Abstract
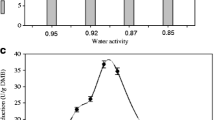
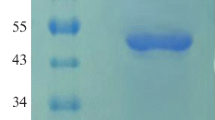
TheAspergillus ficuum phytase genephyA was overexpressed inPichia pastoris X33 after replacing Buffered glycerol-complex medium (BMGY medium) using 1% (v/v) glycerol with fresh Buffered methanol-complex medium (BMMY medium) using 1% (v/v) methanol (on daily basis) as carbon sources. The phytase activity increased evidently with the induction time, and reached 200 U mL−1 after 9 days of induction. We examined the possibility of employing thus obtained phytase to recover phosphorus from the fermented liquid of rice bran. When the 0.1 M sodium acetate buffer was replaced with de-ionised water (pH 5.5±0.1) as an enzyme reaction solution, there was an increase in the phosphorus recovery with respect to time and reached 1.31% after 24 h incubation contributing to 81% release of inorganic P from the rice bran phytate. Studies on hydrolysis of rice bran phytate by the addition of different concentrations of phytase ranging from 0–200 U mL−1 produced through the recombinant yeast shows no significant effect in the rate of phytate hydrolysis at enzyme activities of 200, 100, 50 U mL−1. However, rate of hydrolysis varied significantly at 20, 5, 0 U mL−1 enzyme concentrations.
Similar content being viewed by others
References
Bae H.D., Yanke L.J., Cheng K.J., Selinger L.B. (1999). A novel staining method for detecting phytase activity. J. Microbiol. Methods, 39: 17–22.
Chen C.C., Wu P.H., Huang C.T., Cheng K.J. (2004). APichia pastoris fermentation strategy for enhancing the heterologous expression of anEscherichia coli phytase. Enzyme Microbiol. Technol., 35: 315–320.
Erdman L.W., Poneros S.K. (1989). Phytic acid interaction with divalent cations in foods and in the gastrointestinal tract. Adv. Exp. Med. Biol., 249:161–171.
Han Y., Lei X.G. (1999). Role of glycosylation in functional expression of anAspergillus niger phytase (phyA) inPichia pastoris. Arch. Biochem. Biophys., 364:83–90.
Han W.Y., Wilfred, A.G. (1988). Phytate hydrolysis in soybean and cottonseed meals byAspergillus ficuum phytase. J. Agric. Food Chem., 36:259–262.
Haefner S., Knietsch A., Scholten E., Braun J., Lohscheidt M., Zelder O. (2005). Biotechnological production and applications of phytases. Appl. Microbiol. Biotechnol., 68:588–597.
Howson S. J., Davis R. P. (1983). Production of phytate-hydrolysing enzyme by some fungi, Enzyme Microb. Technol., 5:377–382.
Lei X.G., Ku P.K., Miller E.R., Yokoyama M.T. (1993). Supplementing corn-soybean meal diets with microbial linearly improves phytate P utilization by weanling pigs. J. Anim. Sci., 71: 3359–3367.
Maga J.A. (1982). Phytate: its chemistry, occurrence, food interactions, nutritional significance, and methods of analysis. J. Agric. Food Chem., 30:1–9.
Mayer A.F., Hellmuth K., Schlieker H., Lopez-Ulibarri Oertel R.S., Dahlems U., Strasser A. W. M., van Loon A. P. G. M. (1999). An expression system matures: a highly efficient and cost-effective process for phytase production by recombinant strains ofHansenula polymorpha. Biotech. Bioeng., 63:373–381.
Reddy N.R., Sathe S.K., Salunkhe D.K., (1982). Phytates in legumes and cereals. Adv. Food Res., 28:1–92.
Rimbach G.H., Ingelmann J., Pallauf J. (1994). The role of phytase in the dietary bioavailability of minerals and trace elements. Ernahrungsforschung. 39:1–10.
Shieh E.T., Ware J.H. (1968). Survey of microorganisms for the production of extracellular phytase. Appl Microbiol., 16:1348–1351.
Shimizu M. (1992). Purification and characterization of phytase fromBacillus subtilis (natto) N-77. Biosci. Biotechnol. Biochem., 56:1266–1269.
Quan C.L., Zhang Y., Wang Y., Ohta Y. (2001). Production of phytase in low phosphate medium by a novel yeastCandida krusei. J. Biosci. Bioeng., 92:154–160.
Ullah A.H.J., Sethumadhavan K. (2003).PhyA gene product ofAspergillus ficuum andPeniophora lycii produces dissimilar phytases. Biochem. Biophy. Res. Commun., 303: 463–468.
Van Hartingsveldt W., van Zeijl C.M.J.G.M., Harteveld M., Gouka R.J., Guykerbuyk Luiten M.E.G., van Paridon R.G.M.M., Selten G.C., Veenstra A.E., van Gorcom R.F.M., van den Hondel C.A.M.J.J. (1993). Cloning, characterization and expression of the phytase-encoding genephyA ofAspergillus niger. Gene. 127; 87–94.
Zhang Y., Liu R., Wu X. (2007). The proteolytic systems and heterologous proteins degradation in the methylotrophic yeastPichia pastoris. Ann. Microbiol., 57: 553–560.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chang, MH., Young, CC., Chien, SY. et al. Expression of recombinantPichia pastoris X33 phytase for dephosphorylation of rice bran fermented liquid. Ann. Microbiol. 58, 233–238 (2008). https://doi.org/10.1007/BF03175322
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF03175322