Abstract
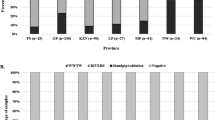
A variety of viruses can cause acute flaccid paralysis (AFP). However, the causative agent, sometimes, remains undetermined. Metagenomics helps in identifying viruses not diagnosed by conventional methods. Stool samples from AFP (n = 104) and non-AFP (n = 114) cases that tested enterovirus-negative by WHO standard methods were investigated. A metagenomics approach, first used on five pools of four samples each, revealed the presence of adenovirus sequences. Amplification in A549 cells and full-genome sequencing were used for complete virus identification and for designing a PCR assay to screen individual related samples. Metagenomic analysis showed that adenovirus sequences that were closely to the A31 and A61 genotypes were the most abundant. Two out of the corresponding 20 individual samples were found positive by PCR, and isolates were obtained in cell culture. Phylogenetic analysis based on complete genome sequences showed that the viruses belong to HAdV-A31 genotype (98-100% nucleotide sequence identity). PCR analysis of stool samples from all AFP and non-AFP cases revealed that a larger proportion of the positive samples were from AFP cases (17.3%) than from non-AFP cases (2.4%). These results open the way to studies aiming to test a possible role of HAdV-A31 in the pathogenesis of AFP.



Similar content being viewed by others
References
Centers for Disease Control and Prevention (CDC) (2009) Progress toward interruption of wild poliovirus transmission—worldwide, 2008. MMWR Morb Mortal Wkly Rep 58:308–312. https://www.ncbi.nlm.nih.gov/pubmed/19343011
da Silva EE, Winkler MT, Pallansch MA (1996) Role of enterovirus 71 in acute flaccid paralysis after the eradication of poliovirus in Brazil. Emerg Infect Dis 2:231–233. https://doi.org/10.3201/eid0203.960312
Junttila N, Lévêque N, Kabue JP et al (2007) New enteroviruses, EV-93 and EV-94, associated with acute flaccid paralysis in the Democratic Republic of the Congo. J Med Virol 79:393–400. https://doi.org/10.1002/jmv.20825
Saeed M, Zaidi SZ, Naeem A et al (2007) Epidemiology and clinical findings associated with enteroviral acute flaccid paralysis in Pakistan. BMC Infect Dis 7:6. https://doi.org/10.1186/1471-2334-7-6
Saad M, Youssef S, Kirschke D et al (2005) Acute flaccid paralysis: the spectrum of a newly recognized complication of West Nile virus infection. J Infect 51:120–127. https://doi.org/10.1016/j.jinf.2004.10.005
Singh SS, Manimunda SP, Sugunan AP et al (2008) Four cases of acute flaccid paralysis associated with chikungunya virus infection. Epidemiol Infect 136:1277–1280. https://doi.org/10.1017/s0950268807009739
Rose G, Wooldridge DJ, Anscombe C et al (2015) Challenges of the unknown: clinical application of microbial metagenomics. Int J Genom 2015:292950. https://doi.org/10.1155/2015/292950
Xie GC, Yu JM, Duan ZJ (2013) New strategy for virus discovery: viruses identified in human feces in the last decade. Sci China 56:688–696. https://doi.org/10.1016/j.jcv.2015.02.017
Victoria JG, Kapoor A, Li L et al (2009) Metagenomic analyses of viruses in stool samples from children with acute flaccid paralysis. J Virol 83:4642–4651. https://doi.org/10.3201/eid1904.121390
Kapoor A, Simmonds P, Slikas E et al (2010) Human bocaviruses are highly diverse, dispersed, recombination prone, and prevalent in enteric infections. J Infect Dis 201:1633–1643. https://doi.org/10.1086/652416
Li L, Victoria J, Kapoor A et al (2009) A novel picornavirus associated with gastroenteritis. J Virol 83:12002–12006. https://doi.org/10.1128/JVI.01241-09
Rezig D, Ben Farhat E, Touzi H et al (2015) Prevalence of human cosaviruses in Tunisia, North Africa. J Med Virol 87:940–943. https://doi.org/10.1002/jmv.24076
Singh G, Zhou X, Lee JY et al (2015) Recombination of the epsilon determinant and corneal tropism: Human adenovirus species D types 15, 29, 56, and 69. Virology 485:452–459. https://doi.org/10.1016/j.virol.2015.08.018
Rosete DP, Manjarrez ME, Barrón BL (2008) Adenoviruses C in non-hospitalized Mexican children older than five years of age with acute respiratory infection. Mem Inst Oswaldo Cruz 103:195–200 (PMID: 18425273)
Robinson CM, Rajaiya J, Zhou X et al (2011) The E3 CR1-gamma gene in human adenoviruses associated with epidemic keratoconjunctivitis. Virus Res 160:120–127. https://doi.org/10.1016/j.virusres.2011.05.022
Liu L, Qian Y, Zhang Y et al (2014) Adenoviruses associated with acute diarrhea in children in Beijing, China. PLoS One 9:e88791. https://doi.org/10.1371/journal.pone.0088791
Venard V, Carret A, Corsaro D et al (2000) Genotyping of adenoviruses isolated in an outbreak in a bone marrow transplant unit shows that diverse strains are involved. J Hosp Infect 44:71–74. https://doi.org/10.1053/jhin.1999.0656
Walls T, Shankar AG, Shingadia D (2003) Adenovirus: an increasingly important pathogen in paediatric bone marrow transplant patients. Lancet Infect Dis 3:79–86 (PMID: 12560192)
Seidemann K, Heim A, Pfister ED et al (2004) Monitoring of adenovirus infection in pediatric transplant recipients by quantitative PCR: report of six cases and review of the literature. Am J Transpl 4:2102–2108. https://doi.org/10.1111/j.1600-6143.2004.00631.x
Kampmann B, Cubitt D, Walls T et al (2005) Improved outcome for children with disseminated adenoviral infection following allogeneic stem cell transplantation. Br J Haematol 130:595–603. https://doi.org/10.1111/j.1365-2141.2005.05649.x
Madisch I, Hofmayer S, Moritz C et al (2007) Phylogenetic analysis and structural predictions of human adenovirus penton proteins as a basis for tissue-specific adenovirus vector design. J Virol 81:8270–8281. https://doi.org/10.1128/JVI.00048-07
Hofmayer S, Madisch I, Darr S et al (2009) Unique sequence features of the Human adenovirus 31 complete genomic sequence are conserved in clinical isolates. BMC Genom 10:557. https://doi.org/10.1186/1471-2164-10-557
Matsushima Y, Shimizu H, Phan TG et al (2011) Genomic characterization of a novel human adenovirus type 31 recombinant in the hexon gene. J Gen Virol 92:2770–2775. https://doi.org/10.1099/vir.0.034744-0
Schnurr D, Bollen A, Crawford-Miksza L et al (1995) Adenovirus mixture isolated from the brain of an AIDS patient with encephalitis. J Med Virol 47:168–71. https://academic.oup.com/cid/article-abstract/31/3/830/299730
Ohtsuki N, Kimura S, Nezu A (2000) Three cases with acute encephalopathy related with adenovirus type 7 infection. No To Hattatsu 32:68–72 (PMID: 10655755)
Nagasawa H, Wada M, Kurita K et al (2006) A case of non-herpetic acute limbic encephalitis associated with a type-2 adenovirus infection. Rinsho Shinkeigaku 46:322–327 (PMID: 16886798)
Frange P, Peffault de Latour R et al (2011) Adenoviral infection presenting as an isolated central nervous system disease without detectable viremia in two children after stem cell transplantation. J Clin Microbiol 49:2361–2364. https://doi.org/10.1128/JCM.00080-11
Dubberke ER, Tu B, Rivet DJ et al (2006) Acute meningoencephalitis caused by adenovirus serotype 26. J Neurovirol 12:235–240. https://doi.org/10.1080/13550280600846633
Ooi MH, Wong SC, Clear D et al (2003) Adenovirus type 21-associated acute flaccid paralysis during an outbreak of hand-foot-and-mouth disease in Sarawak, Malaysia. Clin Infect Dis 36:550–559. https://doi.org/10.1086/367648
Hovi T, Stenvik M (2000) Surveillance of patients with acute flaccid paralysis in Finland: report of a pilot study. Bull World Health Organ 78:298–304 (PMC2560699)
de Azevedo JP, Nascimento LR, Cortinovis MC et al (2004) Characterization of species B adenoviruses isolated from fecal specimens taken from poliomyelitis-suspected cases. J Clin Virol 31:248–252. https://doi.org/10.1016/j.jcv.2004.04.007
Roberts JA, Grant KA, Ibrahim A, Thorley BR (2008) Annual report of the Australian National Poliovirus Reference Laboratory, 2007. Commun Dis Intell Q Rep 32:308–315 (PMID: 19062766)
Bingjun T, Yoshida H, Yan W et al (2008) Molecular typing and epidemiology of non-polio enteroviruses isolated from Yunnan Province, the People’s Republic of China. J Med Virol 80:670–679. https://doi.org/10.1002/jmv.21122
Ivanova OE, Yurashko OV, Eremeeva TP et al (2012) Adenovirus isolation rates in acute flaccid paralysis patients. J Med Virol 84:75–80. https://doi.org/10.1002/jmv.22265
Bayrakdar F, Coşgun Y, Salman Atak T et al (2016) Investigation of adenovirus isolation frequency from the stool samples of patients suspected with acute flaccid paralysis. Mikrobiyol Bul 50:287–292 (PMID: 27175501)
World Health Organization (2004) Dept. of Immunization, Vaccines and Biologicals. Polio laboratory manual 4th edition. World Health Organization, Geneva. http://www.who.int/iris/handle/10665/68762
Gagnieur L, Cheval J, Gratigny M et al (2014) Unbiased analysis by high throughput sequencing of the viral diversity in fetal bovine serum and trypsin used in cell culture. Biologicals 42(3):145–152. https://doi.org/10.1016/j.biologicals.2014.02.002
Besemer J, Borodovsky M (1999) Heuristic approach to deriving models for gene finding. Nucleic Acids Res 27(19):3911–3920 (PMC148655)
Madisch I, Harste G, Pommer H, Heim A (2005) Phylogenetic analysis of the main neutralization and hemagglutination determinants of all human adenovirus prototypes as a basis for molecular classification and taxonomy. J Virol 79:15265–15276. https://doi.org/10.1128/JVI.79.24.15265-15276.2005
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120 (PMID: 7463489)
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599. https://doi.org/10.1093/molbev/msm092
Crawford-Miksza L, Schnurr DP (1996) Analysis of 15 adenovirus hexon proteins reveals the location and structure of seven hypervariable regions containing serotype-specific residues. J Virol 70:1836–1844 (PMID: 8627708)
Greninger AL (2017) A decade of RNA virus metagenomics is (not) enough. Virus Res 244:218–229. https://doi.org/10.1016/j.virusres.2017.10.014
Bodemer C, Sauvage V, Mahlaoui N et al (2014) Live rubella virus vaccine long-term persistence as an antigenic trigger of cutaneous granulomas in patients with primary immunodeficiency. Clin Microbiol Infect 20(10):O656–O663. https://doi.org/10.1111/1469-0691.12573
Frémond ML, Pérot P, Muth E et al (2015) Next-generation sequencing for diagnosis and tailored therapy: a case report of astrovirus-associated progressive encephalitis. J Pediatr Infect Dis Soc 4(3):e53–e57. https://doi.org/10.1093/jpids/piv040
Fredericks DN, Relman DA (1996) Sequence-based identification of microbial pathogens: a reconsideration of Koch’s postulates. Clin Microbiol Rev 9:18–33 (PMC172879)
Jonsson MI, Lenman AE, Frängsmyr L et al (2009) Coagulation factors IX and X enhance binding and infection of adenovirus types 5 and 31 in human epithelial cells. J Virol 83:3816–3825. https://doi.org/10.1128/JVI.02562-08
Acknowledgements
The authors thank the directory of Primary Health Care in Tunisia, and the national staff of the Ministry of Health at all levels, for their efforts in the field work as part of the case investigations and for providing epidemiological and clinical data, and the company PathoQuest (Paris) for the bioinformatics analysis of NGS data from the five pools of stools.
Funding
This study was funded by Laboratoire d’Excellence ‘Integrative Biology of Emerging Infectious Diseases’ [grant number ANR-10-LABX-62-IBEID], “programme Transversal de Recherche PTR484”, Action Concertée des Instituts Pasteur ACIP A22-16, by the Fondation Total [grant number S-CM15010-05B] (http://fondation.total/fr) and the Tunisian Ministry for Scientific Research and Technology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Handling Editor: T. K. Frey.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Haddad-Boubaker, S., Joffret, ML., Pérot, P. et al. Metagenomic analysis identifies human adenovirus 31 in children with acute flaccid paralysis in Tunisia. Arch Virol 164, 747–755 (2019). https://doi.org/10.1007/s00705-018-04141-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-018-04141-5