hepatocellular injury and inflammation that is thought to lead to fibrosis, which is an increase in the extracellular matrix constituents such as collagen.[18] The staging of the liver disease by evaluation of fibrosis is a key component of liver biopsy assessment and the stages are commonly assigned a numerical value to indicate increasing severity (e.g. F0–F4). The stages are determined on the basis of the quantity and location of fibrosis. Chronic HCV infection leads to fibrous expansion of portal tracts that may develop short septa (F1). As the disease progresses, fibrous septa extend to form bridges between portal tracts (F2) and between portal tracts and central veins (F3). With further progression, parenchymal nodules surrounded by fibrosis indicate the development of cirrhosis (F4).[19] Steatosis (fatty liver) is a common histologic feature in chronic HCV infection with around 55% of biopsies showing some steatosis.[20]
Although chronic HCV infection is generally characterized by slowly progressive fibrosis, large cross-sectional studies have shown that the rate at which liver injury progresses varies substantially.[21–23] The rate of fibrosis progression can be estimated using a ratio of the stage of fibrosis to the duration of infection (fibrosis units per year). Approximately one-third of patients with chronic HCV infection may progress to cirrhosis within 20 years, one-third within 50 years and one-third may not progress at all.[21,22] A number of demographic and environmental factors have been identified that influence the rate of fibrosis development. An increased rate of fibrosis progression has been associated with older age at infection, male gender and excessive alcohol consumption,[21–23] co-infection with hepatitis B virus (HBV),[24,25] or immunosuppression related to organ transplantation,[26,27] or co-infection with human immunodeficiency virus (HIV).[26,28,29] Inflammation on biopsy contributes to fibrosis progression.[20] It is also now recognized that obesity-related steatosis[30,31] and insulin resistance[32,33] impact fibrosis progression in chronic HCV infection. While numerous studies have examined the effects of viral related factors (HCV genotype, viral load, viral quasispecies) on fibrosis progression, the weight of evidence indicates that these factors play no role in progression once chronic hepatitis has developed.[34–36]
3. Evidence for Host Genetic Factors Determining Fibrosis Progression
While it is now accepted that the rate of development of fibrosis in HCV infection is influenced by demographic and environmental factors, these account for only a small proportion of the variability. Wright and colleagues[4] examined the influence of age at infection, age at biopsy, duration of infection, gender, viral genotype, alcohol consumption, ethnicity, and mode of acquisition in HIV/HBV-negative HCV-infected patients. Using bivariate and multivariate analyses to construct a regression model to predict rate of fibrosis progression, the model accounted for only 30% of the variability in fibrosis rate. In an earlier study, demographic and environmental factors explained only 17% of fibrosis variability.[21] Thus, there has been increasing interest in the influence of host genetic factors on liver fibrosis.
Fibrogenesis is a complex process (figure 1), mediated by necroinflammation and activated hepatic stellate cells (HSCs) and driven by a number of concurrent pathways involving oxidative stress, inflammation, and steatosis.[36–38] More than 100 reports have examined the relationship between host single nucleotide polymorphisms (SNPs) or other genetic mutations and fibrosis in chronic HCV infection. These have mainly been case-control, candidate gene, allele-association studies where the candidate gene has been selected on the basis of its putative role in fibrogenic pathways and disease pathogenesis. These studies have been extensively reviewed.[39–42] Only genetic mutations that have been examined in at least two independent studies in which fibrosis was assessed by biopsy will be discussed further in this paper (see table I).
Fibrogenesis is a complex process, mediated by necroinflammation and activated hepatic stellate cells (HSC) and driven by a number of concurrent cell types and pathways involving cytokines, chemokines, hormones, oxidative stress, and steatosis.[36–38] Polymorphisms in genes within any of these pathways may potentially influence fibrogenesis. CTGF = connective tissue growth factor; EDN1 = endothelin-1 (also known as ET1); EGF = epidermal growth factor; FGF = fibroblast growth factor; HCV = hepatitis C virus; HFE = hemochromatosis gene; HSC = hepatic stellate cell; IGF = insulin-like growth factor; IL-6 = interleukin-6; MTTP = microsomal triglyceride transfer protein; PDGF = platelet-derived growth factor; RAS = renin-angiotensin system; ROS = reactive oxygen species; TGF-β1 = transforming growth factor-β1; TLR = toll-like receptor; TNFα = tumor necrosis factor-α VEGF = vascular endothelial growth factor.
3.1 Cytokine and Immune Response Genes
Several studies have focused on polymorphisms in cytokine and chemokine genes as these may influence inflammation and fibrosis.
Tumor necrosis factor-α (TNFα) has a major role as a mediator of the inflammatory response, and may promote apoptosis of inflammatory and fibrogenic cells but may also downregulate collagen synthesis.[75,76] Polymorphisms in the promoter of the TNF gene that influence the level of transcription have been associated with advanced liver disease,[43,44] but other studies have not confirmed these findings.[45–49] Interleukin 10 (IL-10) is an anti-inflammatory cytokine that inhibits activation or function of T cells, macrophages, antigen presenting cells, and HSC.[77,78] The promoter region of the IL10 gene contains three biallelic polymorphisms (−1082G>A, −819C>T, and −592C>A) that produce three different haplotypes with differing effects on IL-10 production: GCC (high), ACC (intermediate), and ATA (low).[79] The low IL-10-producing genotypes have been associated with more[50] or less[51] fibrosis in some studies, but not in others.[46,47,52,53]
Chemokine (C-C motif) receptor 5 (CCR5) is a receptor for proinflammatory chemokines that have key roles in host responses to viruses. A common 32-bp deletion mutation in the CCR5 gene (CCR5Δ32) causes truncation and loss of CCR5 receptors on lymphoid cells of homozygotes.[80] Interest in the Δ32 mutation was greatly increased when it was found to be of paramount importance for protection against HIV infection.[80,81] Studies of CCR5Δ32 in HCV have led to discrepant results. Hellier et al.[54] found that possession of the mutant allele was associated with more advanced fibrosis. However, other studies have found the opposite[55] or no association.[56–61]
The dominant fibrogenic cytokine in hepatic fibrosis is transforming growth factor-β1 (TGF-β1), which contributes to the activation of HSCs and their production of extracellular matrix proteins.[82] Its action is exerted through two complementary pathways, one that stimulates matrix accumulation and the other that reduces matrix degradation.[76] Several polymorphic sites have been described within the TGFB1 gene, including two in the promoter region (−800G>A and −509C>T), one at position +72 (C insertion) in a nontranslated region, and two in the signal sequence at codons 10 (Leu/Pro) and 25 (Arg/Pro).[83] Studies of TGFB1 polymorphisms in HCV have led to discrepant results. One study has reported that individuals with the high TGF-β1-producing genotype (homozygosity for the Arg25 allele) are more likely to have increased fibrosis.[47] Another study has suggested that the presence of proline at this position leads to more rapid fibrosis progression,[62] while a third study found no association between codon 25 genotype and fibrosis.[46]
Chemokine (C-C motif) ligand 2 (CCL2; also known as monocyte chemotactic protein type 1 [MCP1], acts on multiple leukocyte populations to promote recruitment.[84] CCL2 production by HSCs regulates leukocyte trafficking and may have a direct profibrogenic action via HSC chemotaxis.[85] A G>A SNP in the CCL2 gene at position −2518 results in increased production of CCL2.[86] Muhlbauer et al.[63] found that HCV-infected patients carrying the −2518G allele were more likely to have advanced fibrosis and severe inflammation on liver biopsy. Other studies have not been able to replicate this finding.[54,64,65]
3.2 Genes that Influence Steatosis, Oxidative Stress, or Fibrogenic Pathways
Recognition of the role of steatosis in increased fibrosis progression in patients with HCV has led to the investigation of genes that influence its development and severity.
Microsomal triglyceride transfer protein (MTTP) plays a role in the synthesis and secretion of very low-density lipoprotein (VLDL) in the liver. A SNP in the promoter of the MTTP gene (−493G>T) has been associated with lower levels of transcription, resulting in lower levels of MTTP and decreased secretion of triacylglycerol from the liver. Association studies for this polymorphism have produced conflicting results.[55,66]
Heavy iron overload may cause significant liver injury, including progressive fibrosis, cirrhosis, and hepatocellular carcinoma. This is well described in patients with hereditary forms of hemochromatosis, where the most common known genetic factor is homozygosity for the 282 Cys>Tyr (C282Y) variant in the HFE gene.[87,88] Heterozygosity for C282Y or other mutations in the HFE gene (e.g. His63Asp [H63D]) may result in relatively modest cellular iron accumulation[89] that may nevertheless increase the production of reactive oxygen species leading to activation of HSCs.[90] A number of studies have examined the association of mutations in the HFE gene with fibrosis in patients who are chronically infected with HCV, but are not homozygous for C282Y (i.e. nonhemochromatosis). Some studies,[55,67–70,91] but not others,[71–73] have reported an association between HFE mutations and advanced fibrosis. The heterogeneity of clinical penetrance of HFE mutations[88] may contribute to the discrepancies in these studies.
In cardiac and renal fibrosis, TGF-β1 production may be enhanced by angiotensin II, the principal effector molecule of the renin-angiotensin system (RAS). Angiotensin II may also augment the accumulation of extracellular matrix.[92] Functional polymorphisms of genes of the RAS have been described, including −6G>A in the promoter of the AGT gene, encoding the precursor peptide, angiotensinogen.[93] Powell et al.,[47] but not Forrest et al.,[74] found an association between the high-producing angiotensinogen avariant allele AGT-6A and increased fibrosis.
4. Evaluation of Study Design
4.1 Patient Categorization
The interpretation of studies that have examined the role of genetic factors in fibrosis progression in HCV is confounded by the method used to categorize subjects as having slow or rapid disease progression. Most studies have been cross-sectional in design, and the associations were assessed using the stage of fibrosis from a single liver biopsy, without taking into account the duration of HCV infection.[17] Within a single center, the majority of patients with chronic HCV infection have minimal or mild fibrosis, but it may not be valid to assign them to a slow fibrosis progression category if they have a relatively short duration of disease. A stronger, more relevant approach is to use strict criteria to categorize subjects based on both the histologic stage of fibrosis and duration of infection. Richardson et al.[55] categorized patients as ‘slow’ progressors if they had no or minimal fibrosis ≥20 years after acquisition of HCV (figure 2). Patients with rapidly progressive disease had stage 3 or 4 fibrosis on liver biopsy or stage 2 fibrosis ≤10 years after acquisition of HCV. Although this limited the number of subjects in the analysis (149 from an initial cohort of 326 patients), it provided a more robust assessment of identifying high- and low-risk patients based on the rate of disease progression.
Strategy for selection of a training cohort from patients (n = 326) with chronic hepatitis C virus (HCV) infection. Patients with slowly progressive disease had no or minimal fibrosis (stage 0 or 1) on liver biopsy ≥20 years after acquisition of HCV. Patients with rapidly progressive disease had stage 3 or 4 fibrosis on liver biopsy or stage 2 fibrosis ≤10 years after acquisition of HCV (reproduced from Richardson et al.,[55] with permission).
4.2 Multiple Gene Testing
For complex diseases, genetic associations are usually of small magnitude with odds ratios of 1.1–1.5, and any single polymorphism accounts for only 1–8% of the overall disease risk in the population.[94] Although the risk attributed to an individual polymorphism is very small, the additive effect of several genetic variants from different loci may account for a greater proportion of the disease risk.[94] However, few studies in chronic HCV infection have examined the combined effect of multiple predisposing alleles.
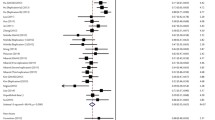
Richardson et al.[55] analyzed associations between polymorphisms in six genes and more rapidly progressing fibrosis. Individual adjusted odds ratios ranged from 2.1 to 4.5. However, for possession of ≥3, ≥4, or ≥5 pro-fibrotic alleles, the adjusted odds ratios were 9.1, 15.5, and 24.1, respectively. Using results from the logistic regression analysis (that adjusted for potential confounding by gender, age at infection, age at biopsy, viral genotype, and BMI), a predictive equation and a corresponding receiver-operating characteristic (ROC) curve was constructed to assess the ability to predict fibrosis progression. The area under the ROC curve was 0.868 and the predictive equation correctly classified 80% of patients in a validation cohort.[55]
Agnostic genome-wide association studies (GWAS) represent a major advance in the study of complex diseases.[95] It is now possible, using high throughput DNA microarrays, to simultaneously examine variations among thousands of genes. SNP chips to examine 500 000 to 1 million SNPs are now routinely analyzed. Huang and colleagues[96] used a method similar to that described above for categorizing patients with chronic HCV infection. In conjunction with a GWAS of 24 823 gene-centric SNPs and sophisticated statistical analysis, a cirrhosis risk score was developed from a 7-gene signature. The cirrhosis risk score was a better predictor than clinical factors for differentiating high- versus low-risk for cirrhosis, although steatosis, BMI, and insulin resistance were not taken into account. There are advantages and disadvantages with using a gene-centric approach. While this methodology should be more complete with regard to the coverage of genes, many putative causal variants lie outside genes and are in relatively low linkage disequilibrium with genic-tag SNPs.[97]
It may be suggested simplistically that the identification of a gene with an odds ratio of less than 1.5 may not usefully contribute to a genetic test to identify patients at risk of advanced fibrosis. However, the identification of a number of such genes, with additive or synergistic effects, may account for up to 70% of the overall disease risk.[94] Indeed, in the studies by Richardson et al.[55] and Huang et al.[96] that used a combination of SNPs in just 6 or 7 genes, 80% of patients in validation cohorts were correctly classified.
While the functionality of many of the SNPs may be unclear, it has been proposed that data from functional studies cannot substitute for compelling statistical genetic evidence of the existence and location of a susceptibility locus.[95] At present, experimental systems may not be sophisticated enough to fully determine the function of the very many SNPs that have been identified. Indeed, a tagged SNP may just be linked to the true functional polymorphism.
5. Genetic Tests and the Potential for Personalized Therapy
Despite enormous effort, to date studies that have investigated the contribution of host genetic factors to fibrosis progression in chronic HCV have been irreproducible and disappointing. This outcome is not, however, restricted to studies involving HCV.[94,98] As seen with genetic studies for other diseases, small study cohorts and poor study design (including mis-phenotyping and heterogeneous study populations) have resulted in limited meaningful findings.[99]
Investigation of factors contributing to fibrosis progression in chronic HCV is particularly challenging for a number of reasons. The restricted ability to collect incident cases and natural history of a disease that spans >20 years limits prospective long-term longitudinal studies. Because of the nature of the mode of acquisition, for most patients it is not possible to obtain a precise duration of infection. The lack of accurate non-invasive tests to assess disease progression necessitates liver biopsy, which is not without risk. The factors that influence the progression of established cirrhosis are also unknown. Early cirrhosis progresses to advanced cirrhosis in a minority of patients[100] and there is a need to uncover the genetic determinants.
Because of the high cost (both financial and in terms of adverse effects) of antiviral therapy, it has been suggested that a genetic test predicting the likelihood of rapid fibrosis progression could be used to decide which patients should be selected to undergo treatment. Recent improvements in antiviral therapy have resulted in 80% of patients with HCV genotype 2 or 3, and 50% of patients with genotype 1 or 4, achieving a SVR.[12,13] With recent reports that obesity and insulin resistance impair response to antiviral therapy,[14,101,102] the possibility is raised that weight loss prior to treatment will further improve the SVR rate. Is there still a need for a genetic test to aid in selection of patients to treat? Until recently, histology-based strategies were in widespread use, with treatment being determined by the degree of fibrosis and inflammation on presenting liver biopsy. However, an increasing number of health systems no longer require a liver biopsy for access to treatment, and the decision to treat is a more consultative process between patient and physician that balances the prospects of enduring treatment morbidity in the hope of preventing advanced fibrosis. Knowledge of a patient’s risk of developing severe disease will remain an important adjunct to the decision-making process.
The primary goal of treatment for patients with HCV is the eradication of the virus. However, for those patients who are unable to undertake or do not respond to antiviral therapy, the aim is to limit fibrosis progression. There are currently no approved therapeutic options designed to delay or reverse the progression of fibrosis. Many of the genes identified through association studies have provided mechanistic clues that may lead to potential targets for antifibrotic drug development. For example, following the association of the RAS with increased hepatic fibrosis,[47] animal models,[103,104] and, more recently, preliminary human studies,[105,106] have demonstrated that RAS inhibitors can attenuate development of hepatic fibrosis. As newer treatment options become available, with attendant clinical trials to evaluate efficacy, it will be vital that equal numbers of patients with a given risk of fibrosis progression are distributed in both control and treatment arms.[107] Since demographic and environmental factors account for only a small proportion of the variability in fibrosis progres sion,[4] the development of a genetic profile or other test that accurately predicts risk will be central to the establishment of drug efficacy.
Will there be a ‘one size fits all’ genetic signature that can predict rapid fibrosis progression? Racial, ethnic, and geographic variations in polymorphism frequency and function are already well documented for a number of genes[108,109] and it is recognized that study populations for genetic studies should be ethnically homogenous[110] and without residual confounding from population stratification. While viral-related factors apparently do not contribute to the rate of disease progression,[34–36] it is not known whether the same predisposing host genetic factors contribute to rapid fibrosis progression for the different viral genotypes. For example, it is now generally accepted that steatosis increases the rate of fibrosis progression in patients with chronic HCV infection (as well as in a number of other liver diseases).[111] However, for patients infected with HCV genotype 3, steatosis is thought to principally be a viral effect, while obesity related factors are considered to be the predominant influence on development of steatosis for patients infected with genotype 1.[35,111]
Similarly, with such a dramatic difference between males and females in the rate of fibrosis progression (independent of alcohol consumption), is it possible that genetic predisposition in causal pathways may differ? It must also be established whether histologic[112] and demographic information can increase the accuracy of a genetic signature.
6. The Way Forward
Successful determination of genetic signatures for fibrosis progression in patients with chronic HCV infection will require multicenter collaborations using GWAS, with large, phenotypically well-defined sample sets. Large sample sets (n > 1000 for the smallest subgroup) are needed to detect even moderate effects in order to eliminate the possibility of false-positive associations.[99] The approach will also need to be multidisciplinary, as the skill sets required are now beyond the scope of an individual specialist physician, scientist, geneticist, or epidemiologist.
As GWAS technology advances, it is becoming more affordable, but currently remains extremely expensive, and it will be necessary to convince funding agencies that the significant financial commitment required will provide ‘value for money.’ In this regard it will be prudent to take advantage of experience gained in other disease settings when designing and analyzing these studies.[99]
Although the strategy shift from hypothesis-driven to hypothesis-generating research[113] may not appeal to all, the outcome offers the potential for personalized therapy and better patient management.
References
WHO. Hepatitis C [online]. Available from URL: http://www.who.int/csr/disease/hepatitis/whocdscsrlyo2003/en/index3.html#prevalence [Accessed 2008 Jul 4]
Marcellin P. Hepatitis C: the clinical spectrum of the disease. J Hepatol 1999; 31: 9–16
Kim WR. The burden of hepatitis C in the United States. Hepatology 2002 Nov; 36(5): S30–4
Wright M, Goldin R, Fabre A, et al. Measurement and determinants of the natural history of liver fibrosis in hepatitis C virus infection: a cross sectional and longitudinal study. Gut 2003 Apr; 52(4): 574–9
Choo QL, Kuo G, Weiner AJ, et al. Isolation of a cDNA clone derived from a blood-borne non-A, non-B viral-hepatitis genome. Science 1989 Apr; 244(4902): 359–62
Penin F, Dubuisson J, Rey FA, et al. Structural biology of hepatitis C virus. Hepatology 2004 Jan; 39(1): 5–19
Simmonds P, Bukh J, Combet C, et al. Consensus proposals for a unified system of nomenclature of hepatitis C virus genotypes. Hepatology 2005 Oct; 42(4): 962–73
Hughes CA, Shafran SD. Chronic hepatitis C virus management: 2000–2005 update. Ann Pharmacother 2006 Jan; 40(1): 74–82
Fried MW. Side effects of therapy of hepatitis C and their management. Hepatology 2002 Nov; 36(5): S237–44
Russo MW, Fried MW. Side effects of therapy for chronic hepatitis C. Gastroenterology 2003 May; 124(6): 1711–9
Tomer Y, Blackard JT, Akeno N. Interferon alpha treatment and thyroid dysfunction. Endocrinol Metab Clin North Am 2007 Dec; 36(4): 1051–61
Manns MP, McHutchison JG, Gordon SC, et al. Peginterferon alfa-2b plus ribavirin compared with interferon alfa-2b plus ribavirin for initial treatment of chronic hepatitis C: a randomised trial. Lancet 2001 Sep; 358(9286): 958–65
Fried MW, Shiffman ML, Reddy KR, et al. Peginterferon alfa-2a plus ribavirin for chronic hepatitis C virus infection. N Engl J Med 2002 Sep; 347(13): 975–82
Bressler BL, Guindi M, Tomlinson G, et al. High body mass index is an independent risk factor for nonresponse to antiviral treatment in chronic hepatitis C. Hepatology 2003 Sep; 38(3): 639–44
Herrine SK, Rossi S, Navarro VJ. Management of patients with chronic hepatitis C infection. Clin Exper Med 2006 Mar; 6(1): 20–6
Dienstag JL, McHutchison JG. American Gastroenterological Association technical review on the management of hepatitis C. Gastroenterology 2006 Jan; 130(1): 231–64
Yee LJ. Host genetic determinants in hepatitis C virus infection. Genes Immun 2004 Jun; 5(4): 237–45
Friedman SL. Liver fibrosis: from bench to bedside. J Hepatol 2003; 38Suppl. 1: S38–53
Goodman ZD. Grading and staging systems for inflammation and fibrosis in chronic liver diseases. J Hepatol 2007 Oct; 47(4): 598–607
Kleiner DE. The liver biopsy in chronic hepatitis C: a view from the other side of the microscope. Semin Liver Dis 2005 Feb; 25(1): 52–64
Poynard T, Bedossa P, Opolon P. Natural history of liver fibrosis progression in patients with chronic hepatitis C. Lancet 1997 Mar; 349(9055): 825–32
Poynard T, Ratziu V, Charlotte F, et al. Rates and risk factors of liver fibrosis progression in patients with chronic hepatitis C. J Hepatol 2001 May; 34(5): 730–9
Roudot-Thoraval F, Bastie A, Pawlotsky JM, et al. Epidemiological factors affecting the severity of hepatitis C virus-related liver disease: a French survey of 6,664 patients. Hepatology 1997 Aug; 26(2): 485–90
Pontisso P, Gerotto M, Benvegnu L, et al. Coinfection by hepatitis B virus and hepatitis C virus. Antiviral Ther 1998; 3: 137–42
Tsai JF, Jeng JE, Ho MS, et al. Independent and additive effect modification of hepatitis C and B viruses infection on the development of chronic hepatitis. J Hepatol 1996 Mar; 24(3): 271–6
Pol S, Fontaine H, Carnot F, et al. Predictive factors for development of cirrhosis in parenterally acquired chronic hepatitis C: a comparison between immunocompetent and immunocompromised patients. J Hepatol 1998 Jul; 29(1): 12–9
Berenguer M, Ferrell L, Watson J, et al. HCV-related fibrosis progression following liver transplantation: increase in recent years. J Hepatol 2000 Apr; 32(4): 673–84
Martin P, Dibisceglie AM, Kassianides C, et al. Rapidly progressive non-A, non-B hepatitis in patients with human immunodeficiency virus infection. Gastroenterology 1989 Dec; 97(6): 1559–61
Soto B, SanchezQuijano A, Rodrigo L, et al. Human immunodeficiency virus infection modifies the natural history of chronic parenterally-acquired hepatitis C with an unusually rapid progression to cirrhosis. J Hepatol 1997 Jan; 26(1): 1–5
Hourigan LF, Macdonald GA, Purdie D, et al. Fibrosis in chronic hepatitis C correlates significantly with body mass index and steatosis. Hepatology 1999 Apr; 29(4): 1215–9
Adinolfi LE, Gambardella M, Andreana A, et al. Steatosis accelerates the progression of liver damage of chronic hepatitis C patients and correlates with specific HCV genotype and visceral obesity. Hepatology 2001 Jun; 33(6): 1358–64
Hickman IJ, Powell EE, Prins JB, et al. In overweight patients with chronic hepatitis C, circulating insulin is associated with hepatic fibrosis: implications for therapy. J Hepatol 2003 Dec; 39(6): 1042–8
Fartoux L, Poujol-Robert A, Guechot J, et al. Insulin resistance is a cause of steatosis and fibrosis progression in chronic hepatitis C. Gut 2005 Jul; 54(7): 1003–8
Seeff LB. Natural history of chronic hepatitis C. Hepatology 2002 Nov; 36(5): S35–46
Massard J, Ratziu V, Thabut D, et al. Natural history and predictors of disease severity in chronic hepatitis C. J Hepatol 2006; 44: S19–24
Teixeira R, Marcos LA, Friedman SL. Immunopathogenesis of hepatitis C infection and hepatic fibrosis: new insights into antifibrotic therapy in chronic hepatitis C. Hepatol Res 2007 Aug; 37(8): 579–95
Bataller R, Brenner DA. Liver fibrosis. J Clin Invest 2005 Feb; 115(2): 209–18
Friedman SL. Mechanisms of hepatic fibrogenesis. Gastroenterology 2008 May; 134(6): 1655–69
Bataller R, North KE, Brenner DA. Genetic polymorphisms and the progression of liver fibrosis: a critical appraisal. Hepatology 2003 Mar; 37(3): 493–503
Feld JJ, Liang TJ. Hepatitis C: identifying patients with progressive liver injury. Hepatology 2006 Feb; 43(2): S194–206
Asselah T, Bieche I, Paradis V, et al. Genetics, genomics, and proteomics: implications for the diagnosis and the treatment of chronic hepatitis C. Semin Liver Dis 2007 Feb; 27(1): 13–27
Osterreicher CH, Stickel F, Brenner AA. Genomics of liver fibrosis and cirrhosis. Semin Liver Dis 2007 Feb; 27(1): 28–43
Yee LJ, Tang J, Herrera J, et al. Tumor necrosis factor gene polymorphisms in patients with cirrhosis from chronic hepatitis C virus infection. Genes Immun 2000 Aug; 1(6): 386–90
Valenti L, Pulixi E, Fracanzani AL, et al. TNF alpha genotype affects TNF alpha release, insulin sensitivity and the severity of liver disease in HCV chronic hepatitis. J Hepatol 2005 Dec; 43(6): 944–50
Bahr MJ, el Menuawy M, Boeker KHW, et al. Cytokine gene polymorphisms and the susceptibility to liver cirrhosis in patients with chronic hepatitis C. Liver Int 2003 Dec; 23(6): 420–5
Barrett S, Collins M, Kenny C, et al. Polymorphisms in tumour necrosis factor-alpha, transforming growth factor-beta, interleukin-10, interleukin-6, interferon-gamma, and outcome of hepatitis C virus infection. J Med Virol 2003 Oct; 71(2): 212–8
Powell EE, Edwards-Smith CJ, Hay JL, et al. Host genetic factors influence disease progression in chronic hepatitis C. Hepatology 2000 Apr; 31(4): 828–33
Rosen HR, McHutchison JG, Conrad AJ, et al. Tumor necrosis factor genetic polymorphisms and response to antiviral therapy in patients with chronic hepatitis C. Am J Gastroenterol 2002 Mar; 97(3): 714–20
Yu ML, Dai CY, Chiu CC, et al. Tumor necrosis factor alpha promoter polymorphisms at position −308 in Taiwanese chronic hepatitis C patients treated with interferon-alpha. Antiviral Res 2003 Jun; 59(1): 35–40
Paladino N, Fainboim H, Theiler G, et al. Gender susceptibility to chronic hepatitis C virus infection associated with interleukin 10 promoter polymorphism. J Virol 2006 Sep; 80(18): 9144–50
Abbas Z, Moatter T, Hussainy A, et al. Effect of cytokine gene polymorphism on histological activity index, viral load and response to treatment in patients with chronic hepatitis C genotype 3. World J Gastroenterol 2005; 11: 6656–61
Knapp S, Hennig BJW, Frodsham AJ, et al. Interleukin-10 promoter polymorphisms and the outcome of hepatitis C virus infection. Immunogenetics 2003 Sep; 55(6): 362–9
Abbott WGH, Rigopoulou E, Haigh P, et al. Single nucleotide polymorphisms in the interferon-gamma and interleukin-10 genes do not influence chronic hepatitis C severity or T-cell reactivity to hepatitis C virus. Liver Int 2004 Apr; 24(2): 90–7
Hellier S, Frodsham AJ, Hennig BJW, et al. Association of genetic variants of the chemokine receptor MRS and its ligands, RANTES and MCP-2, with outcome of HCV infection. Hepatology 2003 Dec; 38(6): 1468–76
Richardson MM, Powell EE, Barrie HD, et al. A combination of genetic polymorphisms increases the risk of progressive disease in chronic hepatitis C. J Med Genet 2005 Jul; 42(7): 6
Promrat K, McDermott DH, Gonzalez CM, et al. Associations of chemokine system polymorphisms with clinical outcomes and treatment responses of chronic hepatitis C. Gastroenterology 2003 Feb; 124(2): 352–60
Wasmuth HE, Werth A, Mueller T, et al. CC chemokine receptor 5 Delta 32 polymorphism in two independent cohorts of hepatitis C virus infected patients without hemophilia. J Mol Med 2004 Jan; 82(1): 64–9
Mascheretti S, Hinrichsen H, Ross S, et al. Genetic variants in the CCR gene cluster and spontaneous viral elimination in hepatitis C-infected patients. Clin Exper Immunol 2004 May; 136(2): 328–33
Ruiz-Ferrer M, Barroso N, Antinolo G, et al. Analysis of CCR5-Delta 32 and CCR2-V64I polymorphisms in a cohort of Spanish HCV patients using realtime polymerase chain reaction and fluorescence resonance energy transfer technologies. J Viral Hepat 2004 Jul; 11(4): 319–23
Goyal A, Suneetha PV, Kumar GT, et al. CCRS Delta 32 mutation does not influence the susceptibility to HCV infection, severity of liver disease and response to therapy in patients with chronic hepatitis C. World J Gastroenterol 2006 Aug; 12(29): 4721–6
Goulding C, Murphy A, MacDonald G, et al. The CCR5-D32 mutation: impact on disease outcome in individuals with hepatitis C infection from a single source. Gut 2005 Aug; 54(8): 1157–61
Tag CG, Mengsteab S, Hellerbrand C, et al. Analysis of the transforming growth factor-beta 1 (TGF-beta 1) codon 25 gene polymorphism by LightCycler-analysis in patients with chronic hepatitis C infection. Cytokine 2003 Dec; 24(5): 173–81
Muhlbauer M, Bosserhoff AK, Hartmann A, et al. A novel MCP-1 gene polymorphism is associated with hepatic MCP-1 expression and severity of HCV-related liver disease. Gastroenterology 2003 Oct; 125(4): 1085–93
Glas J, Torok HP, Tonenchi L, et al. The-2518 promoter polymorphism in the MCP-1 gene is not associated with liver cirrhosis in chronic hepatitis C virus infection. Gastroenterology 2004 Jun; 126(7): 1930–1
Bonkovsky HL, Salek J. No role of the-2518 promoter polymorphism of monocyte chemotactic protein-1 in chronic hepatitis C. Gastroenterology 2005 Oct; 129(4): 1361–2
Petit JM, Masson D, Minello A, et al. Lack of association between microsomal triglyceride transfer protein gene polymorphism and liver steatosis in HCV-infected patients. Mol Genet Metab 2006 Jun; 88(2): 196–8
Martinelli ALC, Franco RF, Villanova MG, et al. Are haemochromatosis mutations related to the severity of liver disease in hepatitis C virus infection? Acta Haematol 1999; 102(3): 152–6
Smith BC, Grove J, Guzail MA, et al. Heterozygosity for hereditary hemochromatosis is associated with more fibrosis in chronic hepatitis C. Hepatology 1998 Jun; 27(6): 1695–9
Erhardt A, Maschner-OIberg A, Mellenthin C, et al. HFE mutations and chronic hepatitis C: H63D and C282Y heterozygosity are independent risk factors for liver fibrosis and cirrhosis. J Hepatol 2003 Mar; 38(3): 335–42
Gehrke SG, Stremmel W, Mathes I, et al. Hemochromatosis and transferrin receptor gene polymorphisms in chronic hepatitis C: impact on iron status, liver injury and HCV genotype. J Mol Med 2003 Dec; 81(12): 780–7
Hezode C, Cazeneuve C, Coue O, et al. Liver iron accumulation in patients with chronic active hepatitis C: prevalence and role of hemochromatosis gene mutations and relationship with hepatic histological lesions. J Hepatol 1999 Dec; 31(6): 979–84
Thorburn D, Curry G, Spooner R, et al. The role of iron and haemochromatosis gene mutations in the progression of liver disease in chronic hepatitis C. Gut 2002; 50(2): 248–52
Lebray P, Zylberberg H, Hue S, et al. Influence of HFE gene polymorphism on the progression and treatment of chronic hepatitis C. J Viral Hepat 2004 Mar; 11(2): 175–82
Forrest EH, Thorburn D, Spence E, et al. Polymorphisms of the renin-angiotensin system and the severity of fibrosis in chronic hepatitis C virus infection. J Viral Hepat 2005 Sep; 12(5): 519–24
Baker SJ, Reddy EP. Modulation of life and death by the TNF receptor superfamily. Oncogene 1998 Dec; 17(25): 3261–70
Verrecchia F, Mauviel A. TGF-beta and TNF-alpha: antagonistic cytokines controlling type I collagen gene expression. Cell Signal 2004 Aug; 16(8): 873–80
Moore KW, Malefyt RD, Coffman RL, et al. Interleukin-10 and the interleukin-10 receptor. Ann Rev Immunol 2001; 19: 683–765
Wang SC, Ohata M, Schrum L, et al. Expression of interleukin-10 by in vitro and in vivo activated hepatic stellate cells. J Biol Chem 1998 Jan; 273(1): 302–8
Turner DM, Williams DM, Sankaran D, et al. An investigation of polymorphism in the interleukin-10 gene promoter. Eur J Immunogenet 1997 Feb; 24(1): 1–8
Liu R, Paxton WA, Choe S, et al. Homozygous defect in HIV-1 coreceptor accounts for resistance of some multiply-exposed individuals to HIV-1 infection. Cell 1996 Aug; 86(3): 367–77
Samson M, Libert F, Doranz BJ, et al. Resistance to HIV-1 infection in Caucasian individuals bearing mutant alleles of the CCR-5 chemokine receptor gene. Nature 1996; 382(6593): 722–5
Bedossa P, Paradis V. Transforming growth factor-beta (TBF-beta): a key role in liver fibrogenesis. J Hepatol 1995; 22: 37–42
Awad MR, El Gamel A, Hasleton P, et al. Genotypic variation in the transforming growth factor-betal gene: association with transforming growth factor-pi production, fibrotic lung disease, and graft fibrosis after lung transplantation. Transplantation 1998 Oct; 66(8): 1014–20
Baggiolini M, Dewald B, Moser B. Human chemokines: an update. Annu Rev Immunol 1997; 15: 675–705
Marra F, Romanelli RG, Giannini C, et al. Monocyte chemotactic protein-1 as a chemoattractant for human hepatic stellate cells. Hepatology 1999 Jan; 29(1): 140–8
Rovin BH, Lu L, Saxena R. A novel polymorphism in the MCP-1 gene regulatory region that influences MCP-1 expression. Biochem Biophys Res Commun 1999 Jun; 259(2): 344–8
Feder JN, Gnirke A, Thomas W, et al. A novel MHC class I-like gene is mutated in patients with hereditary haemochromatosis. Nature Genet 1996 Aug; 13(4): 399–408
Powell LW, Subramaniam VN, Yapp TR. Haemochromatosis in the new millennium. J Hepatol 2000; 32: 48–62
Bomford A. Genetics of haemochromatosis. Lancet 2002 Nov; 360(9346): 1673–81
Pietrangelo A. Iron-induced oxidant stress in alcoholic liver fibrogenesis. Alcohol 2003 Jun; 30(2): 121–9
Tung BY, Emond MJ, Bronner MP, et al. Hepatitis C, iron status, and disease severity: relationship with HFE mutations. Gastroenterology 2003 Feb; 124(2): 318–26
Noble NA, Border WA. Angiotensin II in renal fibrosis: should TGF-beta rather than blood pressure be the therapeutic target? Semin Nephrol 1997 Sep; 17(5): 455–66
Inoue I, Nakajima T, Williams CS, et al. A nucleotide substitution in the promoter of human angiotensinogen is associated with essential hypertension and affects basal transcription in vitro. J Clin Invest 1997 Apr; 99(7): 1786–97
Ioannidis JPA. Genetic associations: false or true? Trends Mol Med 2003 Apr; 9(4): 135–8
Todd JA. Statistical false positive or true disease pathway? Nature Genet 2006 Jul; 38(7): 731–3
Huang HJ, Shiffman ML, Friedman S, et al. A 7 gene signature identifies the risk of developing cirrhosis in patients with chronic hepatitis C. Hepatology 2007 Aug; 46(2): 297–306
Jorgenson E, Witte JS. A gene-centric approach to genome-wide association studies. Nature Rev Genet 2006 Nov; 7(11): 885–91
Frayling TM. Commentary: genetic association studies see light at the end of the tunnel. Int J Epidemiol 2008 Feb; 37(1): 133–5
Ioannidis JP, Boffetta P, Little J, et al. Assessment of cumulative evidence on genetic associations: interim guidelines. Int J Epidemiol 2008 Feb; 37(1): 120–32
Fattovich G, Giustina G, Degos F, et al. Morbidity and mortality in compensated cirrhosis type C: a retrospective follow-up study of 384 patients. Gastroenterology 1997 Feb; 112(2): 463–72
Poustchi H, Negro F, Hui J, et al. Insulin resistance and response to therapy in patients infected with chronic hepatitis C virus genotypes 2 and 3. J Hepatol 2008 Jan; 48(1): 28–34
Walsh MJ, Jonsson JR, Richardson MM, et al. Non-response to antiviral therapy is associated with obesity and increased hepatic expression of suppressor of cytokine signalling 3 (SOCS-3) in patients with chronic hepatitis C, viral genotype 1. Gut 2006 Apr; 55(4): 529–35
Jonsson JR, Clouston AD, Ando Y, et al. Angiotensin-converting enzyme inhibition attenuates the progression of rat hepatic fibrosis. Gastroenterology 2001 Jul; 121(1): 148–55
Paizis G, Gilbert RE, Cooper ME, et al. Effect of angiotensin II type 1 receptor blockade on experimental hepatic fibrogenesis. J Hepatol 2001 Sep; 35(3): 376–85
Terui Y, Saito T, Watanabe H, et al. Effect of angiotensin receptor antagonist on liver fibrosis in early stages of chronic hepatitis C. Hepatology 2002 Oct; 36(4): 1022
Yokohama S, Yoneda M, Haneda M, et al. Therapeutic efficacy of an angiotensin II receptor antagonist in patients with nonalcoholic steatohepatitis. Hepatology 2004 Nov; 40(5): 1222–5
Friedman SL, Bansal MB. Reversal of hepatic fibrosis: fact or fantasy? Hepatology 2006 Feb; 43(2): S82–8
Burchard EG, Ziv E, Coyle N, et al. The importance of race and ethnic background in biomedical research and clinical practice. N Engl J Med 2003 Mar; 348(12): 1170–5
Lazarus R, Vercelli D, Palmer LJ, et al. Single nucleotide polymorphisms in innate immunity genes: abundant variation and potential role in complex human disease. Immunol Rev 2002 Dec; 190(1): 9–25
Amos CI. Successful design and conduct of genome-wide association studies. Hum Mol Genet 2007 Oct; 16: R220–5
Powell EE, Jonsson JR, Clouston AD. Steatosis: co-factor in other liver diseases. Hepatology 2005 Jul; 42(1): 5–13
Paradis V, Mathurin P, Laurent A, et al. Histological features predictive of liver fibrosis in chronic hepatitis C infection. J Clin Pathol 1996 Dec; 49(12): 998–1004
El-Serag HB, White DL, Mitra N. Genetic association studies: from “searching under the lamppost” to “fishing in the pond”. Gastroenterology 2008; 134(3): 662–4
Acknowledgments
Funding for the authors’ research was provided by the National Health and Medical Research Council of Australia, the Sasakawa Foundation (Royal Children’s Hospital), the Queensland Government’s Smart State Health and Medical Research Fund, and the Princess Alexandra Hospital Research and Development Foundation.
The authors have no conflicts of interest that are directly relevant to the content of this review.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jonsson, J.R., Purdie, D.M., Clouston, A.D. et al. Recognition of Genetic Factors Influencing the Progression of Hepatitis C. Mol Diag Ther 12, 209–218 (2008). https://doi.org/10.1007/BF03256286
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF03256286