Abstract
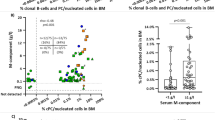
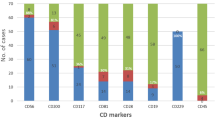
In an attempt to optimise stem cell graft evaluation we have developed a method of quantifying the number of cells in a phenotypically defined population of cells, expressing a gene of interest by combining an RT-PCR method working on whole single cells with flow cytometry. The clinical potential is illustrated by two examples. First, the phenotypes of clonal cells in the bone marrow (BM) of a patient with multiple myeloma (MM), were determined by sorting cells phenotypically defined by their expression of surface antigens and then performing RT-PCR on the individual sorted cells using the rearranged immunogiobulin heavy chain (IgH) gene as clonai marker. All plasma cells with the phenotype CD38++/CD45RA-expressed the clonai marker, whereas it could not be detected in plasma cells with the phenotype CD38++/CD45RA+ A minor population of clonai cells with the CD38+CD45RA- phenotype was found. Second, the number of committed (CD34+/CD38+) and non-committed (CD34+/CD38-) stem cells, expressing the chimeric fusion gene p210 BCR/ABL in the autografi from a patient with chronic myeloid leukemia (CML), was determined. The number of cells expressing BCR/ABL mRNA was nearly equal in the CD34VCD38+ and CD34+CD38- compartment (8.1 and 8.5%). The method presented can easily be applied to determine the phenotype of malignant cells, where a unique mRNA species exist. Furthermore, the method allows one to predict the outcome of antibody mediated purging experiment.
Similar content being viewed by others
References
Shpall EL, Jones RB. Release of tumor cells from bone marrow [editorial; comment].Blood 1994;83:623–6255.
Deisseroth ABet al. Genetic marking shows that Ph+ cells present in autologous transplants of chronic myelo- genous leukemia (CML) contribute to relapse after autologous bone marrow in CML.Blood 1994;83:3068–30766.
Gribben JGet al. Immunological purging of marrow assessed by PCR before autologous bone marrow transplantation for B-cell lymphoma.N Engl J Med 1991;325:1525–15333.
Rasmussen T, Johnsen HE. A quantitative analysis for CD34 mRNA+ haematopoietic stem cells.Int J Hematol 1996;64: Supplement 1, 28 (Abstract).
Satterthwaite AB, Burn TC, Le Beau MM, Tenen DG. Structure of the gene encoding CD34, a human hemato- poietic stem cell antigen.Genomics 1992;12:788–7944.
Cross NCet al. Minimal residual disease after allogeneic bone marrow transplantation for chronic myeloid leukaemia in first choice phase: correlations with acute graft-versus-host disease and relapse.Br J Haematol 1993;84:67–744.
Sambrook J, Fritch EF, Maniatis T.Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory Press: New York, 1989.
Deane M, Norton JD. Immunoglobulin gene ‘fingerprinting’: an approach to analysis of B lymphoid clonality in lymphoproliferative disorders.Br J Haematol 1991;77:274–2811.
Cook GR, Tomlinson IM. The human immunoglobulin VH repertoire.Immunol Today 1995;16:237–2422.
Sanz I. Multiple mechanisms participate in the generation of diversity of human H chain CDR3 regions. Jtmmunol 1991;147:17200.
Hoppe BL, Conti-Tronconi BM, Horton RM. Gel-loading dyes compatible with PCR.Biotechniques 1996;12:679–6800.
Molesh DA, Hall JM. Quantitative analysis of CD34 + stem cells using RT-PCR on whole cells.PCR Methods Appl 1994;3:278–2844.
Harada Het al. Phenotypic difference of normal plasma cells from mature myeloma cells.Blood 1993;81:2658–26633.
Urashima M, Chauhan D, Uchiyama H, Anderson CD40 ligand triggered Interleukin-6 secretion in multiple myeloma.Blood 1995;85:1903–19122.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rasmussen, T., Honoré, L. & Johnsen, H.E. Identification and characterisation of malignant cells using PT-PCR on single flow-sorted cells. Med Oncol 15, 96–102 (1998). https://doi.org/10.1007/BF02989586
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF02989586