Abstract
Genetic testing of an Irish kindred identified an exonic nucleotide substitution c.1664T>C (p.Leu555Pro) in the MLH1 mismatch repair (MMR) gene. This previously unreported variant is classified as a “variant of uncertain significance” (VUS). Immunohistochemical (IHC) analysis and microsatellite instability (MSI) studies, genetic testing, a literature and online MMR mutation database review, in silico phenotype prediction tools, and an in vitro MMR activity assay were used to study the clinical significance of this variant. The MLH1 c.1664T>C (p.Leu555Pro) VUS co-segregated with three cases of classic Lynch syndrome-associated malignancies over two generations, with consistent loss of MLH1 and PMS2 protein expression on IHC, and evidence of the MSI-High mutator phenotype. The leucine at position 555 is well conserved across a number of species, and this novel variant has not been reported as a normal polymorphism in the general population. In silico and in vitro analyses suggest that this variant may have a deleterious effect on the MLH1 protein and abrogate MMR activity. Evidence from clinical, histological, immunohistochemical, and molecular genetic data suggests that MLH1 c.1664T>C (p.Leu555Pro) is likely to be the pathogenic cause of Lynch syndrome in this family.
Similar content being viewed by others
References
Lynch HT, de la Chapelle A (2003) Hereditary colorectal cancer. N Engl J Med 348:919–932
Kovacs ME, Papp J, Szentirmay Z et al (2009) Deletions removing the last exon of TACSTD1 constitute a distinct class of mutations predisposing to Lynch Syndrome. Hum Mutat 30(2):197–203
Ligtenberg MJL, Kuiper RP, Chan TL et al (2009) Heritable somatic methylation and inactivation of MSH2 in families with Lynch syndrome due to deletion of the 3′ exons of TACSTD1. Nat Genet 41(1):114–117
Crépin M, Dieu M-C, Lejeune S et al (2012) Evidence of constitutional MLH1 epimutation associated to transgenerational inheritance of cancer susceptibility. Hum Mutat 33:180–188
Engel C, Forberg J, Holinski-Feder E et al (2006) Novel strategy for optimal sequential application of clinical criteria, immunohistochemistry and microsatellite analysis in the diagnosis of hereditary nonpolyposis colorectal cancer. Int J Cancer 118:115–122
Peltomaki P, Vasen HF (2004) Mutations associated with HNPCC predisposition—update of ICG-HNPCC/InSiGHT mutation database. Dis Markers 20:269–276
Rasmussen LJ, Heinen CD, Royer-Pokora B, Drost M, Tavtigian S, Hofstra RM, de Wind N (2012) Pathological assessment of mismatch repair gene variants in Lynch syndrome: past, present and future. Hum Mutat 33(12):1617–1625
Plon SE, Eccles DM, Easton D et al (2008) Sequence variant classification and reporting: recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum Mutat 29(11):1282–1291
Goldgar DE, Easton DF, Byrnes GB et al (2008) Genetic evidence and integration of various data sources for classifying uncertain variants into a single model. Hum Mutat 29:1282–1291
Vasen HF, Mecklin JP, Khan PM, Lynch HT (1991) The international collaborative group on hereditary non-polyposis colorectal cancer (ICG-HNPCC). Dis Colon Rectum 34:424–425
Woods MO, Williams P, Careen A et al (2007) A new variant database for mismatch repair genes associated with Lynch Syndrome. Hum Mutat 28(7):669–673
Ou J, Niessen RC, Vonk J et al (2008) A database to support the interpretation of human mismatch repair gene variants. Hum Mutat 29(11):1337–1341
Drost M, Zonneveld JBM, van Dijk L et al (2010) A cell-free assay for the functional analysis of variants of the mismatch repair protein MLH1. Hum Mutat 31:247–253
Domingo E, Laiho P, Ollikainen M et al (2004) BRAF screening as a low-cost effective strategy for simplifying HNPCC genetic testing. J Med Genet 41:664–668
Boland CR, Fishel R (2005) Lynch syndrome: form, function, proteins, and basketball. Gastroenterology 129(2):751–755
Spurdle AB (2013) Department of Genetics and Computational Biology, Queensland Institute of Medical Research, Brisbane, Australia (personal communication)
Hofstra RMW, Spurdle AB, Eccles D et al (2008) Tumor characteristics as an analytic tool for classifying genetic variants of uncertain clinical significance. Hum Mutat 29(11):1292–1303
Cleary SP, Cotterchio M, Jenkins MA et al (2009) Germline MutY human homologue mutations and colorectal cancer: a multisite case-control study. Gastroenterology 136:1251–1260
Boparai KS, Dekker E, Van Eeden S et al (2008) Hyperplastic polyps and sessile serrated adenomas as a phenotypic expression of MYH-associated polyposis. Gastroenterology 135:2014–2018
Strachan T, Read AP (1999) Human molecular genetics. Wiley, New York
Westphalen AA, Russell AM, Buser M et al (2005) Evidence for genetic anticipation in hereditary non-polyposis colorectal cancer. Hum Genet 116:461–465
Stella A, Surdo NC, Lastella P et al (2007) Germline novel MSH2 deletions and a founder MSH2 deletion associated with anticipation effects in HNPCC. Clin Genet 71:130–139
Nilbert M, Timshel S, Bernstein I, Larsen K (2009) Role for genetic anticipation in Lynch Syndrome. J Clin Oncol 27:360–364
Spurdle AB, Thompson BA (2013) Department of Genetics and Computational Biology, Queensland Institute of Medical Research, Brisbane, Australia (personal communication)
Plotz G, Raedle J, Spina A et al (2008) Evaluation of the MLH1 I219V alteration in DNA mismatch repair activity and ulcerative colitis. Inflamm Bowel Dis 14(5):605–611
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix 1
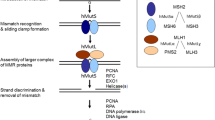
See Fig. 1.
Appendix 2
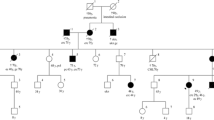
See Fig. 2.
Appendix 3
See Fig. 3.
MMR activity of MLH1 VUS as measured in the in vitro MMR assay. Results are shown as mean ± SEM of 4 fully independent experiments. Mock: Mock expression. I219V represents an innocuous polymorphism [25]. Asterisks Significantly higher than repair-deficient control G67R. (p < 0.001, student’s one-tailed t test). For “Mock” and “PMS2 only” reactions no repair was detected in all experiments
Rights and permissions
About this article
Cite this article
Farrell, M.P., Hughes, D.J., Drost, M. et al. Multivariate analysis of MLH1 c.1664T>C (p.Leu555Pro) mismatch repair gene variant demonstrates its pathogenicity. Familial Cancer 12, 741–747 (2013). https://doi.org/10.1007/s10689-013-9652-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10689-013-9652-9