Abstract
Breeding for disease resistance or tolerance is a viable option for disease management programmes and is important for the continued success and resilience of planted forests. Red needle cast (RNC) is a disease that affects radiata pine (Pinus radiata) and is caused by Phytophthora pluvialis. Knowledge is still very limited regarding the potential for genetic tolerance to this pathogen. The application of controlled screening techniques is clearly required. Using a detached needle assay, we screened 392 clonally replicated individuals (clones) from an elite P. radiata population for quantitative tolerance to RNC. Data was highly skewed and required logarithmic data transformation and Poisson distributions for the estimation of best linear unbiased predictions. These estimates revealed a broad range in susceptibility/tolerance to RNC, and enabled the identification of clones that were clearly susceptible and clones that were clearly tolerant. There was a high correlation between the number and length of lesions that developed in response to inoculation with P. pluvialis. Broad-sense heritability estimates were low to moderate, indicating that there is potential for improving tolerance through breeding. These results provide evidence that breeding for tolerance to P. pluvialis is possible, although continued work into understanding and minimising causes for variance are required.
Similar content being viewed by others
Introduction
Phytophthora species have been recognised as foliar pathogens of radiata pine (Pinus radiata D. Don) since reports of Daño Foliar de Pino (DFP) in Chile in 2008, closely followed by observations of red needle cast (RNC) in New Zealand. Both P. kernoviae (Brasier et al. 2005) and P. pluvialis (Reeser et al. 2013) are known to be present in New Zealand, with the latter shown to be the primary cause of RNC (Dick et al. 2014). This disease is characterised by the development of olive lesions with resinous black bands on infected needles, which turn red and are prematurely cast. This defoliation has the potential to negatively affect growth and productivity (Ganley et al. 2014).
The selection of disease resistant/tolerant cultivars has long been a priority for breeding programmes (Bernatzky and Mulcahy 1992; Bridgwater 2005; Carson and Carson 1989; Devey et al. 2004; Sweeney 2013; Vivas et al. 2012). Resistance to other foliar diseases, such as dothistroma needle blight and cyclaneusma needle cast, has been extensively assessed and incorporated as selection criteria for several decades (Bulman 1993; Carson 1989; Wilcox 1982). In other forest tree species, breeding has also helped significantly in overcoming damage from disease, such as fusiform rust affecting southern pines and white pine blister rust in white pines, for which breeding efforts have been underway for more than 50 years (Bridgwater 2005; Sniezko et al. 2014).
In contrast, RNC has a shorter history (Dick et al. 2014) and information to guide selection programmes is still being collected. Despite being widespread across the majority of New Zealand (Fig. 1), outbreaks within genetic trials have been sporadic. Thus, opportunities to analyse the genetics of resistance or tolerance have been restricted, and further confounded by considerable variation in disease expression within and between sites and seasons. Likewise, reliable data around impacts on productivity are difficult to obtain, although annual incremental growth losses of 35% have been reported on some sites (Ganley et al. 2014). Field-based assessments are less informative in younger plantations due to limited availability of host material and opportunity for inoculum to build up. Consequently, RNC phenotypes are best assessed from 6 to 8 years of age when disease is more severe, which can delay the screening of new clonal and seedling material.
As a result, quantification of genetic variation of RNC tolerance has so far been restricted. Dungey et al. (2014) reported moderate heritabilities for percentage needle loss in an RNC-infected 6-year-old stand. While these results indicate that selection for tolerance could result in improved health, they require confirmation, which relies on information from further outbreaks in structured populations with extensive disease exposure within the trial. Until sufficient reliable evidence is collected regarding the susceptibility of lines within both production and breeding populations to enable accurate predictions of breeding values, incorporation of RNC tolerance as a selection trait in current breeding programmes will likely be limited, and potential genetic gains will not be fully realised.
In the absence of naturally occurring outbreaks in genetic trials, lab-based methods using a detached needle assay can offer an alternative means for screening for tolerance to RNC. Detached leaf assays have been employed in screening a number of different tree species for susceptibility to foliar Phytophthora species (Brasier et al. 2005; Denman et al. 2005; Dorrance and Inglis 1997; Hansen et al. 2005; Nyassé et al. 2002; Tahi et al. 2000), including P. pluvialis (Rolando et al. 2014). Such methods allow for a more controlled infection process with higher inoculum pressure, with subsequent incubation conditions that favour the development of lesions. The ability to screen targeted populations in a controlled manner provides a tool for breeders to understand the genetics of tolerance and the relative level of genetic gain that may be expected from tolerance-focused breeding programmes. This has practical application by enabling the targeted deployment of tolerant clones to disease-prone areas, and the identification of superior parents for further breeding.
The objective of this study was to apply in vitro screening of Pinus radiata for tolerance to RNC within a population of cloned elite germplasm to establish preliminary phenotypic data and expedite the realisation of genetic gain.
Materials and methods
Plant material—Pinus radiata clones
Plant material was sourced from a cloned elite population from the New Zealand Radiata Pine Breeding Company (RPBC). This population is representative of the third generation of the New Zealand P. radiata breeding programme elite germplasm (Dungey et al. 2009) and consisted of 63 full-sib families from 55 parents, with an average of 24 full-sibs per family. In this study, a total of 392 individuals (clones) were screened at an average of 6.2 clones per family.
Plants were clonally propagated from stool beds as bare-rooted cuttings, planted in Scion’s nursery in Rotorua, New Zealand. Five of these replicate plants (ramets) were tested for each clone that was screened. The initial two experiments each screened 24 clones, with subsequent experiments expanded to screen 48 clones. Four reference clones (RC 03_16, 15_07, 23_18, and 29_22) were included in every experiment to help assess between-experiment variations; these clones are part of the same cloned elite population and were chosen to represent a range of responses to RNC based on their observed performance in the first experiment. A total of 10 individual experiments were conducted over a period of 11 months (Jan. 2014–Nov. 2014) in order to screen all 392 clones. Additional rescreens of selected clones were performed to confirm reproducibility, giving a total of 13 experimental datasets.
Inoculum preparation
Zoospore inoculum was prepared as described previously by Rolando et al. (2014) with modifications to produce sufficient inoculum for the scale of the trial. Isolates of P. pluvialis were cultured on carrot agar at 17 °C for 3 days. Plugs of agar and mycelium were taken from the leading edge of the colonies, flooded with 55 ml of clarified carrot broth (Erwin and Ribeiro 1996) in vented 175 cm2 Nunc EasYFlasks (Thermo Scientific, Waltham, MA, USA) and incubated for 3 days in the dark at 17 °C. The resulting mycelial mats were rinsed thoroughly with a gentle stream of deionised water applied through a distribution manifold and run for 4 h. Thereafter, the deionised water was drained and 55 ml of sterile pond water was added. The pond water was collected from a local pond and autoclaved before being used to induce sporangia production and sporulation of Phytophthora pluvialis cultures. The flasks were re-incubated in the dark for a further 3 days before zoospore release was induced with 45 min intervals at 4 °C in the dark, then at room temperature (21–22 °C) on a light box. Zoospore concentrations were determined using a haemocytometer to ensure that a minimum of 5 × 103 zoospores per milliliter were applied in the assay. Zoospore suspensions were used within 2 h of preparation.
Experimental design
Each individual experiment was set up as an alpha design (balanced incomplete blocks) with five ramets per clone, applying the inoculation treatment (P. pluvialis-inoculated or water control) as a whole-plot factor and the clone identity as a sub-plot factor. The data set for a single experiment comprised 4800 observations, with 13 experiments performed in this study.
Detached needle assay
Twenty healthy fascicles (a bundle of needles held together by a sheath, usually containing two or three individual needles) were collected from each ramet (five ramets per clone). Ten fascicles were assigned at random to each treatment (P. pluvialis-inoculated and water control), with two independent sets (A and B) of tubes containing five fascicles each within each treatment. Each tube was inoculated overnight (18 h) with 4.5 ml of either a P. pluvialis zoospore suspension or sterile pond water. Thereafter, fascicles were removed from the tubes and placed on trays moistened with wet paper towels and incubated in a controlled environment (growth chamber set at 17 °C, with 65–70% relative humidity, and a 14-h photoperiod) for 10 days. Each tray contained a total of 12 independent sets of five fascicles each; within trays, treatments were not mixed to ensure that control-inoculated needles were not exposed to P. pluvialis zoospores during incubation. Positions of trays within the growth chamber were randomised. After 10 days, the needles within each fascicle were separated, and lesions counted and measured for length.
Statistical analysis
The data were structured as fascicles within ramets within clones within trays. Both response variables were analysed, with the total number of lesions and the average lesion length measured per fascicle within a set making up each experimental unit. Initial model development was performed using the using the lme4 R package (Bates et al. 2015; R Core Team 2017), while subsequent analyses were done in ASReml-R (Butler et al. 2009).
It was observed that the control treatments had a high proportion of zero lesions rendering their frequency distributions different from the frequency distributions of the infection treatments. On this basis, it was decided to exclude water treatments from the statistical analyses, with the estimates from the control treatment providing a baseline for lesion development without the introduction of the pathogen.
For lesion counts, a Poisson distribution was a natural candidate for the analysis. A Poisson distribution has a mean equal to its variance, which could result in over or underdispersion where the data does not match this feature. An alternative is the application of a negative binomial distribution which includes an extra parameter to deal with the discrepancy between the mean and the variance. In this study, using a Poisson distribution resulted in negligible overdispersion, while the negative binomial distribution had stability problems due to the long tail of the data. Given the experimental design, a Poisson distribution provided the most natural fit (Faraway 2006). As a Poisson formulation requires that the mean equals the variance, variations in best linear unbiased predictions (BLUPs) represent shifts in the locations of the random variables that have the same variance as the grand mean (Hilbe 2014). A generalised linear mixed model (GLMM) was initially used for the analysis as follows:
where β is a vector of fixed effects (experiment) with design matrix X, and b is a vector of random effects (clone, tray within clone, and fascicle within ramet) with design matrix Z, and f is the residual variance.
The average lesion length was a continuous variable with substantial positive skewness representing the high frequencies of large lesion lengths. A simple logarithmic transformation failed to address this problem satisfactorily, with the transformed data still retaining a notable skewness. The addition of a constant to each datum prior to logarithmic transformation was an option to address this (https://stats.stackexchange.com/users/919/whuber 2012). This constant was chosen to make the frequency distribution as symmetrical as possible according to the formula:
where the 10% quantile of the raw data is q10, the 90% quantile of the raw data is q90, and m is the mean of the raw data. From this, a constant of 50 was found to transform the raw data to an appropriate symmetry, as shown in Fig. 2.
The following mixed linear model, using the lme4 R package (Bates et al. 2015; R Core Team 2017), was initially adopted:
where y is a vector of lesion length measurements, β is a vector of all fixed effects (intercept and experiment) with design matrix X, and u is a vector of all random effects (clone, clone within tray, and fascicle within ramet) with design matrix Z, and e is the random residual effect.
The analyses determined that there was no significant effect on either response variable due to fascicles and ramet. This was established through fitting the full and reduced models by maximum likelihood rather than the restricted maximum likelihood approach (REML), and comparing the reduction in likelihood to an appropriate chi-squared distribution (Bates et al. 2015). The final models, therefore, consisted of experiment as a fixed effect, and clone and tray within clone as random effects.
Using these models, three analyses were conducted in ASReml-R (Butler et al. 2009): all experiments together in one model, selected experiments (rescreening experiments from 17 July, 14 August, and 5 June 2014, and 24 September 2015) in one model, and a model for each experiment. The same models were used to produce estimates of variance components and heritability across all experiments.
Clonal heritability and ranking of clones
Clonal (broad-sense) heritability (H2) was estimated from the variance components produced in the previously described model as follows:
where \( {\widehat{\sigma}}_u^2 \) is the clonal variance,\( {\widehat{\sigma}}_{ct}^2 \) is the tray within clone variance and \( {\widehat{\sigma}}_e^2 \) is the residual variance. The residual variance for the Poisson model was estimated following σ2e = φln(1/yp + 1), where φ is the overdispersion parameter, i.e. variance heterogeneity factor from the GLMM models, and yp is the mean of the lesion count (McKown et al. 2014).
Rankings of clones are presented as BLUP solutions produced from the previously described models.
Reference clone consistency
The consistency of lesion development on the four reference clones included in each experiment was explored further in a model containing experiment, inoculation treatment (H2O control, pathogen-inoculated), clone, and their interaction as fixed effects. The nested random term contained tray nested within ramet reflecting the alpha design.
The significance of the fixed terms was assessed using a backward selection procedure based on likelihood ratio tests (Zuur et al., 2009). Graphical model validation tools were used to test the underlying assumptions of variance homogeneity and normality (plots of standardised residuals vs. fitted values and against all explanatory variables to evaluate variance patterns, quantile-quantile plots to assess the normality criterion). The residual plots indicated strong heteroscedasticity and the variance pattern was modelled using a constant variance function (varIdent) using inoculation treatment as a grouping factor. The significant batch × treatment × clone interaction in the lesion length model for the reference clones was followed up applying a multiple comparison procedure using Tukey contrasts (R-package multcomp) (Hothorn et al. 2008).
Results
In total, 392 P. radiata clones were screened for tolerance to RNC using lesion development in a detached needle assay. There were significant interactions (likelihood ratio test, p < 0.01) between the experiment and the clones for all analyses. However, noting that there were a large number of clones tested and that there was no consistency in the relative rankings of these interactions between clones, this implied that these interactions should be treated as unexplained variation. This effectively put these interactions on the same footing as error terms, that is, biological elements that were not abstracted into the model. Thus, attention was directed to the mean clone response over all experiments while the interactions, though significant and with possible biological origins that were not experimentally controlled, were treated as noise.
Pathogen-inoculated samples showed substantially higher lesion development in comparison to the uninoculated controls. On average across all experiments, 51% of the inoculated fascicles developed observable lesions compared to 6% of the water control fascicles (Table 1). The control treatments showed negligibly small lesions induced during mock inoculation, indicating a physiological or biotic response to experimental conditions which was supported by a different frequency distribution to those of the infected treatments (data not shown). One experiment (27 February 2014) seemed to have a much higher percentage of non-zero lesion counts (35%) for the water controls; the reason for this is not clear. A baseline of 3–5% fascicles with lesions was observed across the remaining experiments in association with the expression of other pathogens, independent of inoculation with P. pluvialis.
We observed a very high correlation (r = 0.89; Spearman rank correlation) between the number of lesions and the average length of the lesions in a sample. This implies that the degree of infection might be measured by either the lesion count or the average lesion length equally well. However, the level of response observed for these two variables varied widely across experiments, with average lesion lengths and counts per experiment ranging from 0.44–21.50 and 0.31–2.81, respectively (Table 1). This highlights the need to transform the data to enable merging and analysis of data across multiple experiments.
The residual standard deviations of the average lesion length and the square roots of the deviations for the number of lesions, for each of the experiments after analysis, are shown in Tables 2 and 3, respectively. The results are very similar, and do not support a hypothesis that error variances differ between experiments. However, the deviations for the lesion counts for the water treatment show significantly lower values than that for the corresponding inoculation treatments in a number of cases. This may not be surprising, and could reflect bias when scoring needles from control trays that were not expected to have lesions.
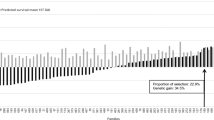
All clones were ranked according to their BLUPs estimated across all experiments for both lesion counts (Fig. 3) and lengths (Fig. 4). Clonal (broad-sense) heritability was moderate for average lesion length (H2 = 0.387 ± 0.02) but lower for lesion count (H2 = 0.239 ± 0.02) (Table 4). Where clones were tested in multiple experiments (5 June, 17 July, and 14 August 2014), greater variation was observed between replicate runs for those clones with highest rank susceptibility to infection (Fig. 5).
In terms of reference clone performance, 29_22 tended to display the longest lesions, followed by 23_18, and clones 15_07 and 03_16 tended to display shorter lesions. Figure 6 shows the average BLUPs for the four reference clones across all experiments, and supports the observation that 29_22 is the more susceptible and 15_07 the more tolerant reference clone.
While performance of the four reference clones in relation to each other was consistent when looking at the average lesion length across multiple experiments for reference clones, we noted seasonal variability (Fig. 7). These differences were significant (p < 0.02), with the level of infection, as measured by the lesions, increasing in the spring (late Aug/Sep) compared to the summer/autumn period (Apr-Jul) of New Zealand. Increases in lesion numbers and lengths are likely the result of the changing physiological state of both the host and pathogen isolates. While inoculations were carried out across the year, fresh isolates were only sourced during seasonal outbreaks in late winter and spring (July–Sep.). Between outbreaks, isolates gradually attenuated with successive culturing.
Average lesion length per fascicle in the four Pinus radiata reference clones in each experiment across a year, indicating the date of inoculation for each of the 13 experiments. Water control (upper panel), Phytophthora pluvialis infected needles (lower panel). Different lower case letters within an experiment indicate statistically significant differences at α = 0.05 (multiple comparison procedure using Tukey contrasts)
Discussion
Our results demonstrate a broad range in clonal susceptibility to RNC in this population, with evidence for some clones having either greater susceptibility or tolerance than the population average (Figs. 3 and 4). This study, therefore, gives support to the use of in vitro screening of radiata pine needles for breeding and selection of material with greater tolerance to infection by P. pluvialis. The plants tested here represent the third generation of RPBC elite germplasm for New Zealand, for which a limited amount of plant material was available for this study from previously established stool beds. No RNC tolerance data was yet available for this population. We present results from 392 clones across the 63 full-sib families of this cloned elite population, screened for tolerance to P. pluvialis using a detached needle assay.
Although it was useful to understand the baseline for lesion development not induced by the pathogen inoculation treatment, there does not appear to be further value in continually incorporating water control treatments. This exclusion of water controls would have the added benefit of essentially doubling screening capacity per experiment. Furthermore, significant experiment effects and insignificant ramet effects suggest greater value in reducing the number of ramets tested per clone in favour of increasing the total number of clones per experiment. Replicate screening runs could then address the need for replication within clones and address experimental effects.
Heritability
Of the two traits measured in this study, lesion length proved more useful with a greater ability to resolve differences between clones, ranging from 0 to 434 (total millimetres per fascicle) in the inoculated treatments, while lesion counts had a range from 0 to 40 (total per fascicle). Clonal heritability was moderate for lesion length (H2 = 0.387 ± 0.02), indicating that the genetic variation of tolerance to RNC can be detected using this trait in a detached needle assay. For lesion counts, clonal heritability was lower (H2 = 0.239 ± 0.02), rendering this trait potentially less useful for breeding or deployment selections. As the correlations between lesion length and lesion count are high (0.89), it may be possible to assess only the trait of higher broad-sense (clonal) heritability without losing important information on RNC tolerance in this population.
Our observations for clonal heritability are comparable to those previously reported for RNC in radiata pine. Dungey et al. (2014) showed narrow-sense heritability of RNC-associated needle loss in the field to be between 0.21 and 0.31 and broad-sense heritability between 0.23 and 0.59. Expression of other diseases on radiata pine in New Zealand show similar or slightly higher heritability estimates: dothistroma needle blight between 0.17 and 0.69 (Carson and Carson 1989; Ivković et al. 2010; Wilcox 1982), and cyclaneusma needle cast between 0.32 and 0.68 (Beets et al. 1997; King and Burdon 1991). However, estimates for these diseases have varied widely between trials, and are most consistently observed in the range of 0.1–0.4 (Dungey et al. 2006), which corresponds to the range we are reporting. In addition, heritability of disease resistance or tolerance is reportedly lower when assessed across sites (h2 = 0.07–0.11, H2 = 0.07–0.11) when compared to within-site estimates (h2 = 0.40–0.47, H2 = 0.40–0.55) (Kumar et al. 2008). The vast majority of studies in which tree species have been screened for resistance or tolerance to Phytophthora have shown clonal responses (Table 5), but many have not tracked or reported the heritability of this trait across a defined population. These studies have also focused primarily on root-infecting species of Phytophthora, with limited screening for resistance or tolerance to aerial Phytophthora species that would be directly comparable to needle infection of radiata pine with P. pluvialis. In the few studies which have reported heritability, estimates for Phytophthora resistance/tolerance have tended to be higher for shoot, stem, and root assays, and lower in studies that measured leaf lesions, but this depends greatly on the underlying mechanisms and genetics of resistance.
Experimental noise
In this study, we also noted a significant interaction between experiment and clone, which was likely to have biological origins that were not experimentally controlled, and was therefore treated as noise. This interaction was found to be inconsistent between clones, which is further supported by the variations in BLUP estimates for those clones that were tested in multiple experiments (Fig. 5). Some clones were more stable in their response to inoculation, regardless of the experiment in which they were tested, whereas some clones varied widely, more so towards the susceptible end of the rankings. Leaf age can affect disease susceptibility (Franich et al. 1986; Harris and Webber 2016); therefore, care was taken to select fully formed needles that best represent those observed to be the most affected in the field. Due to the time interval in screening all 392 clones, however, the age of the material and progressive crowding of plants in the nursery beds differed from the first to the final experiment. In addition, seasonal effects could be further affecting host physiology and culture attenuation of the pathogen, as has been reported in other Phytophthora studies (Dodd et al. 2005; Harris and Webber 2016). This issue of isolate attenuation was addressed where possible by using composite inoculum suspensions of a range of seasonal isolates in each inoculation run. The availability of fresh isolates was thus swayed by the natural expression of disease in the field, and inability to recover isolates consistently across the year. Prolonged culturing in non-host-based axenic media can result in attenuation of isolate aggressiveness, as has been shown in P. infestans (Caten 1971; Langcake 1974). Periodic passaging through radiata pine needles did mitigate this (data not shown), but the availability of fresh strains from naturally occurring infections in late winter/early spring coincided with notable improvements in infection levels, giving support to the importance of using fresh field isolates (Fig. 7).
Limitations of detached assays
Although detached leaf and needle assays have been used broadly in host testing with Phytophthora species (Hüberli et al. 2008), their application must take into account the senescence of the plant material having been removed from the plant. With such removal of plant material, many key metabolic and defence responses normally present in the plant are likely to be compromised. In addition, the detachment of the needle provides a wound point through which infection may be initiated. This was not observed to be significant in this study or other studies, with most of the infections focussed along the needle shaft and at the meniscus point of the inoculum rather than via the wound point, due to gravitaxis and chemotaxis (Burgess et al. 1998; Häder 1999).
All methods have inherent advantages and disadvantages that should be considered when choosing a method for screening for disease resistance/tolerance. If the goal is the identification of resistant/tolerant trees for breeding programmes, then the inoculation method must ensure that all trees are exposed, so that a susceptible tree is not accidentally identified as resistant/tolerant. Conditions that favour the development of disease, i.e., availability of susceptible tissue, consistent inoculum density, and favourable incubation conditions, minimise the potential for disease escape of susceptible clones (Schmidt 1972). The use of detached needle assays allows for such controlled inoculations, with high inoculum pressure and incubation conditions which favour the development of lesions. The ability to detect a quantitative range of responses to infection, as shown across the 13 experiments presented here, demonstrates the utility of the assay in identifying clones with greater relative tolerance. However, we recognise that the assay conditions could result in overestimation of susceptibility. This is supported by some preliminary work (unpublished data), comparing parallel results from live plant and detached needle inoculations. Similar outcomes have been observed when comparing artificial inoculation systems for other pine diseases (Schmidt 1972). The detached needle assay may be less useful at differentiating more subtle differences in susceptibility between clones, such as those arising from multigenic effects. This could partially explain why we are observing heritabilities in this study that are at the lower end of those previously reported from field observations (Dungey et al. 2014). However, the detached needle assay remains useful for primary screening of a broad range of material not yet exposed to RNC in the field and identification of tolerant candidates for further investigation. In addition, it remains quite possible that, in spite of observing a broad range of tolerance in this study, we have not yet captured the full range of responses possible for radiata pine as the clones tested represent only a portion of the entire breeding programme. In other pine species, detecting resistant/tolerant phenotypes often requires the screening of thousands of individuals (Sniezko et al. 2014). The clones screened in this study have now been established by the RPBC in regional clonal trials across New Zealand, including some RNC-prone sites, which will hopefully present opportunities to validate the results in this study with field-based observations of resistance.
Implications for field tolerance
Further work is still needed to better understand the actual mechanisms that contribute to tolerance in the field. This study measured lesion length and number, which reflect the early stages of infection and disease, but it is unclear how this will correlate to field tolerance where the end stage of disease, i.e. casting of needles, is what is measured, and is ultimately the phenotype that industry are interested in due to the impact on productivity. Lesions may reflect secondary host responses rather than indicate the extent of infection, and other measures of pathogen growth and establishment within the host could potentially be more informative (Harris and Webber 2016). Non-systematic microscopy observations made throughout this study (data not shown) indicate that the level of sporangia production varies considerably across host genotypes. Preliminary results contrasting more tolerant and more susceptible clones identified in this study have indicated that pathogen spore production within the host could be a key factor in susceptibility, with more sporangia observed in highly susceptible clones. Sporulating hosts are also likely to be drivers of disease in natural outbreaks, as they contribute to the build-up of inoculum in the forest and impact the spread of disease (Carson et al. 1991; Harris and Webber 2016). The potential for utilising sporulation as a trait in breeding value assessments for RNC susceptibility is currently under investigation.
This is the first demonstration of a genetic and thus heritable basis for RNC tolerance using direct inoculation of P. pluvialis on detached needles with quantification of infection. This tolerance shows evidence of consistency in differentiating genotype performance between experiments, and future work will seek to account for seasonality and other environmental variables where possible. Broad-sense heritability estimates from this study support previous work in understanding the genetics of RNC tolerance in radiata pine (Dungey et al. 2014) and shows that breeding has the potential to improve tree health in relation to RNC.
References
Bates D, Mächler M, Bolker B, Walker S (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67:48. https://doi.org/10.18637/jss.v067.i01
Beets P, Oliver G, Kimberley M (1997) Genotypic variation in symptoms of upper mid-crown yellowing and Cyclaneusma minus in a Pinus radiata stand. N Z J For Sci 27:69–75
Bernatzky R, Mulcahy DL (1992) Marker-aided selection in a backcross breeding program for resistance to chestnut blight in the American chestnut. Can J For Res 22:1031–1035
Brasier CM, Beales PA, Kirk SA, Denman S, Rose J (2005) Phytophthora kernoviae sp. nov., an invasive pathogen causing bleeding stem lesions on forest trees and foliar necrosis of ornamentals in the UK. Mycol Res 109:853–859
Bridgwater FKTBTMS (2005) Risk assessment with current deployment strategies for fusiform rust-resistant loblolly and slash pines. South J Appl For 29:80–87
Bryan WC (1965) Testing shortleaf pine seedlings for resistance to infection by Phytophthora cinnamomi vol 50. Southeastern Forest Experiment Station, US Department of Agriculture, Forest Service
Bulman L (1993) Cyclaneusma needle-cast and Dothistroma needle blight in NZ pine plantations. New Zealand Forestry 38:21–24
Burgess T, McComb J, Hardy G, Colquhoun I (1998) Influence of low oxygen levels in aeroponics chambers on eucalypt roots infected with Phytophthora cinnamomi. Plant Dis 82:368–373
Butcher TB, Stukely MJC, Chester GW (1984) Genetic variation in resistance of Pinus radiata to Phytophthora cinnamomi. For Ecol Manag 8:197–220. https://doi.org/10.1016/0378-1127(84)90053-7
Butler D, Cullis BR, Gilmour A, Gogel B (2009) ASReml-R reference manual. The State of Queensland, Department of Primary Industries and Fisheries, Brisbane
Carson MJ, Carson SD (1989) Breeding for resistance in forest trees: a quantitative genetic approach. Annual Rev Phytopathlogy 27:375–395
Carson S (1989) Selecting Pinus radiata for resistance to Dothistroma needle blight. N Z J For Sci 19:3–21
Carson S, Dick A, West G (1991) Benefits of the Dothistroma-resistant breed of radiata pine. New directions in forestry costs and benefits of change. JC Allen and AGD Whyte, eds Aust & NZ Inst Forestry, Christchurch, New Zealand:251–262
Caten C (1971) Single zoospore variation in Phytophthora infestans and attenuation of strains in culture. Trans Br Mycol Soc 56:1–7
Chandelier A, Husson C, Druart P, Marçais B (2016) Assessment of inoculation methods for screening black alder resistance to Phytophthora ×alni. Plant Pathol 65:441–450. https://doi.org/10.1111/ppa.12418
Denman S, Kirk SA, Brasier CM, Webber JF (2005) In vitro leaf inoculation studies as an indication of tree foliage susceptibility to Phytophthora ramorum in the UK. Plant Pathol 54:512–521. https://doi.org/10.1111/j.1365-3059.2005.01243.x
Devey ME, Groom KA, Nolan MF, Bell JC, Dudzinski MJ, Old KM, Matheson AC, Moran GF (2004) Detection and verification of quantitative trait loci for resistance to Dothistroma needle blight in Pinus radiata. TAG Theor Appl Genet 108:1056–1063
Díaz R, Fernández-López J (2005) Genetic variation at early ages for several traits of interest for timber-production breeding of Juglans regia. Can J For Res 35:235–243. https://doi.org/10.1139/x04-162
Dick M, Williams N, Bader M, Gardner J, Bulman L (2014) Pathogenicity of Phytophthora pluvialis to Pinus radiata and its relation with red needle cast disease in New Zealand. N Z J For Sci 44:1–12. https://doi.org/10.1186/s40490-014-0006-7
Dodd RS, Hüberli D, Douhovnikoff V, Harnik TY, Afzal-Rafii Z, Garbelotto M (2005) Is variation in susceptibility to Phytophthora ramorum correlated with population genetic structure in coast live oak (Quercus agrifolia)? New Phytol 165:203–214. https://doi.org/10.1111/j.1469-8137.2004.01200.x
Dorrance AE, Inglis DA (1997) Assessment of greenhouse and laboratory screening methods for evaluating potato foliage for resistance to late blight. Plant Dis 81:1206–1213. https://doi.org/10.1094/PDIS.1997.81.10.1206
Douhan GW, Fuller E, McKee B, Pond E (2011) Genetic diversity analysis of avocado (Persea americana miller) rootstocks selected under greenhouse conditions for tolerance to phytophthora root rot caused by Phytophthora cinnamomi. Euphytica 182:209–217. https://doi.org/10.1007/s10681-011-0433-y
Dungey H, Brawner J, Burger F, Carson M, Henson M, Jefferson P, Matheson A (2009) A new breeding strategy for Pinus radiata in New Zealand and New South Wales. Silvae Genetica 58:28–38
Dungey H, Low C, Bulman L Needle cast in New Zealand–are there opportunities for improvement in plantation growth. In: Breeding for success: diversity in action’, Proceedings of the 13th Australasian Plant Breeding Conference, Ed CF Mercer Christchurch, 2006. pp 497–504
Dungey H, Williams N, Low C, Stovold G (2014) First evidence of genetic-based tolerance to red needle cast caused by Phytophthora pluvialis in radiata pine. N Z J For Sci 44:31
Erwin DC, Ribeiro O (1996) Phytophthora Diseases Worldwide. APS Press, Minnesota
Faraway J (2006) Extending the linear model with R. Chapman Hall. CRC, Boca Raton
Frampton J, Isik F, Benson DM (2013) Genetic variation in resistance to Phytophthora cinnamomi in seedlings of two Turkish Abies species. Tree Genetics Genomes 9:53–63
Franich RA, Carson MJ, Carson SD (1986) Synthesis and accumulation of benzoic acid in Pinus radiata needles in response to tissue injury by dothistromin, and correlation with resistance of P. radiata families to Dothistroma pini. Physiol Mol Plant Pathol 28:267–286
Ganley RJ, Williams NM, Rolando CA, Hood IA, Dungey HS, Beets PN, Bulman LS (2014) Management of red needle cast, caused by Phytophthora pluvialis, a new disease of radiata pine in New Zealand. New Zealand Plant Protection 67:48–53
Green S, Brasier CM, Schlenzig A, McCracken A, MacAskill GA, Wilson M, Webber JF (2013) The destructive invasive pathogen Phytophthora lateralis found on Chamaecyparis lawsoniana across the UK. For Pathol 43:19–28. https://doi.org/10.1111/j.1439-0329.2012.00788.x
Häder D-P (1999) Gravitaxis in unicellular microorganisms. Adv Space Res 24:843–850
Hansen E, Hamm PB, Roth LF (1989) Testing port-Orford-cedar for resistance to phytophthora. Plant Dis 73:791–794
Hansen EM, Parke JL, Sutton W (2005) Susceptibility of Oregon forest trees and shrubs to Phytophthora ramorum: a comparison of artificial inoculation and natural infection. Plant Dis 89:63–70. https://doi.org/10.1094/pd-89-0063
Harris A, Webber J (2016) Sporulation potential, symptom expression and detection of Phytophthora ramorum on larch needles and other foliar hosts. Plant Pathol 65:1441–1451
Hayden KJ (2010) Variation and heritability of host susceptibility to the introduced pathogen Phytophthora ramorum in tanoak (Lithocarpus densiflorus, proposed name Notholithocarpus densiflorus) populations. University of California, Berkeley
Hilbe JM (2014) Modeling count data. Cambridge University Press, Cambridge. https://doi.org/10.1017/CBO9781139236065
Hoover BK, Bates R (2013) Susceptibility of Fraser, Canaan, and nordmann fir to root rot incited by Phytophthora cactorum and Phytophthora drechsleri. HortTechnology 23:44–50
Hothorn T, Bretz F, Westfall P (2008) Simultaneous inference in general parametric models. Biometrical J Biometrische Zeitschrift 50:346–363. https://doi.org/10.1002/bimj.200810425
https://stats.stackexchange.com/users/919/whuber (2012) Choosing c such that log(x + c) would remove skew from the population. https://stats.stackexchange.com/q/41415 (version: 2012-11-05). 2018
Hüberli D, Lutzy B, Voss B, Calver M, Ormsby M, Garbelotto M (2008) Susceptibility of New Zealand flora to Phytophthora ramorum and pathogen sporulation potential: an approach based on the precautionary principle. Australas Plant Pathol 37:615–625
Ivković M, Baltunis B, Gapare W, Sasse J, Dutkowski G, Elms S, Wu H (2010) Breeding against Dothistroma needle blight of radiata pine in Australia. Can J For Res 40:1653–1660
King JN, Burdon RD (1991) Time trends in inheritance and projected efficiencies of early selection in a large 17-year-old progency test of Pinus radiata. Can J For Res 21:1200–1207
Kumar S, Burdon R, Stovold G, Gea L (2008) Implications of selection history on genetic architecture of growth, form, and wood-quality traits in Pinus radiata. Can J For Res 38:2372–2381
Langcake P (1974) Sterols in potato leaves and their effects on growth and sporulation of Phytophthora infestans. Trans Br Mycol Soc 63:573–586
McKown AD, Guy RD, Quamme L, Klápště J, La Mantia J, Constabel CP, El-Kassaby YA, Hamelin RC, Zifkin M, MS Azam (2014) Association genetics, geography and ecophysiology link stomatal patterning in with carbon gain and disease resistance trade-offs. Molecular Ecology 23(23):5771–5790
McWilliams MG (2000) Port-Orford-cedar and Phytophthora lateralis: grafting and heritability of resistance in the host, and variation in the pathogen. Oregon State University
Nyassé S, Despréaux D, Cilas C (2002) Validity of a leaf inoculation test to assess the resistance to Phytophthora megakarya in a cocoa (Theobroma cacao L.) diallel mating design. Euphytica 123:395–399
Ploetz R, Schnell RJ, Haynes J (2002) Variable response of open-pollinated seedling progeny of avocado to Phytophthora root rot. Phytoparasitica 30:262–268
R Core Team (2017) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org
Reeser PW, Sutton W, Hansen EM (2013) Phytophthora pluvialis, a new species found in mixed tanoak–Douglas-fir forests of western Oregon, U.S.A. North American Fungi 8 https://doi.org/10.2509/naf2013.008.007
Robin C, Brasier C, Reeser P, Sutton W, Vannini A, Vettraino AM, Hansen E (2014) Pathogenicity of Phytophthora lateralis lineages on different selections of Chamaecyparis lawsoniana. Plant Dis 99:1133–1139. https://doi.org/10.1094/PDIS-07-14-0720-RE
Robin C, Morel O, Vettraino A-M, Perlerou C, Diamandis S, Vannini A (2006) Genetic variation in susceptibility to Phytophthora cambivora in European chestnut (Castanea sativa). For Ecol Manag 226:199–207
Rolando C, Gaskin R, Horgan D, Williams N, Bader MK-F (2014) The use of adjuvants to improve uptake of phosphorous acid applied to Pinus radiata needles for control of foliar Phytophthora diseases. N Z J For Sci 44:1–7. https://doi.org/10.1186/s40490-014-0008-5
Santos C, Machado H, Correia I, Gomes F, Gomes-Laranjo J, Costa R (2015) Phenotyping Castanea hybrids for Phytophthora cinnamomi resistance. Plant Pathol 64:901–910. https://doi.org/10.1111/ppa.12313
Schmidt RA (1972) A literature review of inoculation techniques used in studies of fusiform rust. Biology of Rust Resistance in Forest Trees 1221:341–356
Sniezko R, Smith J, Liu J-J, Hamelin R (2014) Genetic resistance to fusiform rust in southern pines and white pine blister rust in white pines—a contrasting tale of two rust Pathosystems—current status and future prospects. Forests 5:2050–2083
Stukely M, Crane C (1994) Genetically based resistance of Eucalyptus marginata to Phytophthora cinnamomi. Phytopathology 84:650–656
Sweeney K (2013) Histological comparisons of needle tissues of four species of white pine infected by Cronartium ribicola. Oregon State University
Tahi M, Kebe I, Eskes AB, Ouattara S, Sangare A, Mondeil F (2000) Rapid screening of cacao genotypes for field resistance to Phytophthora palmivora using leaves, twigs and roots. Eur J Plant Pathol 106:87–94. https://doi.org/10.1023/a:1008747800191
Utkhede R, Quamme H (1988) Use of the excised shoot assay to evaluate resistance to Phytophthora cactorum of apple rootstock cultivars. Can J Plant Sci 68:851–857
Vivas M, Zas R, Solla A (2012) Screening of maritime pine (Pinus pinaster) for resistance to Fusarium circinatum, the causal agent of pitch canker disease. Forestry 85:185–192
Wilcox MD (1982) Genetic variation and inheritance of resistance to Dothistroma needle blight in Pinus radiata. N Z J For Sci 12:14–35
Zuur AFIE, Walker NJ, Saveliev AA, Smith GM (2009) Mixed effects models and extensions in ecology with R (2009) mixed effects models and extensions in ecology with R. Springer, New York
Acknowledgements
Pinus radiata clonal material was sourced from the Radiata Pine Breeding Company. Thank you to Alvin Yanchuk for his contributions to experimental design. The authors also wish to acknowledge the invaluable contributions of the team who assisted with inoculum production, needle collections and inoculations, lesion assessments and data capture.
Data archiving statement
The phenotypic data used for the analysis in this study will be submitted to TreeGenes (https://treegenesdb.org) upon acceptance of the manuscript.
Funding
This work was funded by the New Zealand Ministry of Business, Innovation and Employment (CO4X1305), Scion Core funding, the Forest Growers Levy Trust (Administered by the Forest Owners Association) and the Radiata Pine Breeding Company (RPBC1001).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. Beaulieu
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Graham, N.J., Suontama, M., Pleasants, T. et al. Assessing the genetic variation of tolerance to red needle cast in a Pinus radiata breeding population. Tree Genetics & Genomes 14, 55 (2018). https://doi.org/10.1007/s11295-018-1266-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-018-1266-9