Abstract
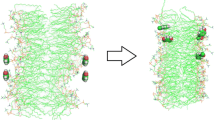
Emodin is one of the most abundant anthraquinone derivatives found in nature. It is the active principle of some traditional herbal medicines with known biological activities. In this work, we combined experimental and theoretical studies to reveal information about location, orientation, interaction and perturbing effects of Emodin on lipid bilayers, where we have taken into account the neutral form of the Emodin (EMH) and its anionic/deprotonated form (EM−). Using both UV/Visible spectrophotometric techniques and molecular dynamics (MD) simulations, we showed that both EMH and EM− are located in a lipid membrane. Additionally, using MD simulations, we revealed that both forms of Emodin are very close to glycerol groups of the lipid molecules, with the EMH inserted more deeply into the bilayer and more disoriented relative to the normal of the membrane when compared with the EM−, which is more exposed to interfacial water. Analysis of several structural properties of acyl chains of the lipids in a hydrated pure DMPC bilayer and in the presence of Emodin revealed that both EMH and EM− affect the lipid bilayer, resulting in a remarkable disorder of the bilayer in the vicinity of the Emodin. However, the disorder caused by EMH is weaker than that caused by EM−. Our results suggest that these disorders caused by Emodin might lead to distinct effects on lipid bilayers including its disruption which are reported in the literature.










Similar content being viewed by others

References
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002) Molecular biology of the cell, 4th edn. Garland, New York
Almeida JG, Preto AJ, Koukos PI, Bonvin AMJJ, Moreira IS (2017) Membrane proteins structures: a review on computational modeling tools. Biochim Biophys Acta 1859:2021–2039
Alves DS, Pérez-Fons L, Estepa A, Micol V (2004) Membrane-related effects underlying the biological activity of the anthraquinones emodin and barbaloin. Biochem Pharmacol 68:549–561
Anke H, Kolthum I, Laaatsch H (1980) Metabolic products of microorganisms. 192. The anthraquinones of the Aspergillus glaucus group. II. Biological activity. Arch Microbiol 126:231–236
Apostolova E, Krumova S, Tuparev N, Molina MT, Filipova T, Petkanchin I, Taneva SG (2003) Interaction of biological membranes with substituted 1,4-anthraquinones. Colloids Surf B 29:1–12
Barnard DL, Huffman JH, Morris JL, Wood SG, Hughes BG, Sidwell RW (1992) Evaluation of the antiviral activity of anthraquinones, anthrones and anthraquinone derivatives against human cytomegalovirus. Antivir Res 17:63–77
Becke AD (1993) Density-functional thermochemistry. III The role of exact exchange. J Chem Phys 98:5648–5652
Bemporad D, Essex JW, Luttmann C (2004) Permeation of small molecules through a lipid Bilayer: a computer simulation study. J Phys Chem B 108:4875–4884
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81:3684–3690
Berendsen HJC, Postma JPM, van Gunsteren WF, Hermans J (1981) Interaction models for water in relation to protein hydration. In: Pullman, B (ed) Intermolecular forces. Reidel, Dordrecht
Bi S, Zhang H, Qiao C, Sun Y, Liu C (2008) Studies of interaction of emodin and DNA in the presence of ethidium bromide by spectroscopic method. Spectrochim Acta Part A 69:123–129
Boggara MB, Krishnamoorti B (2010) Partitioning of nonsteroidal Antiinflammatory drugs in lipid membranes: a molecular dynamics simulation study. Biophys J 98:586–595
Breneman CM, Wiberg KB (1990) Determining atom-centered monopoles from molecular electrostatic potentials. The need for high sampling density in formamide conformational analysis. J Comp Chem 11:361–373
Brown MF (1996) Membrane structure and dynamics studied with NMR spectroscopy. In: Merz K, Roux B (eds) Biological membranes. Birkhäuser, Boston
Canuto S, Coutinho K (2000) From hydrogen bond to bulk: Solvation analysis of the n-π* transition of formaldehyde in water. Int J Quantum Chem 77:192–198
Chan TC, Chang CJ, Koonchanok NM, Geahlen RL (1993) Selective inhibition of the growth of ras-transformed human bronchial epithelial cells by emodin, a protein-tyrosine inhibitor. Biochem Biophys Res Commun 193:1152–1158
Chen YC, Shen SC, Lee WR, Hsu FL, Lin HY, Ko CH, Tseng SW (2002) Emodin induces apoptosis in human promyeloleukemic HL-60 cells accompanied by activation of caspase 3 cascade but independent of reactive oxygen species production. Biochem Pharmacol 64:1713–1724
Choi RJ, Ngoc TM, Bae K, Cho HJ, Kim DD, Chun J, Khan S, Kim YS (2013) Anti-inflammatory properties of anthraquinones and their relationship with the regulation of P-glycoprotein function and expression. Eur J Pharmacol Sci 48:272–281
Costigan SC, Booth PJ, Templer RH (2000) Estimations of lipid bilayer geometry in fluid lamellar phases. Biochim Biophys Acta 1468:41–54
Cuendet MA, van Gunsteren WF (2007) On the calculation of velocity-dependent properties in molecular dynamics simulations using the leapfrog integration algorithm. J Chem Phys 127:184102–184110
da Cunha AR, Duarte EL, Lamy MT, Coutinho K (2014) Protonation/deprotonation process of Emodin in aqueous solution and pKa determination: UV/visible spectrophotometric titration and quantum/molecular mechanics calculations. Chem Phys 440:69–79
Davis JE, Rahaman O, Patel S (2009) Molecular dynamics simulations of a DMPC Bilayer using nonadditive interaction models. Biophys J 96:385–402
De Young LR, Dill KA (1988) Solute partitioning into lipid bilayer membranes. Biochemist 27:5281–5289
Ditchfield R, Hehre WJ, Pople JA (1971) Self-consistent molecular-orbital methods. IX An Extended Gaussian Type Basis for Molecular Orbital Studies of Organic Molecules. J Chem Phys 54:724–728
Dong X, Fu J, Yin X, Cao S, Li X, Lin L, Huyiligegi NJ (2016) Emodin: a review of its pharmacology, Toxicity and Pharmacokinetics. Phytother Res 30:1207–1218
Douliez JP, Léonard A, Dufoure EJ (1995) Restatement of order parameters in biomembranes: calculation of C-C bond order parameters from C-D quadrupolar splittings. Biophys J 68:1727–1739
Duarte EL, Oliveira TR, Alves DS, Micol V, Lamy MT (2008) On the interaction of the Anthraquinone Barbaloin with negatively charged DMPG Bilayers. Langmuir 24:4041–4049
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577–8593
Fabriciova G, Cortés SS, Ramos JVG, Miskovsky P (2004) Surface-enhanced Raman spectroscopy study of the interaction of the Antitumoral drug Emodin with human serum albumin. Biopolymers 74:125–130
Falck E, Hautala JT, Karttunen M, Kinnunen PKJ, Patra M, Saaren-Seppälä H, Vattulainen I, Wiedmer SK, Holopainen JM (2006) Interaction of Fusidic acid with lipid membranes: implications to the mechanism of antibiotic activity. Biophys J 91:1787–1799
Francke B, Moss H, Timbury MC, Hay J (1978) Alkaline DNase activity in cells infected with a temperature-sensitive mutant of herpes simplex virus type 2. J Virol 26:209–213
Frisch MJ et al (2004) Gaussian 03. Gaussian, Wallingford, CT
Fuchs J, Milbradt R, Zimmer G (1990) Multifunctional analysis of the interaction of anthralin and its metabolites anthraquinone and anthralin dimer with the inner mitochondrial membrane. Arch Dermatol Res 282:47–55
Gabdouline RR, Vanderkooi G, Zheng C (1996) Comparison of structures of dimyrstoylphosphatidylcholine in the presence and absence of cholesterol by molecular dynamics simulation. J Phys Chem 96:15942–15946
Ghomi M (2012) Applications of Raman spectroscopy to biology, vol 5. IOS, Amsterdam
Gierula MP, Takaoka Y, Miyagawa H, Kitamura K, Kusumi A (1999) Charge pairing of headgroups in phosphatidylcholine membranes: a molecular dynamics simulation study. Biophys J 73:1228–1240
Goldstein EB, Evron Z, Frenkel M, Cohen K, Meiron KN, Peer D, Roichman Y, Flescher E, Fridman M (2011) Targeting Anthracycline-resistant tumor cells with synthetic aloe-Emodin glycosides. ACS Med Chem Lett 2:528–531
Heimburg T (2007) Thermal biophysics of membranes. Wiley-VCH, Weinheim
Hernandez M, Recio G, Palma RJM, Ramos JVG, Domingo C, Sevilla P (2012) Surface enhanced fluorescence of anti-tumoral drug emodin adsorbed on silver nanoparticles and loaded on porous silicon. Nanoscale Res Lett 7:364–371
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472
Hofsäß C, Lindahl E, Edholm O (2003) Molecular dynamics simulations of Phospholipid Bilayers with cholesterol. Biophys J 84:2192–2206
Högberg CJ, Maliniak A, Lyubartsev AP (2007) Dynamical and structural properties of charged and uncharged lidocaine in a lipid bilayer. Biophys Chem 125:416–424
Hsiang CY, Ho TY (2008) Emodin is a novel alkaline nuclease inhibitor that suppresses herpes simplex virus type 1 yields in cell cultures. Br J Pharmacol 155:227–235
Huang HC, Chu SH, Chao PD (1991) Vasorelaxants from Chinese herbs, emodin and scoparone, possess immunosuppressive properties. Eur J Pharmacol 198:211–213
Huang Q, Shen HM, Ong CN (2004) Inhibitory effect of emodin on tumor invasion through suppression of activator protein-1 and nuclear factor-kappaB. Biochem Pharmacol 68:361–371
Huang Q, Shen HM, Ong CN (2005) Emodin inhibits tumor cell migration through suppression of the phosphatidylinositol 3-kinase-Cdc42/Rac1 pathway. Cell Mol Life Sci 62:1167–1175
Huang Q, Shen HM, Shui G (2006) Emodin inhibits tumor cell adhesion through disruption of the membrane lipid raft-associated Integrin signaling pathway. Cancer Res 66:5807–5815
Huang W, Lin Z, van Gunsteren WF (2011) Validation of the GROMOS 54A7 force field with respect to beta-peptide folding. J Chem Theory Comput 7:1237–1243
Imamura Y, Otsuka T, Nakai H (2007) Description of Core excitations by time-dependent density functional theory with local density approximation, generalized gradient approximation, meta-generalized gradient approximation, and hybrid Hunctionals. J Comput Chem 28:2067–2074
Izhaki I (2002) Emodin – a secondary metabolite with multiple ecological functions in higher plants. New Phytol 155:205–217
Jalili S, Saeedi M (2016) Study of curcumin behavior in two different lipid bilayer models of liposomal curcumin using molecular dynamics simulation. J Biomol Struct Dyn 34:327–340
Jämbeck JPM, Lyubartsev AP (2012) Derivation and systematic validation of a refined all-atom force field for Phosphatidylcholine lipids. J Phys Chem B 116:3164–3179
Jayasuriya H, Koonchanok NM, Geahlen RL, McLaughlin L, Chang CJ (1992) Emodin, a protein tyrosine kinase inhibitor from Polygonum Cuspidatum. J Nat Prod 55:696–698
Jendrossek V, Handrick R (2003) Membrane targeted anticancer drugs: potent inducers of apoptosis and putative Radiosensitisers. Curr Med Chem Anticancer Agents 3:343–353
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935
Kawai K, Kato T, Mori H, Kitamura J, Nozawa Y (1984) A comparative study on cytotoxicities and biochemical properties ofanthraquinone mycotoxinsemodin and skyrin from Penicillium islandicum. Toxicol Lett 20:155–160
Koenig BW, Strey HH, Gawrisch K (1997) Membrane lateral compressibility determined by NMR and x-ray diffraction: effect of acyl chain polyunsaturation. Biophys J 73:1954–1966
Koukoulitsa C, Durdagi S, Siapi E, Villalonga-Barber C, Alexi X, Steele BR, Micha-Screttas M, Alexis MN (2011) Comparison of thermal effects of stilbenoid analogs in lipid bilayers using differential scanning calorimetry and molecular dynamics: correlation of thermal effects and topographical position with antioxidant activity. Eur Biophys J 40:865–875
Koyama M, Kelly TR, Watanabe KA (1988) Novel type of potential anticancer agents derived from chrysophanol and emodin. Some structure-activity relationship studies. J Med Chem 31:283–284
Kumar A, Dhawan S, Aggarwal BB (1998) Emodin (3-methyl-1,6,8-trihydroxyanthraquinone) inhibits TNF-induced NF-B activation, IB degradation, and expression of cell surface adhesion proteins in human vascular endothelial cells. Oncogene 17:913–918
Kuo YC, Meng HC, Tsai WJ (2001) Regulation of cell proliferation, inflammatory cytokine production and calcium mobilization in primary human T lymphocytes by emodin from Polygonum Hypoleucum Ohwi. Inflamm Res 50:073–082
Lis LJ, McAlister M, Fuller N, Rand RP, Parsegian VA (1982) Interactions between neutral phospholipid bilayer membranes. Biophys J 37:657–665
Lopez CF, Nielsen SO, Klein ML, Moore PB (2004) Hydrogen bonding structure and dynamics of water at the Dimyristoylphosphatidylcholine lipid Bilayer surface from a molecular dynamics simulation. J Phys Chem B 108:6603–6610
Loverde SM (2014) Molecular simulation of the transport of drugs across model membranes. J Phys Chem Lett 5:1659–1665
Lucio M, Nunes C, Gaspar D, Ferreira H, Lima JLFC, Reis S (2009) Antioxidant activity of vitamin E and Trolox: understanding of the factors that govern lipid peroxidation studies in vitro. Food Biophys 4:312–320
MacCallum JL, Tieleman DP (2006) Computer simulation of the distribution of hexane in a lipid Bilayer: spatially resolved free energy, entropy, and enthalpy profiles. J Am Chem Soc 128:125–130
Malde AK, Zuo L, Breeze M, Stroet M, Poger D, Nair PC, Oostenbrink C, Mark AE (2011) An automated force field topology builder (ATB) and repository: version 1.0. J Chem Theory Comput 7:4026–4037
Marković ZS, Manojlović NT (2009) DFT study on the reactivity of OH groups in emodin: structural and electronic features of emodin radicals. Monatsh Chem 140:1311–1318
Marsh D (1990) CRC handbook of lipid Bilayers. CRC, Boca Raton
Miertus S, Scrocco E, Tomasi J (1981) Electrostatic interaction of a solute with a continuum. A direct utilizaion of AB initio molecular potentials for the prevision of solvent effects. Chem Phys 55:117–129
Miyamoto S, Kollman PA (1992) An analytical version of the SHAKE and RATTLE algorithm for rigid water models. J Comput Chem 13:952–962
Moore PB, Lopez CF, Klein ML (2001) Dynamical properties of a hydrated lipid Bilayer from a multinanosecond molecular dynamics simulation. Biophys J 81:2484–2494
Nagle JF, Tristram-Nagle S (2000) Structure of lipid bilayers. Biochim Biophys Acta 1469:159–195
Nguyen SC, Hansen BKV, Hoffmann SV, Spanget-Larsen J (2008) Electronic states of emodin and its conjugate base. Synchrotron linear dichroism spectroscopy and quantum chemical calculations. Chem Phys 352:67–174
Nitschke WK, Suplicy CCV, Coutinho K, Stassen H (2012) Molecular dynamics investigations of PRODAN in a DLPC Bilayer. J Phys Chem B 116:2713–2721
Nunes C, Brezesinski G, Lopes D, Lima JLFC, Reis S, Lúcio M (2011) Lipid–drug interaction: biophysical effects of Tolmetin on membrane mimetic Systems of Different Dimensionality. J Phys Chem B 115:12615–12623
Omote H, Al-Shawi MK (2006) Interaction of transported drugs with the lipid Bilayer and P-glycoprotein through a Solvation exchange mechanism. Biophys J 90:4046–4059
Orsi M, Essex JW (2010) Permeability of drugs and hormones through a lipid bilayer: insights from dual-resolution molecular dynamics. Soft Matter 6:3797–3808
Pal T, Jana NR (1993) Emodin (1,3,8-trihydroxy-6-methylanthraquinone): a spectrophotometric reagent for the determination of beryllium(II), magnesium(II) and calcium(II). Analyst 118:1337–1342
Parr RG, Yang W (1994) Density functional theory of atoms and molecules. Oxford University Press, Oxford
Peetla C, Stine A, Labhasetwar V (2009) Biophysical interactions with model lipid membranes: applications in drug discovery and drug delivery. Mol Pharm 6:1264–1276
Pereira CS, Lins RD, Chandrasekhar I, Freitas LCG, Hünenberger PH (2004) Interaction of the disaccharide Trehalose with a Phospholipid Bilayer: a molecular dynamics study. Biophys J 86:2273–2285
Petrache HI, Dodd SW, Brown MF (2000) Area per lipid and acyl length distributions in fluid phosphatidylcholines determined by (2)H NMR spectroscopy. Biophys J 79:3172–3192
Petrache HI, Tristram-Nagle S, Nagle JF (1998) Fluid phase structure of EPC and DMPC bilayers. Chem Phys Lipid 95:83–94
Poger D, Mark AE (2010) On the validation of molecular dynamics simulations of saturated and cis-monounsaturated Phosphatidylcholine lipid Bilayers: a comparison with experiment. J Chem Theory Comput 6:325–336
Poger D, Mark AE (2012) Lipid Bilayers: the effect of force field on ordering and dynamics. J Chem Theory Comput 8:4807–4817
Poger D, Mark AE (2013) The relative effect of sterols and Hopanoids on lipid Bilayers: when comparable is not identical. J Phys Chem B 117:16129–16140
Poger D, van Gunsteren WF, Mark AE (2010) A new force field for simulating phosphatidylcholine bilayers. J Comput Chem 31:1117–1125
Rajarathnam K, Rösgen J (2014) Isothermal titration calorimetry of membrane proteins - progress and challenges. Biochim Biophys Acta 1838:69–77
Rand RP, Parsegian VA (1989) Hydration forces between phospholipid bilayers. Biochim Biophys Acta 988:351–376
Rissanen S, Kumorek M, Martinez-Seara H, Li YC, Jamroz D, Bunker A, Nowakowska M, Vattulainen I, Kepczynski M, Roǵ T (2014) Effect of PEGylation on drug entry into lipid Bilayer. J Phys Chem B 118:144–151
Robinson AJ, Richards WG, Thomas PJ, Hann MM (1995) Behavior of cholesterol and its effect on headgroup and chain conformations in lipid bilayers: a molecular dynamics study. Biophys J 68:164–170
Sabín J, Prieto G, Ruso JM, Messina PV, Salgado FJ, Nogueira M, Costas M, Sarmiento F (2009) Interactions between DMPC Liposomes and the serum blood proteins HSA and IgG. J Phys Chem B 113:1655–1661
Sachs JN, Petrache HI, Woolf TB (2003) Interpretation of small angle X-ray measurements guided by molecular dynamics simulations of lipid bilayers. Chem Phys Lipid 126:211–223
Saito ST, Silva G, Pungartnik C, Brendel MJ (2012) Study of DNA–emodin interaction by FTIR and UV–Vis spectroscopy. Photochem Photobiol B 111:59–63
Schmid N, Eichenberger AP, Choutko A, Riniker S, Winger M, Mark AE, van Gunsteren WF (2011) Definition and testing of the GROMOS force-field versions 54A7 and 54B7. Eur Biophys J 40:843–856
Seddon AM, Casey D, Law RV, Gee A, Templera RH, Cesab O (2009) Drug interactions with lipid membranes. Chem Soc Rev 38:2509–2519
Sevilla P, Blanco FG, Ramos JVG, Cortés SS (2009) Aggregation of antitumoral drug emodin on Ag nanoparticles: SEF, SERS and fluorescence lifetime experiments. Phys Chem Chem Phys 11:8342–8348
Sevilla P, De-Llanos R, Domingo C, Sanchez-Cortes S, Garcia-Ramos JV (2010) SERS plus MEF of the anti tumoral drug emodin adsorbed on silver nanoparticles. In: VoDinh T, Lakowicz JR (eds) Plasmonics in biology and medicine. Proceedings of SPIE, Volume 7577, 25 - 28 January, 2010, Bellingham, USA
Seydel JK, Wiese M (2002) Drug-membrane interactions: analysis, drug distribution, modeling, vol 15. , Weinheim
Sirk TW, Brown EF, Friedman M, Sum AK (2009) Molecular binding of Catechins to biomembranes: relationship to biological activity. J Agric Food Chem 57:6720–6728
Sirka KK (2014) Separation of molecules, macromolecules and particles: principles, phenomena and processes. Cambridge University Press, New York
Smaby JM, Momsen MM, Brockman HL, Brown RE (1997) Phosphatidylcholine acyl unsaturation modulates the decrease in interfacial elasticity induced by cholesterol. Biophys J 73:1492–1505
Smondyrev AM, Berkowitz ML (1999) Structure of Dipalmitoylphosphatidylcholine/cholesterol Bilayer at Lowand high cholesterol concentrations: molecular dynamics simulation. Biophys J 77:2075–2089
Smondyrev AM, Berkowitz ML (2001) Molecular dynamics simulation of the structure of Dimyristoylphosphatidylcholine Bilayers with cholesterol, Ergosterol, and Lanosterol. Biophys J 80:1649–1658
Spoel DVD, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJC (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718
Srinivas G, Anto RJ, Srinivas P, Vidhyalakshmi S, Senan VP, Karunagaran D (2003) Emodin induces apoptosis of human cervical cancer cells through poly(ADP-ribose) polymerase cleavage and activation of caspase-9. Eur J Pharmacol 473:117–125
Thomson RH (1987) Naturally occurring Quinones, vol 3. Chapman and Hall, London
Tieleman DP, Marrink SJ, Berendsen HJC (1997) A computer perspective of membranes: molecular dynamics studies of lipid bilayer systems. Biochim Biophys Acta 1331:235–270
To LV (1984) Emodin - a fungal metabolite - and the effects of Emodin on the growth of some soil microorganisms. Acta Agrar Silv Ser Agrar 23:235–242
Vargas F, Rivas C, Medrano M (2004) Interaction of Emodin, aloe-Emodin, and Rhein with human serum albumin: a fluorescence spectroscopic study. Toxicol Mech Methods 14:227–231
Vermeer LS, de Groot BL, Réat V, Milon A, Czaplicki J (2007) Acyl chain order parameter profiles in phospholipid bilayers: computation from molecular dynamics simulations and comparison with 2H NMR experiments. Eur Biophys J 36:919–931
Wang L, Lin L, Ye BJ (2006) Electrochemical studies of the interaction of the anticancer herbal drug emodin with DNA. Pharm Biomed Anal 42:625–629
Wang WH, Chung JG (1997) Emodin-induced inhibition of growth and DNA damage in the helicobacter pylori. Curr Microbiol 35:262–266
Witzke S, Duelund L, Kongsted J, Petersen M, Mouritsen OG, Khandelia H (2010) Inclusion of Terpenoid plant extracts in lipid Bilayers investigated by molecular dynamics simulations. J Phys Chem B 114:15825–15831
Xiang TX, Anderson BD (2006) Liposomal drug transport: a molecular perspective from molecular dynamics simulations in lipid bilayers. Adv Drug Deliv Rev 58:1357–1378
Yamamoto E, Akimoto T, Shimizu H, Hirano Y, Yasui M, Yasuoka K (2012) Diffusive nature of xenon anesthetic changes properties of a lipid Bilayer: molecular dynamics simulations. J Phys Chem B 116:8989–8995
Zhang L, Hung MC (1996) Sensitization of HER-2/neu-overexpressing non-small cell lung cancer cells to chemotherapeutic drugs by tyrosine kinase inhibitor emodin. Oncogene 12:571–576
Zhou XM, Chen QH (1988) Biochemical study of Chinese rhubarb. XXII. Inhibitory effect of anthraquinone derivatives on Na+−K+−ATPase of the rabbit renal medulla and their diuretic action. Yao Xue Xue Bao 23:17–20
Zubrzycki IZ, Xu Y, Madrid M, Tang P (2000) Molecular dynamics simulations of a fully hydrated dimyristoylphosphatidylcholine membrane in liquid-crystalline phase. J Chem Phys 112:3437–3441
Acknowledgements
This work was partially supported by CNPq, CAPES, FAPESP, INCT-FCx, NAP-FCx(USP) and BioMol (Brazil). Additionally, ARC acknowledges the fellowship from CNPq/CAPES, and HS, MTL and KC research fellowships from CNPq.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Antonio R. da Cunha declares that he has no conflicts of interest. Evandro L. Duarte declares that he has no conflicts of interest. Hubert Stassen declares that he has no conflicts of interest. M. Teresa Lamy declares that she has no conflicts of interest. Kaline Coutinho declares that she has no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
This article is part of a Special Issue on ‘Latin America’ edited by Pietro Ciancaglini and Rosangela Itri.
Electronic supplementary material
ESM 1
(DOCX 2370 kb)
Rights and permissions
About this article
Cite this article
da Cunha, A.R., Duarte, E.L., Stassen, H. et al. Experimental and theoretical studies of emodin interacting with a lipid bilayer of DMPC. Biophys Rev 9, 729–745 (2017). https://doi.org/10.1007/s12551-017-0323-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12551-017-0323-1