Abstract
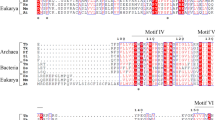
The nuclease NurA is present in all known thermophilic archaea and has been implicated to facilitate efficient DNA double-strand break end processing in Mre11/Rad50-mediated homologous recombinational repair. To understand the structural and functional relationship of this enzyme, we constructed five site-directed mutants of NurA from Sulfolobus tokodaii (StoNurA), D56A, E114A, D131A, Y291A, and H299A, at the conserved motifs, and four terminal deletion mutants, StoNurAΔN (19–331), StoNurAΔNΔC (19–303), StoNurAΔC (1–281), and StoNurAΔC (1–303), and characterized the proteins biochemically. We found that mutation at the acidic residue, D56, E114, D131, or at the basic residue, H299, abolishes the nuclease activity, while mutation at the aromatic residue Y291 only impairs the activity. Interestingly, by chemical cross-linking assay, we found that the mutant Y291A is unable to form stable dimer. Additionally, we demonstrated that deletion of the C-terminal amino acid residues 304–331 of StoNurA results in loss of the physical and functional interaction with the single-stranded DNA-binding protein (StoSSB). These results established that the C-terminal conserved aromatic residue Y291 is involved in dimer formation and the C-terminal residues 304–331 of NurA are involved in the interaction with single-stranded DNA-binding protein.





Similar content being viewed by others
References
Constantinesco F, Forterre P, Elie C (2002) NurA, a novel 5′–3′ nuclease gene liked to rad50 and mre11 homologs of thermophilic Archaea. EMBO Rep 3:537–542
Constantinesco F, Forterre P, Koonin EV, Aravind L, Elie C (2004) A bipolar DNA helicase gene, herA, clusters with rad50, mre11 and nurA genes in thermophilic archaea. Nucleic Acids Res 32:1439–1447
Frols S, Gordon PM, Panlilio MA, Duggin IG, Bell SD, Sensen CW, Schleper C (2007) Response of the hyperthermophilic archaeon Sulfolobus solfataricus to UV damage. J Bacteriol 189:8708–8718
Haber JE (2000) Partners and pathways repairing a double-strand break. Trends Genet 16:259–264
Hopfner KP, Karcher A, Shin D, Fairley C, Tainer JA, Carney JP (2000) Mre11 and Rad50 from Pyrococcus furiosus: cloning and biochemical characterization reveal an evolutionarily conserved multiprotein machine. J Bacteriol 182:6036–6041
Hopfner KP, Putnam CD, Tainer JA (2002) DNA double-strand break repair from head to tail. Curr Opin Struct Biol 12:115–122
Hopkins BB, Paull TT (2008) The P. furiosus Mre11/Rad50 complex promotes 5′ strand resection at a DNA double-strand break. Cell 135:250–260
Iyer LM, Makarova KS, Koonin EV, Aravind L (2004) Comparative genomics of the Ftsk-HerA superfamily of pumping ATPases: implications for the origins of chromosome segregation, cell division and viral capsid packing. Nucleic Acids Res 32:52–5260
Karran P (2000) DNA double strand break repair in mammalian cells. Curr Opin Genet Dev 10:144–150
Khanna KK, Jackson SP (2001) DNA double-strand breaks: signaling, repair and the cancer connection. Nat Genet 27:247–254
Kowalczykowski SC, Dixon DA, Eggleston AK, Lauder SD, Rehrauer WM (1994) Biochemistry of homologous recombination in Escherichia coli. Microbiol Rev 58:401–465
Manzan A, Pfeiffer G, Hefferin ML, Lang CE, Carney JP, Hopfner KP (2004) MlaA, a hexameric ATPase linked to the Mre11 complex in archaeal genomes. EMBO Rep 5:54–59
Mimitou EP, Symington LS (2008) Sae2, Exo1 and Sgs1 collaborate in DNA double-strand break processing. Nature 455:770–774
Mimitou EP, Symington LS (2009) Nuclease and helicase take center stage in homologous recombination. Trends Biochem Sci 34:264–272
Nishino T, Morikawa K (2002) Structure and function of nucleases in DNA repair: shape, grip and blade of the DNA scissors. Oncogene 21:9022–9032
Niu H, Raynard S, Sung P (2009) Multiplicity of DNA end resection machineries in chromosome break repair. Genes Dev 23:1481–1486
Paques F, Haber JE (1999) Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 63:349–404
Rolfsmeier ML, Laughery MF, Haseltine CA (2010) Repair of DNA double-strand breaks following UV damage in three Sulfolobus solfataricus strains. J Bacteriol 192:4954–4962
Wadsworth RIM, White MF (2001) Identification and properties of the crenarchaeal single-stranded DNA binding protein from Sulfolobus solfataricus. Nucleic Acids Res 29:914–920
Wei T, Zhang S, Zhu S, Ni J, Sheng D, Shen Y (2008) Physical and functional interaction between archaeal single-stranded DNA-binding protein and the 5′–3′ nuclease NurA. Biochem Biophys Res Commun 367:523–529
Zhang S, Wei T, Hou G, Zhang C, Liang P, Ni J, Sheng D, Shen Y (2008) Archaeal DNA helicase HerA interacts with Mre11 homologue and unwinds blunt-ended double-stranded DNA and recombination intermediates. DNA Repair 8:380–391
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (30930002 and 30870046 to Y.S., 30770050 to J.N.).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Driessen.
Electronic supplementary material
Below is the link to the electronic supplementary material.
792_2010_351_MOESM2_ESM.tif
Supplementary Fig. 2. Construction of deletion mutants of StoNurA. (A) Limited proteolysis of StoNurA by trypsin. Purified StoNurA (50 μg) was incubated at 30°C with 0.5 μg trypsin as described in Materials and Methods. At the indicated times, aliquots were quenched and analyzed by SDS-PAGE. (B) A schematic representation of the wild type StoNurA and its truncated fragments StoNurAΔN(19-331), StoNurAΔN(19-303), StoNurAΔN(1-281), and StoNurAΔN(19-303). (C) SDS-PAGE of the purified recombinant deletion mutant proteins of StoNurA. (TIFF 547 kb)
792_2010_351_MOESM3_ESM.tif
Supplementary Fig. 3. Single-stranded DNA-binding and nuclease activities of the wild type and mutant StoNurA proteins. (A) DNA-binding activity. Increasing concentrations of proteins (0, 1.0, 5.0, and 10.0 pmol, lanes 1-4) were incubated with 0.2 pmol ssDNA. The reaction products were separated by 6% non-denaturing gels and visualized using a FluoImager 585. (B) Nuclease activity. The reaction mixtures (20 μl) contained 2 pmol ssDNA and increasing amounts (0.5, 1.0, 2.0, and 5.0 pmol) of each protein. The products were resolved on 15% polyacrylamide gels containing 7 M urea and visualized using a FluoImager 585. (C) Quantification of results in (B). Each value was calculated based on the results of three independent reactions. (TIFF 844 kb)
792_2010_351_MOESM4_ESM.tif
Supplementary Fig. 4. Double-stranded DNA-binding and exonuclease activities of the wild type and mutant StoNurA proteins. (A) DNA-binding activity of StoNurA and its truncated forms. (B) Exonuclease activity. (C) Quantification of results in (B). Each value was calculated based on the results of three independent reactions. (TIFF 958 kb)
Rights and permissions
About this article
Cite this article
Wei, T., Zhang, S., Hou, L. et al. The carboxyl terminal of the archaeal nuclease NurA is involved in the interaction with single-stranded DNA-binding protein and dimer formation. Extremophiles 15, 227–234 (2011). https://doi.org/10.1007/s00792-010-0351-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-010-0351-2