Abstract
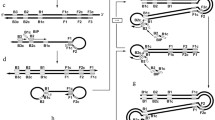
It has been considered that target DNA is a forgiving component for PCR amplification. Herein we present evidence to demonstrate that secondary structure located at the end of a template may interfere with the specificity of amplification. Experiments indicate that nonspecific amplification results from a long stretch of stem and loop structures at the 3′ end of prochymosin cDNA. Based on the sequence of mRNA coding for prochymosin, it is argued that the sequence responsible for the formation of the complex structure described here is most likely generated during synthesis of the second cDNA strand.
Similar content being viewed by others
References
McPherson, M. J., Quirke, P., and Taylor, G. R. (1991),PCR: A Practical Approach. Oxford University Press, Oxford.
Saiki, R. K. (1989), InPCR Technology, Henry A. Erlish (ed.), Stockton Press, New York, pp. 7–16.
Zhang, Y. Y., Zhou, W., Liu, N. J., and Yang, K. Y. (1991),Chin. J. Biotechnol. 7, 195.
Harris, T. J. R., Lowe, P. A., Lyons, A., Thomas, P. G., Eaton, M. A. W., Millican, T. A., Patel, T. P., Boss, C. C., Carey, N. H., and Doel, M. T. (1982),Nucleic Acids Res. 10, 2177.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Zhang, Z., Huang, K., Zhang, Y. et al. Interference by complex structures of target DNA with specific PCR amplification. Appl Biochem Biotechnol 44, 15–20 (1994). https://doi.org/10.1007/BF02921847
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02921847