Abstract
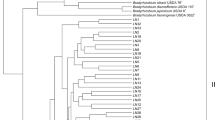
Nodule samples were collected from four alder species:Alnus nepalensis, A. sibirica, A. tinctoria andA. mandshurica growing in different environments on Gaoligong Mountains, Yunnan Province of Southwest China and on Changbai Mountains, Jilin Province of Northeast China. PCR-RFLP analysis of the IGS betweennifD andnifK genes was directly applied to unculturedFrankia strains in the nodules. A total of 21 restriction patterns were obtained. TheFrankia population in the nodules ofA. nepalensis had the highest genetic diversity among all fourFrankia populations; by contrast, the population in the nodules ofA. mandshurica had the lowest degree of divergence; the ones in the nodules ofA. sibirica andA. tinctoria were intermediate. A dendrogram, which was constructed based on the genetic distance between the restriction patterns, indicated thatFrankia strains fromA. sibirica andA. tinctoria had a close genetic relationship.Frankia strains fromA. nepalensis might be the ancestor ofFrankia strains infecting otherAlnus species. From these results and the inference of the ages ofAlnus host species, it is deduced that there was a co-evolution betweenAlnus and its microsymbiontFrankia in China.
Similar content being viewed by others
References
Normand, P., Orso, S., Cournoyer, B. et al., Molecular phylogeny of genusFrankia and related genera and emendation of the family Frankiaceae, Int. J. Syst. Bacteriol., 1996, 46: 1–9.
Guan, C., Pawlowski, P., Bisseling, T., Interaction betweenFrankia and actinorhizal plants, Subcelluar Biochem., 1998, 29: 165–189.
Lumini, E., Bosco, M., Polymerase chain reaction-restriction fragment length polymorphisms for assessing and increasing biodiversityof Frankia culture collections, Can. J. Bot., 1999, 77: 1261–1269.
Chen, Z. D., Origin and disperse of Betulaceae, in the Geography of Spermatophytic Families and Genera (ed. Lu, A. M.), Beijing: Science Press, 1999, 236–258.
Chen, Z. D., Manchester, S. R., Sun, H. Y., Phylogeny and evolution of the Betulaceae as inferred from DNA sequences, morphology, and paleobotany, Am. J. Bot., 1999, 86: 1168–1181.
Xiong, Z., Li, W. J., Zhang, Z. Z. et al., The influence of altitude on the genetic of microsymbionts ofAlnus nepalensis-Frankia, Chin. J. Southwest Forest. Coll., 2001, 21: 205–209.
Wu, S. H., Zhang, H. W., Xiong, Z., Biodiversity ofFrankia strains in nodules fromAlnus andHippophae by ARDRA, Chin. J. App. Ecol., 2001, 12: 883–886.
Nalin, R., Normand, P., Domenach, A. M., Distribution and N2-fixing activity ofFrankia strains in relation to soil depth, Physiol. Plantarum, 1997, 99: 732–738.
Varghese, R., Chauhan, V., Misra, A., Hypervariable spacer regions are good sites for developing specific PCR-RFLP markers and PCR primers for screening actinorhizal symbionts, J. Bioscience, 2003, 28: 437–442.
Hahn, D., Nickel, A., Dawson, J., AssessingFrankia populations in plants and soil using molecular methods, FEMS Microbiol. Ecol., 1999, 29: 215–227.
Li, H., Guo, H. J., Dao, Z. L., Flora of Gaoligong Mountains, Beijing: Science Press, 2000.
Hao, Z. Q., Yu, D. Y., Deng, H. B. et al., Study on complexity of plant communities at different altitudes on the North Slope of Changbai Mountain, Chin. J. Forest. Res., 2002, 13: 17–20.
Ritchie, N. J., Myrold, D. D., Phylogenetic placement of unculturedCeanothus microsymbionts using 16S rRNA gene sequences, Can. J. Bot, 1999, 77: 1208–1213.
Mitchell, L. G., Bodenteich, A., Merril, C. R., Silver staining of DNA, in Basic DNA and RNA Protocols (ed. Harwood, A. J., Sheng, X. Y.), Beijing: Science Press, 2002, 67–71.
Yeh, F. C., Yang, R. C., Royle, T. B. et al., POPGENE V1.31, http://www.ualberta.ca/-fyeh, 1997.
Buntjer, J. B., PHYTOOL (Phylogenetic Computer Tool) version 1.23, Laboratory of Plant Breeding, Wageningen University, Netherlands, 1997–2000.
Felsenstein, J., PHYLP (Phylogeny Inference Package) version 3.5c, Department of Genetics, University of Washington, Seattle, 1993.
Page, R. D. M., TREEVIEW: An application to display phylogenetic trees on personal computers, Comp. Ap. Biosciences, 1996, 12: 357–358.
Qian, W., Ge, S., Analyses of population genetic structure by using dominant markers, Acta Genet. Sinica, 2001, 28: 244–255.
Zou, Y. P., Ge, S., Wang, X.D., Molecular Fingerprint in Systematic and Evolution Botany, Beijing: Science Press, 2001, 190.
Varghese, R., Chauhan, V., Misra, A., Evolutionary implication of nucleotide sequence relatedness betweenAlnus nepalensis andAlnus glutinosa and also between correspondingFrankia microsybionts, Plant Soil, 2003, 254: 219–227.
Navarro, E., Bousquent, J., Moiroud, A. et al., Molecular phylogenyof Alnus (Betulaceae), inferred from nuclear ribosomal DNA ITS sequence, Plant Soil, 2003, 245: 207–217.
Jeong, S. C., Ritchie, N. J., Myrold, D. D., Molecular phylogenies of plants andFrankia support multiple origins of actinorhizal symbioses, Mol. Phylogenet. Evol., 1999, 13: 493–503.
Simonet, P., Navarro, E., Rouvier, C., Co-evolution betweenFrankia populations and host plants in the family Casuarinaceae and consequent patterns of global dispersal, Environ. Microb., 1999, 1: 525–533.
Navarro, E., Jaffre, T., Gauthier, D. et al., Distribution ofGymnostama spp. microsymbioticFrankia strains in New Caledonia is related to soil type and to host-plant species, Mol. Ecol., 1999, 8: 1781–1788.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dai, Y., Zhang, C., Xiong, Z. et al. Correlations between the ages ofAlnus host species and the genetic diversity of associated endosymbioticFrankia strains from nodules. Sci. China Ser. C.-Life Sci. 48 (Suppl 1), 76–81 (2005). https://doi.org/10.1007/BF02889804
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02889804