Abstract
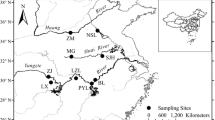
Determining the genetic structure is essential for developing conservation and stock improvement plans. Four dinucleotide microsatellite loci were analysed to reveal population genetic structure of the Indian major carp,Labeo rohita collected from three major rivers namely the Halda, the Jamuna, and the Padma in Bangladesh. The four loci were polymorphic (P 95) in all the populations. The populations varied in the number and frequencies of alleles as well as heterozygosities in the loci analyzed. Population differentiation (F ST) value between the Halda and the Jamuna population was significant (P<0.05). Relatively high level of gene flow and low level ofF ST values were found between the Padma and the Jamuna population. The unweighted pair group method with averages (UPGMA) dendrogram based on genetic distance resulted in two clusters: the Halda population was alone in one cluster whereas the Jamuna and the Padma made another cluster. The results revealed a relatively low level of genetic variability in the river populations ofL. rohita in Bangladesh.
Similar content being viewed by others
References
Alam MA, Akanda MSH, Khan MMR andAlam MS (2002) Comparison of genetic variability between a hatchery and a river population of rohu (Labeo rohita) by allozyme electrophoresis. Pakistan J. Biol. Sci. 5: 959–961.
Alam MA andAlam MS (2003) Comparative studies on developmental instability, growth and genetic variability of selected hatchery and one natural population of rohu (Labeo rohita). BAU Res. Prog. 13: 131–132.
Alam MS andIslam MS (2005) Population genetic structure ofCatla catla (Hamilton) revealed by microsatellite DNA markers. Aquaculture 246: 151–160.
Banglapedia (2006)Banglapedia-the National Encyclopedia of Bangladesh. Asiatic Society of Bangladesh.
Barman HK, Barat A, Yadav BM, Banerjee S, Meher PK, Reddy PVGK andJana RK (2003) Genetic variation between four species of Indian major carps as revealed by random amplified polymorphic DNA assay. Aquaculture 217: 115–123.
BBS (2000)Statistical Year Book. Bangladesh Bureau of Statistics, Department of Statistics, People's Republic of Bangladesh.
Das P, Barat A, Meher PK, Ray PP andMajumdar D (2005) Isolation and characterization of polymorphic microsatelhtes inLabeo rohita and their cross-species amplification in related species. Mol. Ecol Notes 5: 232–233.
DeWoody JA andAvise JC (2000) Microsatellite variation in marine, freshwater and anadromous fishes compared with other animals. J. Fish. Biol. 56: 461–473.
DoF (2008)Compendium- Fisheries Resources Development Campaign 2008, Department of Fisheries, Ministry of Fisheries and livestock, Government of the Peoples' Republic of Bangladesh, 96 pp.
FAO (2006)State of World Aquaculture 2006. FAO Fisheries Technical paper 500.
Ferguson MM (1995) The role of molecular genetic markers in the management of cultured fishes. InMolecular genetics in fisheries, G. R. Carvalho and T. J. Pitcher, Eds., Chapman & Hall Inc., London. pp. 81–103.
Goudet J (2001) FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). Available from http://www.unil.ch/izea/soft-wares/fstat.html.
Islam MS, Ahmed ASI, Azam MS andAlam MS (2005) Genetic analysis of three river populations ofCatla catla (Hamilton) usuing randomly amplified polymorphic DNA markers. Asian-Aust. J. Anim. Sci. 18(4): 453–457.
Islam MS andAlam MS (2004) Randomly Amplified Polymorphic DNA analysis of four different populations of the Indian major carpLabeo rohita (Hamilton). J. Appl. Ichthyol. 20: 407–412.
Khan MMR, Alam MS andBhuiyan MH (2006) Allozyme variation of hatchery and river populations of rohu (Labeo rohita, Hamilton) in Bangladesh. Aquacult. Res. 37: 233–240.
Majumdar KC, Ravinder K andNasaruddin K (1997) DNA fingerprinting in Indian carps and tilapia by Bkm 2(8) and M13 probes. Aquacult. Res. 28: 129–138.
Nash JHE (1991) DNAfrag, Version 3.03. Institute for Biological Sciences, National Research Council of Canada, Ottawa, Ontario, Canada.
Nei M (1972) Genetic distance between populations. Am. Nat. 106: 283–292,
Norris AT.Bradley DG andCunningham EP (1999) Microsatellite genetic variation between and within farmed and wild Atlantic salmon (Salmo salar) using microsatellite markers. Aquaculture 182: 73–83.
Padhi BK andMandai RK (1997) Inadvertent hybridization in a carp hatchery as detected by nuclear DNA RFLP. J. Fish Biol. 50: 906–909.
Paetkau D andStrobeck C (1995) The molecular basis and evolutionary history of a microsatellite null allele in bears. Mol. Ecol. 4: 519–520.
Page RDM (1996) TREEVIEW: An application to display phylogenetic trees on personal computers. Computer Appl. Biosci. 12: 357–358.
Peaka R andSmouse PE (2006) GENEALEX V6: Genetic Analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 6: 288–295.
Siegismund HR (1995) GSTAT, Version 3.1. Genetical statistical programs for the analysis of population data. Botanical Institute, University of Copenhagen, Denmark.
Simonsen V, Hansen MM, Mensberg K-LD, Sarder Rl andAlam S (2005) Widespread hybridization among species of Indian major carps in hatcheries, but not in the wild. J. Fish. Biol. 67: 794–808.
Wang S, Har JJ andUtter F (2002) Salmonid inbreeding: a review. Rev. Fish Biol. Fish., 11: 301–319.
Was A andWenne R (2002) Genetic differentiation in hatchery and wild sea trout (Salmo trutta) in the southern Baltic at microsatellite loci. Aquaculture 204: 493–506.
Welcomme RL (1988)International introductions of inland aquatic species. FAO Fish. Tech. Pap. No. 294, Rome, FAO, 318 pp.
Yeh FC, Yang RC and Boyle T (1999) POPGENE VERSION 1.32: Microsoft Window-based freeware for population genetic analysis (httpV/www.ualber-ta.ca/_fyeh).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Alam, M.S., Jahan, M., Hossain, M.M. et al. Population genetic structure of three major river Populations of rohu,Labeo rohita (Cyprinidae: Cypriniformes) using microsatellite DNA markers. Genes & Genomics 31, 43–51 (2009). https://doi.org/10.1007/BF03191137
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF03191137