Abstract
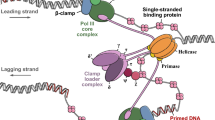
Restriction-modification systems protect bacteria from foreign DNA. Type I restriction-modification enzymes are multifunctional heteromeric complexes with DNA-cleavage and ATP-dependent DNA translocation activities located on endonuclease/motor subunit HsdR. The recent structure of the first intact motor subunit of the type I restriction enzyme from plasmid EcoR124I suggested a mechanism by which stalled translocation triggers DNA cleavage via a lysine residue on the endonuclease domain that contacts ATP bound between the two helicase domains. In the present work, molecular dynamics simulations are used to explore this proposal. Molecular dynamics simulations suggest that the Lys–ATP contact alternates with a contact with a nearby loop housing the conserved QxxxY motif that had been implicated in DNA cleavage. This model is tested here using in vivo and in vitro experiments. The results indicate how local interactions are transduced to domain motions within the endonuclease/motor subunit.







Similar content being viewed by others
References
Murray NE (2002) Immigration control of DNA in bacteria: self versus non-self. Microbiology 148:3–20
Lapkouski M, Panjikar S, Janscak P, Smatanova IK, Carey J, Ettrich R, Csefalvay E (2009) Structure of the motor subunit of type I restriction-modification complex EcoR124I. Nat Struct Mol Biol 16:94–105
Gorbalenya AE, Koonin EV (1991) Endonuclease (R) subunits of type-I and type-III restriction–modification enzymes contain a helicase-like domain. FEBS Lett 291:277–281
Obarska-Kosinska A, Taylor JE, Callow P, Orlowski J, Bujnicki JM, Kneale GG (2008) HsdR subunit of the type I restriction-modification enzyme EcoR124I: biophysical characterisation and structural modelling. J Mol Biol 376(2):438–452
Sisakova E, Stanley LK, Weiserova M, Szczelkun MD (2008) A RecB-family nuclease motif in type I restriction endonuclease EcoR124I. Nucleic Acids Res 36:3939–3949
Niv MY, Ripoll DR, Vila JA, Liwo A, Vanamee ES, Aggarwal AK, Weinstein H, Scheraga HA (2007) Topology of type II REases revisited; structural classes and the common conserved core. Nucleic Acids Res 35:2227–2237
Krieger E, Koraimann G, Vriend G (2002) Increasing the precision of comparative models with YASARA NOVA; a self-parameterizing force field. Proteins 47:393–402
Konagurthu AS, Whisstock JC, Stuckey PJ, Lesk AM (2006) MUSTANG: a multiple structural alignment algorithm. Proteins 64:559–574
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJC (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718
Berendsen HJC, van der Spoel D, van Drunen R (1995) GROMACS: a message-passing parallel molecular dynamics implementation. Comp Phys Comm 91:43–56
Pronk S, Pall S, Schulz R, Larsson P, Bjelkmar P, Apostolov R, Shirts MR, Smith JC, Kasson PM, van der Spoel D, Hess B, Lindahl E (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29(7):845–854
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C (2006) Comparison of multiple AMBER force fields and development of improved protein backbone parameters. Proteins 65:712–725
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general AMBER force field. J Comput Chem 25:1157–1174
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Pople JA et al. (2004) GAUSSIAN 03 (revision C.02). Gaussian, Inc., Wallingford
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926
Darden T, York D, Pedersen L, Ewald P (1993) An N·log(N) method for Ewald sums in large systems. J Chem Phys 98(12):10089–10092
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comp Chem 18(12):1463–1472
Bussi G, Donadio D, Parrinello M (2007) Canonical sampling through velocity rescaling. J Chem Phys 126:014101
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182
Hayward S, Kitao A, Berendsen HJC (1997) Model-free methods of analyzing domain motions in proteins from simulation: a comparison of normal mode analysis and molecular dynamics simulation of lysozyme. Proteins 27:425–437
Hayward S, Berendsen HJC (1998) Systematic analysis of domain motions in proteins from conformational change; new results on citrate synthase and T4 lysozyme. Proteins 30:144–154
Amadei A, Linnssen AB, Berendsen HJ (1993) Essential dynamics of proteins. Proteins 17:412–25
Schrödinger LLC (2011) QSite 5.7. Schrödinger LLC, New York
Grimme S, Antony J, Ehrlich S, Krieg H (2010) A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H–Pu. J Chem Phys 132:154104–154123
Boys SF, Bernardi F (1970) Calculation of small molecular interactions by differences of separate total energies—some procedures with reduced errors. Molec Phys 19:553–556
Schrödinger LLC (2011) Impact 5.7. Schrödinger LLC, New York
Jorgensen WL, Tirado-Rives J (2005) Potential energy functions for atomic-level simulations of water and organic and biomolecular systems. Proc Natl Acad Sci USA 102:6665–6670
Humphrey W, Dalke A, Schulten K (1996) VMD—visual molecular dynamics. J Mol Graph 14:33–38
Janscak P, Abadjieva A, Firman K (1996) The type I restriction endonuclease R.EcoR124I: over-production and biochemical properties. J Mol Biol 257(5):977–991
Holubova I, Vejsadová Š, Firman K, Weiserova M (2004) Cellular localization of type I restriction-modification enzymes is family dependent. Biochem Biophys Res Commun 319:375–380
Patel J, Taylor I, Dutta CF, Kneale G, Firman K (1992) High-level expression of the cloned genes encoding the subunits of and intact DNA methylase, M.EcoR124. Gene 112:21–27
Taylor I, Patel J, Firman K, Kneale GG (1992) Purification and biochemical characterization of the EcoR124 type I modification methylase. Nucleic Acids Res 20:179–186
Jacob F, Wollman EL (1954) Etude génétique d’un bactériophage tempéré d’Escherichia coli. III. Effet du rayonnement ultraviolet sur la recombinaison génétique. Ann Inst Pasteur 87:653–673
Colson C, Glover SW, Symons N, Stanley KA (1965) The location of the genes for host-controlled modification and restriction in Escherichia coli K-12. Genetics 52:1043–1050
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33:103–119
Abramoff MD, Magalhaes PJ, Ram SJ (2004) Image processing with ImageJ. Biophoton Int 11(7):36–42
Chan KM, Delfert D, Junger KD (1986) A direct calorimetric assay for Ca2+ stimulated ATPase activity. Anal Biochem 157:375–380
Davies GP, Martin I, Sturrock SS, Cronshaw A, Murray NE, Dryden DTF (1999) On the structure and operation of type I DNA restriction enzymes. J Mol Biol 290:565–579
Uyen NT, Park S, Choi J, Lee HJ, Nishi K, Kim JS (2009) The fragment structure of a putative HsdR subunit of a type I restriction enzyme from Vibrio vulnificus YJ016: implications for DNA restriction and translocation activity. Nucleic Acids Res 37:6960–6969
Sisakova E, Weiserova M, Dekker C, Seidel R, Szczelkun MD (2008) The interrelationship of helicase and nuclease domains during DNA translocation by the molecular motor EcoR124I. J Mol Biol 384:1273–1286
Studier FW, Bandyopadhyay PK (1988) Model for how type I restriction enzymes select cleavage sites in DNA. Proc Natl Acad Sci USA 85:4677–4681
Acknowledgments
We gratefully acknowledge support from the Czech Science Foundation (P207/12/2323 to RE and MW), the institutional research project RVO 61388971, and joint Czech–US National Science Foundation international research cooperation (DBI10-04830).
Author information
Authors and Affiliations
Corresponding author
Additional information
Dhiraj Sinha and Katsiaryna Shamayeva contributed equally.
This paper belongs to Topical Collection MIB 2013 (Modeling Interactions in Biomolecules VI)
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplemental Fig. 1a–c
RMSD relative to the starting structure. The root-mean-square deviation (RMSD, Å) of Cα positions is derived every 1 ps during the trajectory by overlaying each simulation structure with its corresponding initial structure by Cα superposition. a WT HsdR. b Restrained WT. c Arg182Ala mutant. (GIF 30 kb)
Supplemental Fig. 2a–c
Root-mean-square fluctuation (RMSF, Å) calculated during the last 20 ns of each simulation for each Cα atom is plotted vs. residue number. a WT HsdR. b Restrained WT. c Arg182Ala mutant. (GIF 45 kb)
Supplemental Fig. 3a–c
Root-mean-square fluctuation (RMSF, Å) calculated during the last 20 ns of each simulation for each Cα atom of the endonuclease domain is plotted vs. residue number. a WT HsdR. b Restrained WT. c Arg182Ala mutant. The secondary structures assigned in the PDB file by DSSP are shown under each panel, with red bars representing helical segments, yellow bars representing strand segments, and gray bars representing irregular segments. (GIF 43 kb)
ESM
(PDF 20 kb)
Rights and permissions
About this article
Cite this article
Sinha, D., Shamayeva, K., Ramasubramani, V. et al. Interdomain communication in the endonuclease/motor subunit of type I restriction-modification enzyme EcoR124I. J Mol Model 20, 2334 (2014). https://doi.org/10.1007/s00894-014-2334-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-014-2334-1