Abstract
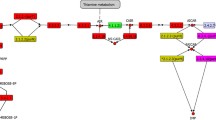
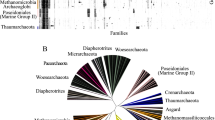
More than 100 sequenced genomes were searched for genes coding for the enzymes involved in glycolysis in an effort to find the most frequently occurring ones. Triosephosphate isomerase (TIM), glyceraldehyde-3-phosphate dehydrogenase (GAPD), phosphoglycerate kinase (PGK) and enolase (ENOL) were found to be present in 90 investigated genomes all together. The final set consisted of 80 prokaryotic and 10 eukaryotic genomes. Of the 80 prokaryotic genomes, 73 were from Bacteria, 7 from Archaea. Two microbial genomes were also from Eucarya (yeasts). Eight genomes of nonmicrobial origin were included for comparison. The amino acid sequences of TIMs, GAPDs, PGKs and ENOLs were collected and aligned, and their individual as well as concatenated evolutionary trees were constructed and discussed. The trees clearly demonstrate a closer relatedness between Eucarya and Archaea (especially the concatenated tree) but they do not support the hypothesis that eukaryotic glycolytic enzymes should be closely related to their α-proteobacterial counterparts. Phylogenetic analyses further reveal that although the taxonomic groups (e.g., α-proteobacteria, γ-proteobacteria, firmicutes, actinobacteria,etc.) form their more or less compact clusters in the trees, the inter-clade relationships between the trees are not conserved at all. On the other hand, several examples of conservative relatedness separating some clades of the same taxonomic groups were observed,e.g., Buchnera along withWigglesworthia and the rest of γ-proteobacteria, or mycoplasmas and the rest of firmicutes. The results support the view that these glycolytic enzymes may have their own evolutionary history.
Similar content being viewed by others
References
Antonyuk S.V., Eady R.R., Strange R.W., Hasnain S.S.: The structure of glyceraldehyde-3-phosphate dehydrogenase fromAlcaligenes xylosoxidans at 1.7 Å resolution.Acta Crystallogr. D59, 835–842 (2003).
Bairoch A., Apweiler R.: The Swiss Prot protein sequence database and its supplement TrEMBL, in 2000.Nucl.Acids Res. 28, 45–48 (2000).
Benson D.A., Karsch-Mizrachi I., Lipman D.J., Ostell J., Rapp B.A., Wheeler D.L.: GenBank.Nucl.Acids Res. 28, 15–18 (2000).
Bernstein B.E., Williams D.M., Bressi J.C., Kuhn P., Gelb M.H., Blackburn G.M., Hol W.G.: A bisubstrate analog induces unexpected conformational changes in phosphoglycerate kinase fromTrypanosoma brucei.J.Mol.Biol. 279, 1137–1148 (1998).
Canback B., Andersson S.G., Kurland C.G.: The global phylogeny of glycolytic enzymes.Proc.Nat.Acad.Sci.USA 99, 6097–6102 (2002).
Cordwell S.J.: Microbial genomes and “missing enzymes”: redefining biochemical pathways.Arch.Microbiol. 172, 269–279 (1999).
Dandekar T., Schuster S., Snel B., Huynen M., Bork P.: Pathway alignment: application to the comparative analysis of glycolytic enzymes.Biochem.J. 343, 115–124 (1999).
Erlandsen H., Abola E.E., Stevens R.C.: Combining structural genomics and enzymology: completing the picture in metabolic pathways and enzyme active sites.Curr.Opin.Struct.Biol. 10, 719–730 (2000).
Felsenstein J.: Confidence limits on phylogenies: an approach using the bootstrap.Evolution 39, 783–791 (1985).
Figge R.M., Cerff R.: GAPDH gene diversity in spirochetes: a paradigm for genetic promiscuity.Mol.Biol.Evol. 18, 2240–2249 (2001).
Fleming T., Littlechild J.: Sequence and structural comparison of thermophilic phosphoglycerate kinases with a mesophilic equivalent.Comp.Biochem.Physiol. A118, 439–451 (1997).
Fothergill-Gilmore L.A.: The evolution of glycolytic pathway.Trends Biochem.Sci. 11, 47–51 (1986).
Fothergill-Gilmore L.A., Michels P.A.M.: Evolution of glycolysis.Progr.Biophys.Mol.Biol. 59, 105–235 (1993).
Galperin M.Y., Koonin E.V.: Functional genomics and enzyme evolution. Homologous and analogous enzymes encoded in microbial genomes.Genetica 106, 159–170 (1999).
Gebbia J.A., Backenson P.B., Coleman J.L., Anda P., Benach J.L.: Glycolytic enzyme operon ofBorrelia burgdorferi: characterization and evolutionary implications.Gene 188, 221–228 (1997).
Gogarten J.P., Olendzenski L., Hilario E., Simon C., Holsinger K.E.: Dating the cenancester of organisms.Science 274, 1750–1751 (1996).
Hannaert V., Brinkmann H., Nowitzki U., Lef J.A., Albert M.A., Sensen C.W., Gaasterland T., Muller M., Michels P., Martin W.: Enolase fromTrypanosoma brucei, from the amitochondriate protistMastigamoeba balamuthi, and from the chloroplast and cytosol ofEuglena gracilis: pieces in the evolutionary puzzle of the eukaryotic glycolytic pathway.Mol.Biol.Evol. 17, 989–1000 (2000).
Henrissat B., Deleury E., Coutinho P.M.: Glycogen metabolism loss: a common marker of parasitic behavior in bacteria?Trends Genet. 18, 437–440 (2002).
Huynen M.A., Dandekar T., Bork P.: Variation and evolution of the citric-acid cycle: a genomic perspective.Trends Microbiol. 7, 281–291 (1999).
Isupov M.N., Fleming T.M., Dalby A.R., Crowhurst G.S., Bourne P.C., Littlechild J.A.: Crystal structure of the glyceraldehyde-3-phosphate dehydrogenase from the hyperthermophilic archaeonSulfolobus solfataricus.J.Mol.Biol. 291, 651–660 (1999).
Keeling P.J., Doolittle W.F.: Evidence that eukaryotic triosephosphate isomerase is of α-proteobacterial origin.Proc.Nat.Acad.Sci.USA 94, 1270–1275 (1997).
Klenk H.P., Clayton R.A., Tomb J.F., White O., Nelson K.E., Ketchum K.A., Dodson R.J., Gwinn M., Hickey E.K., Peterson J.D., Richardson D.L., Kerlavage A.R., Graham D.E., Kyrpides N.C., Fleischmann R.D., Quackenbush J., Lee N.H., Sutton G.G., Gill S., Kirkness E.F., Dougherty B.A., McKenney K., Adams M.D., Loftus B., Peterson S., Reich C.I., McNeil L.K., Badger J.H., Glodek A., Zhou L., Overbeek R., Gocayne J.D., Weidman J.F., McDonald L., Utterback T., Cotton M.D., Spriggs T., Artiach P., Kaine B.P., Sykes S.M., Sadow P.W., D’Andrea K.P., Bowman C., Fujii C., Garland S.A., Mason T.M., Olsen G.J., Fraser C.M., Smith H.O., Woese C.R., Venter J.C.: The complete genome sequence of the hyperthermophilic, sulfate-reducing archaeonArchaeoglobus fulgidus.Nature 390, 364–370 (1997).
Kohlhoff M., Dahm A., Hensel R.: Tetrameric trioscphosphate isomerase from hyperthermophilic Archaea.FEBS Lett. 383, 245–250 (1996).
Kováčová A., Janeček Š.: Evolutionary relationships of glycolytic (β/α)8-barrel enzymes present in completely sequenced genomes.Biologia (Bratislava) 57, 283–288 (2002).
Lebioda L., Stec B., Brewer J.M.: The structure of yeast enolase at 2.25-Å resolution. An 8-fold β+α-barrel with a novel ββαα (βα)6 topology.J.Biol.Chem. 264, 3685–3693 (1989).
Lolis E., Alber T., Davenport R.C., Rose D., Hartman F.C., Petsko G.A.: Structure of yeast triosephosphate isomerase at 1.9-Å resolution.Biochemistry 29, 6609–6618 (1990).
Martin W., Müller M.: The hydrogen hypothesis for the first cukaryote.Nature 392, 37–41 (1998).
Muirhead H., Watson H.: Glycolytic enzymes: from hexose to pyruvate.Curr.Opin.Struct.Biol. 2, 870–876 (1992).
van der Oost J., Huynen M.A., Verhees C.H.: Molecular characterization of phosphoglycerate mutase in archaea.FEMS Microbiol.Lett. 212, 111–120 (2002).
Page R.D.: TreeView: an application to display phylogenetic trees on personal computers.Comput.Applic.Biosci. 12, 357–358 (1996).
Pujadas G., Palau J.: TIM barrel fold: structural, functional and evolutionary characteristics in natural and designed molecules.Biologia (Bratislava) 54, 231–254 (1999).
Ronimus R.S., Morgan H.W.: Distribution and phylogenies of enzymes of the Embden-Meyerhof-Parnas pathway from archaea and hyperthermophilic bacteria support a gluconeogenic origin of metabolism.Archaea 1, 199–221 (2003).
Saiiou N., Nei M.: The neighbor-joining method: a new method for reconstructing phylogenetic trees.Mol.Biol.Evol. 4, 406–425 (1987).
Schmidt S., Sunyaev S., Bork P., Dandekar T.: Metabolites: a helping hand for pathway evolution?Trends Biochem.Sci. 28, 336–341 (2003).
Schuler G.D., Epstein J.A., Ohkawa H., Kans J.A.: Entrez: molecular biology database and retrieval system.Meth.Enzymol. 266, 141–162 (1996).
Skarzynski T., Moody P.C., Wonacott A.J.: Structure of holo-glyceraldehyde-3-phosphate dehydrogenase fromBocillus stearothermophilus at 1.8 Å resolution.J.Mol.Biol. 193, 171–187 (1987).
Stec B., Lebioda L.: Refined structure of yeast apo-enolase at 2.25 å resolution.J.Mol.Biol. 211, 235–248 (1990).
Thompson J.D., Higgins D.G., Gibson T.J.: CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment trough sequence weighting, position specific gap penalties and weight matrix choice.Nucl.Acids Res. 22, 4673–4680 (1994).
Tomb J.F., White O., Kerlavage A.R., Clayton R.A., Sutton G.G., Fleischmann R.D., Ketchum K.A., Klenk H.P., Gill S., Dougherty B.A., Nelson K., Quackenbush J., Zhou L., Kirkness E.F., Peterson S., Loftus B., Richardson D., Dodson R., Khalak H.G., Glodek A., McKenney K., Fitzegerald L.M., Lee N., Adams M.D., Hickey F.K., Berg D.E., Gocayne J.D., Utterback T.R., Peterson J.D., Kelley J.M., Cotton M.D., Weidman J.M., Fujii C., Bowman C., Watthey L., Wallin E., Hayes W.S., Borodovsky M., Karp P.D., Smith H.O., Fraser C.M., Venter J.C.: The complete genome sequence of the gastric pathogenHelicobacter pylori.Nature 388, 539–547 (1997).
Velanker S.S., Ray S.S., Gokhale R.S., Suma S., Balaram H., Balaram P., Murthy M.R.N.: Triosephosphate isomerase fromPlasmodium falciparum: the crystal structure provides insights into antimalarial drug design.Structure 5, 751–761 (1997).
Verhees C.H., Kengen S.W.M., Tuininga J.E., Schut G.J., Adams M.W.W., de Vos W.M., Van der Oost J.: The unique features of glycolytic pathways in Archaea.Biochem.J. 375, 231–246 (2003).
Watson H.C., Walker N.P., Shaw P.J., Bryant T.N., Wendell P.L., Fothergill L.A., Perkins R.E., Conroy S.C., Dobson M.J., Tuite M.F., Kingsman A.J., Kingsman S.M.: Sequence and structure of yeast phosphoglycerate kinase.EMBO J. 1, 1635–1640 (1982).
Wedekind J.E., Poyner R.R., Reed G.H., Rayment I.: Chelation of serine-39 to Mg2+ latches a gate at the active site of enolase: structure of the bis(Mg2+) complex of yeast enolase and the intermediate, phosphonoacetohydroxamate, at 2.1 Å resolution.Biochemistry 33, 9333–9342 (1994).
Author information
Authors and Affiliations
Corresponding author
Additional information
This work was supported by the VEGA grant no. 2/2057/23 from theSlovak Grant Agency.
Rights and permissions
About this article
Cite this article
Oslancová, A., Janeček, Š. Evolutionary relatedness between glycolytic enzymes most frequently occurring in genomes. Folia Microbiol 49, 247–258 (2004). https://doi.org/10.1007/BF02931039
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF02931039