Abstract
Hematopoietic stem cells (HSCs) exhibit multilineage differentiation and self-renewal activities that maintain the entire hematopoietic system during an organism’s lifetime. These abilities are sustained by intrinsic transcriptional programs and extrinsic cues from the microenvironment or niche. Recent studies using metabolomics technologies reveal that metabolic regulation plays an essential role in HSC maintenance. Metabolic pathways provide energy and building blocks for other factors functioning at steady state and in stress. Here we review recent advances in our understanding of metabolic regulation in HSCs relevant to cell cycle quiescence, symmetric/asymmetric division, and proliferation following stress and lineage commitment, and discuss the therapeutic potential of targeting metabolic factors or pathways to treat hematological malignancies.
Similar content being viewed by others
Introduction
Hematopoietic stem and progenitor cells (HSPCs) in bone marrow (BM) produce all differentiated blood cells in adult organisms. HSPCs are composed of hematopoietic stem cells (HSCs) and several types of lineage-biased, but not fully committed, hematopoietic progenitor cells (HPCs). HSC capacities such as self-renewal and multilineage differentiation maintain the whole hematopoietic system over an organism’s lifetime [1], and contribute to blood regeneration under stress condition, whereas HPCs reportedly function to maintain the supply of blood cells at steady state [2, 3]. The surrounding BM microenvironment or “niche” determines HSC fate, namely its quiescence, symmetric/asymmetric division, differentiation, or migration potential, in both steady state and during stress. It is now evident that metabolic programs operating in HSCs play critical roles in maintaining hematopoietic homeostasis [4], often in response to BM niche signals [5]. Cellular pathways generate various metabolites through enzymatic activity that supplies material used as fuel, building blocks for other factors, or modulators of intra- or intercellular activities (Fig. 1). Despite the fact that numerous biochemical studies have addressed HSC metabolic regulation, numerous gaps in our knowledge of that regulation remain.
Recently, novel means to fractionate metabolites using highly sensitive mass spectrometric technology [6] have dramatically facilitated metabolome analysis. These technologies reduce the need for high cell numbers and increase sensitivity of metabolite selection. As a result, metabolome analysis is feasible in rare cell populations such as HSCs. These analyses have revealed that metabolic programs play critical roles in maintaining stem cell “stemness” (Fig. 2). Here we review recent advances in the field of how metabolic changes regulate HSC dynamics under quiescence, proliferation, and differentiation from the viewpoint of each pathway. In addition to the normal HSC system, we also review activity of leukemia-related metabolic pathways, tumor-associated metabolites (oncometabolites), and related epigenetic regulation by oncometabolites in hematological malignancies such as leukemia.
Hypoxia and anaerobic glycolysis in quiescent HSCs
Tissue stem cells are classified into two categories based on cell cycle status: proliferative and quiescent. Most HSCs and mesenchymal stem cells in the BM, as well as epidermal and melanocyte stem cells in skin and neural stem cells in central nervous system are in a quiescent state. These quiescent stem cells suppress generation of reactive oxygen species (ROS), which cause oxidative damage. By contrast, proliferating stem cells will finally undergo replicative senescence and stop cycling permanently, suggesting that cell temporary cycle arrest is an adaptive strategy to preserve stem cell capacity. Accordingly, a quiescent stem cell state positively contributes to organismal homeostasis through an animal’s lifetime. Due to limited blood perfusion and high oxygen consumption by resident hematopoietic cells, the BM shows a much lower pO2 relative to peripheral blood [7, 8]. Thus, energy supplies in HSCs are maintained by a unique metabolic program in an oxygen-limited environment. A fundamental intracellular energy source for cells is glucose, which is catabolized to pyruvate via sequential glycolytic reactions. In anaerobic (hypoxic) conditions, lactate dehydrogenase (LDH) converts pyruvate to lactate. These activities, which do not require molecular oxygen, are called anaerobic glycolysis and generate only two ATP molecules per one molecule of glucose. In contrast, in aerobic (or normoxic) conditions, glycolysis-derived pyruvate enters the mitochondrial tricarboxylic acid (TCA) cycle and produces 36 molecules of ATP per 1 molecule of glucose in oxidative phosphorylation (OXPHOS). Glycolysis can also occur in aerobic conditions and is then called aerobic glycolysis. Thus, the end-product of glycolysis differs between anaerobic (lactate) and aerobic (pyruvate) conditions. The latter plus the TCA cycle and OXPHOS produce cellular energy more efficiently than does anaerobic glycolysis but requires molecular oxygen for completion. Thus hypoxic conditions inhibit OXPHOS, while a by-product of OXPHOS, ROS generation, injures HSCs [9]. Consequently, suppression of OXPHOS in BM cells by environmental hypoxia is advantageous. A glycolytic program in quiescent HSCs under hypoxia is reportedly activated by glycolytic enzyme expression and suppression of pyruvate dehydrogenase (PDH), which when active allows pyruvate to enter the TCA cycle [10]. Both activities are induced by hypoxia-inducible factor-1α (HIF-1α) [11] and function coordinately to maintain HSC capacity.
Regulation of anaerobic glycolysis by HIF-1α
HIF-1α is a basic helix-loop-helix type transcription factor that responds and adapts to hypoxia at cellular and organismal levels. Under normoxia, prolyl residues in the HIF-1α oxygen-dependent degradation domain (ODD) and an asparagine residue in the C-terminal transactivation domain (C-TAD) are hydroxylated [12], activities regulated by two different oxygen-dependent hydroxylases: prolyl hydroxylase domain (PHD) proteins modifying proline and factor-inhibiting HIF-1 proteins (FIH-1) targeting a C-TAD asparagine residue. Prolyl-hydroxylated ODD is recognized by an E3 ubiquitin ligase, von Hippel–Lindau (VHL) protein, and then degraded by the ubiquitin/proteasome pathway. Moreover, the asparagine-hydroxylated form of the T-CAD cannot interact with p300/CBP, inhibiting transactivation. Thus, in normoxia, HIF-1α is relatively unstable and shows downregulated transactivation activity. By contrast, under hypoxia, PHDs and FIH-1 lose hydroxylase activity and non-hydroxylated HIF-1α protein evades degradation and shows full transactivation ability, enabling the formation of a heterodimeric transcriptional complex (called HIF-1) including the oxygen-independent subunit HIF-1β. HIF-1 then translocates to the nucleus where it activates expression of genes required for adaptation to hypoxia. In addition, HIF-1α is transcriptionally induced by Meis1, a transcription factor essential for HSC stemness [13]. Metabolome and flux analysis suggests that HSCs consume less oxygen and activate of glycolysis to a greater extent than their differentiated counterparts [10], an observation partly explained by relatively higher, HIF-1α-dependent expression of factors catalyzing anaerobic glycolysis [14]. HIF-1 also indirectly regulates glycolytic enzyme expression and cell survival through the autocrine Cripto/GRP78 pathway and through expression of Vegf [15]. HIF-1α deletion in mice reduces expression of glycolytic enzymes and enhances the HSC cell cycle and mobilization of HSCs from the BM [10, 11]. Lactate dehydrogenase (LDH), which functions at the last step of anaerobic glycolysis, catalyzes pyruvate conversion to lactate. Loss of one LDH subunit (LDHA) depletes functional HSCs and HPCs from BM [16]. Therefore, glycolytic metabolism sustains HSC survival, quiescence, and BM anchoring.
The differentiation state of hematopoietic cells is also associated with different glycolytic enzymes. One example is pyruvate kinase (PK), an enzyme that transfers a high-energy phosphate group from phosphoenolpyruvate to ADP, resulting in conversion of phosphoenolpyruvate to pyruvate and ATP generation. Alternative splicing of the PK muscle form (PKM) produces two protein isoforms (PKM1 and PKM2) [17]. Selective loss of the PKM2 isoform in mice impairs activity of HPCs but not HSCs [16]. Although both PKM1 and PKM2 are expressed in HSCs and HSPs, these observations indicate that each enzyme plays a unique role in stem cell or progenitor capacity. Stage-specific regulation of glycolytic enzymes remains a topic to be explored.
PDH regulation of metabolite flux into the TCA cycle
At the last step of aerobic glycolysis, pyruvate is converted to acetyl-CoA in mitochondria by the PDH complex. Three serine residues in the E1α subunit of that complex are phosphorylated by PDH kinase (Pdk) [18], a modification that blocks PDH activity and reduces metabolite flux from glycolysis to the TCA cycle. Upregulation of glycolysis by HIF-1α in HSCs increases pyruvate incorporation into the TCA cycle [10]. However, HSCs contain fewer mitochondria than differentiated cells [19, 20], and mitochondrial morphology in HSCs is immature relative to that seen in differentiated cells, indicative of low mitochondrial membrane potential and low NADH levels. In fact, HSCs depend less on OXPHOS, based on findings that ATP-dependent dye efflux activity in HSCs is maintained in the presence of an inhibitor of cytochrome oxidase [10]. These analyses suggest that metabolic flux from anaerobic glycolysis to the TCA cycle/OXPHOS is reduced in HSCs. In support of this idea, PDH-E1α phosphorylation levels are higher in HSCs than in HPCs, suggesting high Pdk activity in HSCs. In mammals, Pdk has four isozymes: Pdk1, Pdk2, Pdk3, and Pdk4 [18], and HIF-1α expression positively correlates with that of two of them, Pdk2 and Pdk4 [10]. Pdk2 or Pdk4 overexpression also rescues repopulation defects in HIF-1α-deficient HSCs following transplantation, suggesting that Pdk2 and Pdk4 function downstream of HIF-1α. Loss of both Pdk2 and Pdk4 in mice promotes loss of HSC quiescence as well as transplantation defects. It is reported that among the three PDH-E1α phosphorylation sites targeted by Pdks, site 3 (Ser 203 in humans and Ser 232 in mice) is phosphorylated only by Pdk1. Thus, Pdk1 might play a unique role in PDH regulation. Accordingly, Pdk1 knockdown reportedly induces increased ROS and promotes transplantation defects in mouse HSCs [21]. Taken together, Pdks play an essential role in regulating pyruvate entry into the TCA cycle. Moreover, phosphorylated PDH-E1α subunit is dephosphorylated by pyruvate dehydrogenase phosphatase (Pdp) [22]. Thus, reversible regulation of PDH activity by Pdks and Pdp enables fine-tuning of metabolic flux from glycolysis to the TCA cycle in various cellular states.
Role of the TCA cycle and OXPHOS in HSC differentiation
As noted, although mitochondrial OXPHOS is an effective cellular energy-generating strategy, normal quiescent HSCs depend heavily on anaerobic glycolysis. However, once stimulated to divide, HSCs start to activate OXPHOS to meet metabolic demands of proliferation and differentiation. Once activated, HSCs produce ROS and enter the cell cycle. Labeling with a ROS-sensitive fluorescent dye enables to classification of HSPCs as ROShi or ROSlo populations [23]. ROShi HSPCs lose cell cycle quiescence and tend to differentiate. Quiescent HSPCs retain higher repopulation capacity when transplanted. Several enzymes critical for mitochondrial metabolism reportedly regulate HSC differentiation and function. For example, the mitochondrial enzyme PTPMT1 (PTEN-like phosphatase) encoded by nuclear DNA is critical for mitochondrial structure and function. Ptpmt1-deficient HSCs show respiratory defects and a differentiation block [24]. On the other hand, loss of fumarate hydratase (Fh1), a TCA cycle enzyme, in HSCs blocks that cycle. Fh1-deficient fetal HSCs fail to self-renew and differentiate into a lymphoid lineage, defects potentially due to increased histone trimethylation, including H3K9me3, H3K27me3, and H3K36me3 but not H3K4me3 [25]. Recently, mitochondrial dynamics as well as activity of mitochondrial enzymes was shown to contribute to HSC heterogeneity. Mitofusin 2 (Mfn2) deficiency mainly affects lymphoid- but not myeloid-dominant HSCs in the HSC population. Mfn2 activity modulates calcium signaling and negatively regulates nuclear translocation of nuclear factor of activated T cells (Nfat) to sustain lymphoid potential [26].
In addition to maintaining a steady state, mitochondrial energy production is crucial for HSC activity under stress. Fanconi anemia is a hereditary disorder associated with BM failure. HSCs from murine models of the disease, Fanca −/− or Fancc −/− mice, show relatively greater dependence on OXPHOS than on glycolysis. As a result, HSCs from Fanconi anemia models are more susceptible to poisoning by sodium azide treatment than are wild-type HSCs [27]. OXPHOS activation requires p53 induction, as p53 loss in Fanconi anemia model HSCs makes them more susceptible to the glycolysis inhibitor 2-deoxy-d-glucose. By contrast, chronological aging induces mitochondrial accumulation in a subset of HSCs [28], likely because autophagic clearance of normal activated mitochondria is augmented in those cells. Thus, mitochondrial oxygen consumption increases in aging HSCs, suggesting a metabolic shift to OXPHOS.
Mitochondrial OXPHOS constitutes a critical “energy factory” due to its ATP-generating activity. By contrast, recent cancer studies reveal that an important role of OXPHOS during cell proliferation is aspartate biosynthesis [29, 30], indicating that OXPHOS is activated not only to produce ATP production. This finding raises the intriguing possibility that in HSCs OXPHOS participates in activities other than energy production, e.g., metabolites production for differentiation or proliferation. Taken together, these observations indicate that while glycolysis regulates quiescence in HSCs, TCA and OXPHOS are essential for their proliferation and differentiation.
Adipocytes, fatty acid metabolism and hematopoiesis
BM adipose tissue (MAT) occupies up to 70% of BM volume in adult humans [31]. MAT volume in bones increases with age and in pathological situations such as aplastic anemia [32]. Factors derived from MAT negatively regulate HSC and progenitor proliferation in vivo [33]. Adipose tissue also serves as an extramedullary HSC niche [34]. Obesity promotes reduction of the adipokine adiponectin and increases TNF-alpha production from BM macrophages. HSPCs exposed to TNF-alpha show defective JAK-STAT signaling and impaired clearance of infectious pathogens. Adiponectin treatment promotes HSPC proliferation in obese or adiponectin-deficient mice by suppressing BM inflammation [35]. These findings suggest that local and systemic factors derived from adipose tissues regulate HSPCs. Adipose tissue also serves as an “energy warehouse”, storing excess energy as triglyceride and releasing fatty acids (FA) to fulfill energy demands. FA is transported to mitochondria by carnitine palmitoyl transferase 1 (CPT1) and catabolized to acetyl-CoA by β-oxidation. Acetyl-CoA is utilized to fuel the TCA cycle to produce ATP [36]. Ex vivo treatment of mouse HSCs with the CPT1 inhibitor etomoxir induces repopulation defects following HSC transplantation [37]. In mouse, loss of PPAR-δ, a master regulator of FA oxidation (FAO) including β-oxidation, in HSCs induces repopulation defects following transplantation [37]. Treatment of HSCs ex vivo with a PPAR-δ inhibitor reduces asymmetric division and loss of transplantation capacity [37]. These findings suggest that FA metabolism maintains stem cell capacity during cell division via cytokine stimulation. Evaluation of FAO function in homeostatic HSC regulation in genetic models is a topic for future investigation. In addition to its role as an energy source, FA functions as a material for the cellular membrane and it also serves as an intracellular signal transducer. Much of the membrane consists of the phospholipids phosphatidylcholine and phosphatidylethanolamine, and phospholipase A2 releases arachnoid acid from membrane phospholipids. Lipoxygenase (LOX) enzymes incorporate oxygen into unsaturated lipids derived from arachnoid acid. Hydroxyeicosatetraenoic acids (HETEs), such as 12-HETE and 15-HETE, are produced by 12/15-LOX and function as signal transducers in various situations, including inflammatory settings [38]. Loss of 12/15-LOX in HSCs induces defective hematopoiesis in a homeostatic context and repopulation deficits following transplantation, suggesting a critical role for 12/15-LOX in normal hematopoiesis and in maintaining HSC capacity [39]. In pathological settings, 12/15-LOX activity suppresses myeloproliferative disorder [40]. Levels of the HETE factors decrease in patients with chronic myeloid leukemia compared to healthy controls [41], suggesting that LOX suppresses various hematological diseases, including HSC disorders. These findings indicate overall that phospholipid metabolism maintains normal hematopoiesis and protects HSCs from stress.
FA metabolism is becoming increasingly relevant to leukemia research. The FA transporter CD36 is reportedly a marker of poor prognosis in AML [42]. FAO is activated in a subset of leukemia cells, especially in leukemia stem cells (LSCs) residing in gonadal adipose tissue, compared to normal blood cells [43]. These LSCs highly express the FA transporter CD36 and activate lipolysis in gonadal adipose tissue, a metabolic adaptation that protects LSCs from chemotherapy. On the other hand, AML cells cocultured with adipocytes express a high level of the FA-binding protein 4 (FABP4), facilitating intracellular transport of free FA [44]. Inhibition of either FABP4 or CPT1 reduces the leukemia burden and improves survival of AML-transplanted mice. Taken together, FAO is essential for survival of leukemia cells, including LSCs.
The interconnection of lipid metabolism, plasma membrane synthesis, and HSC dynamics is a crucial topic for future studies of hematopoietic homeostasis. At end-product of the arachidonic acid pathway, PGE2 can reportedly expand HSCs in vitro and in vivo [45, 46]. Moreover, neutrophil-derived PGE2 inhibits G-CSF-mediated HSPC mobilization [47]. These observations indicate that the relationship of FA-derived lipid molecules to HSC dynamics should be further investigated.
Amino acid-dependence of HSCs and leukemic cells
One function of amino acids (AAs) is to serve as cellular energy sources. Systemic and local regulation of AA levels modulates the hematopoietic system, including HSC activity. Classically, a low-protein diet has been shown to cause severe granulocytopenia and anemia in rats [48, 49]. Measurement of levels of various AAs using high-performance liquid chromatography (HPLC) reveals that BM contains >100-fold higher concentrations of all 20 AAs than does peripheral blood [50]. Among them, valine is required for HSC proliferation in vitro and for repopulation capacity following transplantation in vivo. Dietary restriction of valine results in a relatively selective depletion of HSCs from BM. As a result, transplanted HSCs can effectively engraft in valine-restricted recipient mice under non-myeloablative conditions [50]. Although the molecular basis for these activities remains unclear, they suggest that dietary valine directly regulates HSC maintenance in the BM niche. On the other hand, intracellular glutamine is metabolized into α-ketoglutarate (α-KG) and ten–eleven translocation (TET) proteins, inducing epigenetic changes in an α-KG-dependent manner (described below). Glutamine is essential for HSC differentiation to erythroid lineage cells [51]. In terms of malignant hematopoiesis, asparagine-related pathways have been targeted as therapy in acute lymphocytic leukemia (ALL). The enzyme that catalyzes asparagine biosynthesis, asparagine synthetase, is expressed at relatively low levels in leukemic cells. As a consequence, leukemic cells, including ALL cells, become addicted to asparagine, and treatment of patients with l-asparaginase induces deamination and depletion of serum asparagine, resulting in leukemia cell death and improved ALL prognosis [52]. While these studies suggest a relationship between AA metabolism and the hematopoietic system, metabolic activities of AAs other than valine, glutamine, or asparagine in HSCs the context of leukemia remain to be elucidated.
Nucleic acid-related pathways and HSPC proliferation
Nucleic acid purines (adenosine and guanine) and pyrimidines (uracil, cytosine and thymine) constitute deoxyribonucleic acid (DNA) and ribonucleic acid (RNA). Purines also serve as energy currency as ATP components, or as signal transducers, such as cAMP and cGMP. De novo purine synthesis starts from phosphoribosyl pyrophosphate (PRPP), which is derived from glucose via the pentose phosphate pathway, and PRPP functions in both adenosine and guanine generation. For guanine generation, PRPP is converted to inosine monophosphate (IMP) by using AAs including aspartic acid, glycine, and glutamine, eventually giving rise to xanthosine monophosphate. This reaction is the first step of guanine synthesis and is catalyzed by IMP dehydrogenase (IMPDH), the rate-limiting enzyme of guanine synthesis [53]. As a result, IMPDH is essential for proliferation of multiple cells including tumor cells and activated lymphocytes [54]. An IMPDH inhibitor, mycophenolic acid (MPA), serves as an immunosuppressant in the context of HSC transplantation by reducing activity and proliferation of reactive lymphocytes. Upon hematological stresses, including chemotherapy or transplantation, quiescent HSCs start to proliferate to increase the supply of differentiated hematopoietic cells and promote BM regeneration. HSCs activate p38α, a member of mitogen-activated protein kinase (MAPK) family, to induce Mitf, a transcriptional regulator that upregulates expression of IMPDH2 (an IMPDH isozyme) in stressed HSPCs. IMPDH2 induction promotes HSPC proliferation and BM regeneration following conditions of hematological stress [55]. These observations indicate that “reprogramming” of the metabolic state is required for HSPC function in these contexts. How a steady state is restored after these changes is a topic for future studies.
Pyrimidine metabolism is also reported to be a possible target for AML [56]. Dihydroorotate dehydrogenase (DHODH) functions in pyrimidine synthesis, producing orotate from dihydroorotate. DHODH localizes in the mitochondrial inner membrane and provides electrons to OXPHOS complex III. Treatment of AML animal models with a DHODH inhibitor induces differentiation of immature leukemic cells into mature myeloid cells. Collectively, purine and pyrimidine nucleic acid metabolism is required to activate proliferation and maintenance of an undifferentiated state in HSPCs.
Metabolic pathways contributing to hematopoietic and leukemic stem/progenitor cell maintenance. A schema illustrating various metabolic pathways related to hematopoietic and leukemic stem cell maintenance. Shown are the glycolysis pathway (boxed in red), the TAC and OXPHOS pathway (boxed in green), the fatty acid metabolism pathway (boxed in blue), the PPP (boxed in pink), the purine synthesis pathway (boxed in purple), the amino acid metabolism pathway (boxed in yellow), the epigenetic regulation (boxed in navy) and the oncometabolite pathway (boxed in black). PK pyruvate kinase, LDH lactate dehydrogenase, PDH pyruvate dehydrogenase, PDK pyruvate dehydrogenase kinase, IDH isocitrate dehydrogenase, G6P glucose-6-phosphate, PEP phosphoenolpyruvate, ATP adenosine triphosphate, FH fumarate hydroxylase, TCA tricarboxylic acid, α-KG α-ketoglutarate, 2-HG 2-hydroxyglutarate, FADH flavin adenine dinucleotide, NADH nicotinamide adenine dinucleotide, OXPHOS oxidative phosphorylation, TET ten–eleven translocation, JHDM JmjC domain-containing histone demethylase, PHD prolyl hydroxylase, HIF-1α hypoxia-inducible factor-1α, IMPDH inosine-5′-monophosphate dehydrogenase, PPP pentose phosphate pathway, PRPP phosphoribosyl diphosphate, IMP inosine-5′-monophosphate, XMP xanthosine monophosphate, GMP guanosine monophosphate, AA arachidonic acid, EPA eicosapentaenoic acid, DHA docosahexaenoic acid, HETE hydroxyeicosatetraenoic acid, 12/15LOX 12/15-lipoxygenase, FA fatty acid, CPT1 carnitine palmitoyltransferase I
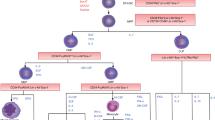
Metabolic dependence of hematopoietic cell compartments. Dependence and characteristics of enzymes or metabolites functioning in various hematopoietic cell compartments. LDHA, PDH/Pkd, and 12/15LOX are essential for quiescent and proliferative HSC, whereas IMPDH, FH, and PKM2 are required for only proliferative HSC. In addition, LSC and leukemia cell demand other pathway differ from normal HSCs; CD36, FABP, and DHODH. PK pyruvate kinase, LDH lactate dehydrogenase, PDH pyruvate dehydrogenase, IDH isocitrate dehydrogenase, IMPDH inosine-5′-monophosphate dehydrogenase, TET ten–eleven translocation enzyme, FABP fatty acid-binding protein, DHODH dihydroorotate dehydrogenase
Epigenetic regulation in leukemic cells by oncometabolites
Some metabolites also directly modify protein or DNA and serve as epigenetic regulators [4], and epigenetic regulation of leukemia cells by tumor-specific metabolites (oncometabolites) has been reported [57, 58]. Most individual AML cases are associated with five different mutations, on average [59]. Mutational analysis of single AML cells suggests stepwise acquisition of mutations [60]. Among them, mutations in metabolism-related enzymes such as TET and isocitrate dehydrogenase (IDH) have been reported primarily in the earliest step of AML. Mutated IDH acquires a neo-enzymatic activity that converts α-KG, a TCA cycle intermediary, into d-2-hydroxyglutarate (d-2HG), while normal IDH lacks this activity [57]. Thus, cells harboring mutant IDH exhibit low α-KG and high d-2HG levels. The JmjC domain-containing histone demethylase (JHDM) demethylates histone lysine residues, an activity requiring α-KG as a cofactor. d-2HG also binds to and competitively inhibits the JHDM α-KG binding site [61]. Consequently, cells harboring mutant IDH show decreased JHDM activity and abnormal patterns of histone methylation [62], outcomes associated with a block in HSC differentiation. Mutant IDH activity also affects DNA methylation. TET family proteins catalyze the conversion of 5-methylcytosine to 5-hydroxymethylcytosine (5-hmC) on DNA and promote DNA demethylation. TET enzymatic activity requires α-KG [63]. As a result, mutation in either TET or IDH impairs DNA demethylation, contributing to AML pathogenesis. Targeting of mutant IDH by inhibitors has been tested as a potential treatment strategy in clinical settings [64]. Mutant IDH induces HSPC expansion in vitro and in vivo [58]. 2HG is a chiral molecule existing as both a d- and l-enantiomer. Both have inhibitory activity against α-KG-dependent dioxygenases. l-2HG is generated from α-KG by LDHA or malate dehydrogenase in an IDH-independent manner in hypoxia or low pH conditions [65, 66]. Mutation of genes encoding the TCA cycle enzymes FH1 or succinate dehydrogenase induces accumulation of fumarate or succinate, respectively. Although these mutations are rare in hematological malignancies, both fumarate and succinate suppress α-KG-dependent dioxygenases, including histone and DNA demethylases, and function as oncometabolites [67]. Thus, a physiological link between glycolysis, mitochondrial metabolism and oncometabolites in HSCs is another important area of investigation.
Conclusion
Multistep metabolic pathways have long been regarded as fundamental cellular “housekeepers”. Recent advances in integrative metabolic analysis, including capillary electrophoresis mass spectrometry, have dramatically changed this classical view and provided solid evidence that metabolism regulates HSC quiescence, differentiation and proliferation (Fig. 2). Moreover, metabolic targeting approaches provide insight into pathophysiology and could suggest therapeutic strategies to treat hematological malignancies. However, we are currently just beginning to elucidate the function of respective metabolic pathways in HSCs and in LSCs: we now know that enzymes or isozymes functioning in one metabolic pathway can function differently in other contexts. Thus improving analytical sensitivity and coverage in metabolomics analysis is critical for understanding of HSC activity. Furthermore, how metabolic features of HSCs change during various conditions, including stress hematopoiesis, require further analysis. Eventually, greater understanding of temporal and spatial metabolic dynamics should lead to novel techniques to manipulate normal HSCs and eradicate leukemic cells.
References
Orkin SH, Zon LI. Hematopoiesis: an evolving paradigm for stem cell biology. Cell. 2008;132(4):631–44.
Sun J, Ramos A, Chapman B, Johnnidis JB, Le L, Ho YJ, et al. Clonal dynamics of native haematopoiesis. Nature. 2014;514(7522):322–7.
Busch K, Klapproth K, Barile M, Flossdorf M, Holland-Letz T, Schlenner SM, et al. Fundamental properties of unperturbed haematopoiesis from stem cells in vivo. Nature. 2015;518(7540):542–6.
Chandel NS, Jasper H, Ho TT, Passegue E. Metabolic regulation of stem cell function in tissue homeostasis and organismal ageing. Nat Cell Biol. 2016;18(8):823–32.
Morrison SJ, Scadden DT. The bone marrow niche for haematopoietic stem cells. Nature. 2014;505(7483):327–34.
Soga T, Igarashi K, Ito C, Mizobuchi K, Zimmermann HP, Tomita M. Metabolomic profiling of anionic metabolites by capillary electrophoresis mass spectrometry. Anal Chem. 2009;81(15):6165–74.
Cipolleschi MG, Dello Sbarba P, Olivotto M. The role of hypoxia in the maintenance of hematopoietic stem cells. Blood. 1993;82(7):2031–7.
Parmar K, Mauch P, Vergilio JA, Sackstein R, Down JD. Distribution of hematopoietic stem cells in the bone marrow according to regional hypoxia. Proc Natl Acad Sci USA. 2007;104(13):5431–6.
Suda T, Takubo K, Semenza GL. Metabolic regulation of hematopoietic stem cells in the hypoxic niche. Cell Stem Cell. 2011;9(4):298–310.
Takubo K, Nagamatsu G, Kobayashi CI, Nakamura-Ishizu A, Kobayashi H, Ikeda E, et al. Regulation of glycolysis by Pdk functions as a metabolic checkpoint for cell cycle quiescence in hematopoietic stem cells. Cell Stem Cell. 2013;12(1):49–61.
Takubo K, Goda N, Yamada W, Iriuchishima H, Ikeda E, Kubota Y, et al. Regulation of the HIF-1alpha level is essential for hematopoietic stem cells. Cell Stem Cell. 2010;7(3):391–402.
Semenza GL. Hypoxia-inducible factor 1 (HIF-1) pathway. Sci STKE. 2007;2007(407):cm8.
Kocabas F, Zheng J, Thet S, Copeland NG, Jenkins NA, DeBerardinis RJ, et al. Meis1 regulates the metabolic phenotype and oxidant defense of hematopoietic stem cells. Blood. 2012;120(25):4963–72.
Cabezas-Wallscheid N, Klimmeck D, Hansson J, Lipka DB, Reyes A, Wang Q, et al. Identification of regulatory networks in HSCs and their immediate progeny via integrated proteome, transcriptome, and DNA methylome analysis. Cell Stem Cell. 2014;15(4):507–22.
Miharada K, Karlsson G, Rehn M, Rorby E, Siva K, Cammenga J, et al. Cripto regulates hematopoietic stem cells as a hypoxic-niche-related factor through cell surface receptor GRP78. Cell Stem Cell. 2011;9(4):330–44.
Wang YH, Israelsen WJ, Lee D, Yu VW, Jeanson NT, Clish CB, et al. Cell-state-specific metabolic dependency in hematopoiesis and leukemogenesis. Cell. 2014;158(6):1309–23.
Mazurek S, Boschek CB, Hugo F, Eigenbrodt E. Pyruvate kinase type M2 and its role in tumor growth and spreading. Semin Cancer Biol. 2005;15(4):300–8.
Harris RA, Bowker-Kinley MM, Huang B, Wu P. Regulation of the activity of the pyruvate dehydrogenase complex. Adv Enzyme Regul. 2002;42:249–59.
Simsek T, Kocabas F, Zheng J, Deberardinis RJ, Mahmoud AI, Olson EN, et al. The distinct metabolic profile of hematopoietic stem cells reflects their location in a hypoxic niche. Cell Stem Cell. 2010;7(3):380–90.
Norddahl GL, Pronk CJ, Wahlestedt M, Sten G, Nygren JM, Ugale A, et al. Accumulating mitochondrial DNA mutations drive premature hematopoietic aging phenotypes distinct from physiological stem cell aging. Cell Stem Cell. 2011;8(5):499–510.
Halvarsson C, Eliasson P, Jonsson JI. Pyruvate dehydrogenase kinase 1 is essential for transplantable mouse bone marrow hematopoietic stem cell and progenitor function. PLoS One. 2017;12(2):e0171714.
Patel MS, Nemeria NS, Furey W, Jordan F. The pyruvate dehydrogenase complexes: structure-based function and regulation. J Biol Chem. 2014;289(24):16615–23.
Jang YY, Sharkis SJ. A low level of reactive oxygen species selects for primitive hematopoietic stem cells that may reside in the low-oxygenic niche. Blood. 2007;110(8):3056–63.
Yu WM, Liu X, Shen J, Jovanovic O, Pohl EE, Gerson SL, et al. Metabolic regulation by the mitochondrial phosphatase PTPMT1 is required for hematopoietic stem cell differentiation. Cell Stem Cell. 2013;12(1):62–74.
Guitart AV, Panagopoulou TI, Villacreces A, Vukovic M, Sepulveda C, Allen L, et al. Fumarate hydratase is a critical metabolic regulator of hematopoietic stem cell functions. J Exp Med. 2017;214(3):719–35.
Luchsinger LL, de Almeida MJ, Corrigan DJ, Mumau M, Snoeck HW. Mitofusin 2 maintains haematopoietic stem cells with extensive lymphoid potential. Nature. 2016;529(7587):528–31.
Du W, Amarachintha S, Wilson AF, Pang Q. SCO2 mediates oxidative stress-induced glycolysis to oxidative phosphorylation switch in hematopoietic stem cells. Stem Cells. 2016;34(4):960–71.
Ho TT, Warr MR, Adelman ER, Lansinger OM, Flach J, Verovskaya EV, et al. Autophagy maintains the metabolism and function of young and old stem cells. Nature. 2017;543(7644):205–10.
Birsoy K, Wang T, Chen WW, Freinkman E, Abu-Remaileh M, Sabatini DM. An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell. 2015;162(3):540–51.
Sullivan LB, Gui DY, Hosios AM, Bush LN, Freinkman E, Vander Heiden MG. Supporting aspartate biosynthesis is an essential function of respiration in proliferating cells. Cell. 2015;162(3):552–63.
Fazeli PK, Horowitz MC, MacDougald OA, Scheller EL, Rodeheffer MS, Rosen CJ, et al. Marrow fat and bone—new perspectives. J Clin Endocrinol Metab. 2013;98(3):935–45.
Masumoto A, Yonekura S, Haida M, Yanagimachi N, Hotta T. Analysis of intramedullary cell density by MRI using the multiple spin-echo technique. Am J Hematol. 1997;55(3):134–8.
Naveiras O, Nardi V, Wenzel PL, Hauschka PV, Fahey F, Daley GQ. Bone-marrow adipocytes as negative regulators of the haematopoietic microenvironment. Nature. 2009;460(7252):259–63.
Han J, Koh YJ, Moon HR, Ryoo HG, Cho CH, Kim I, et al. Adipose tissue is an extramedullary reservoir for functional hematopoietic stem and progenitor cells. Blood. 2010;115(5):957–64.
Masamoto Y, Arai S, Sato T, Yoshimi A, Kubota N, Takamoto I, et al. Adiponectin enhances antibacterial activity of hematopoietic cells by suppressing bone marrow inflammation. Immunity. 2016;44(6):1422–33.
Currie E, Schulze A, Zechner R, Walther TC, Farese RV Jr. Cellular fatty acid metabolism and cancer. Cell Metab. 2013;18(2):153–61.
Ito K, Carracedo A, Weiss D, Arai F, Ala U, Avigan DE, et al. A PML-PPAR-delta pathway for fatty acid oxidation regulates hematopoietic stem cell maintenance. Nat Med. 2012;18(9):1350–8.
Conrad DJ. The arachidonate 12/15 lipoxygenases. A review of tissue expression and biologic function. Clin Rev Allergy Immunol. 1999;17(1–2):71–89.
Kinder M, Wei C, Shelat SG, Kundu M, Zhao L, Blair IA, et al. Hematopoietic stem cell function requires 12/15-lipoxygenase-dependent fatty acid metabolism. Blood. 2010;115(24):5012–22.
Middleton MK, Zukas AM, Rubinstein T, Jacob M, Zhu P, Zhao L, et al. Identification of 12/15-lipoxygenase as a suppressor of myeloproliferative disease. J Exp Med. 2006;203(11):2529–40.
Stenke L, Lauren L, Reizenstein P, Lindgren JA. Leukotriene production by fresh human bone marrow cells: evidence of altered lipoxygenase activity in chronic myelocytic leukemia. Exp Hematol. 1987;15(2):203–7.
Perea G, Domingo A, Villamor N, Palacios C, Junca J, Torres P, et al. Adverse prognostic impact of CD36 and CD2 expression in adult de novo acute myeloid leukemia patients. Leuk Res. 2005;29(10):1109–16.
Ye H, Adane B, Khan N, Sullivan T, Minhajuddin M, Gasparetto M, et al. Leukemic stem cells evade chemotherapy by metabolic adaptation to an adipose tissue niche. Cell Stem Cell. 2016;19(1):23–37.
Shafat MS, Oellerich T, Mohr S, Robinson SD, Edwards DR, Marlein CR, et al. Leukemic blasts program bone marrow adipocytes to generate a protumoral microenvironment. Blood. 2017;129(10):1320–32.
North TE, Goessling W, Walkley CR, Lengerke C, Kopani KR, Lord AM, et al. Prostaglandin E2 regulates vertebrate haematopoietic stem cell homeostasis. Nature. 2007;447(7147):1007–11.
Ikushima YM, Arai F, Hosokawa K, Toyama H, Takubo K, Furuyashiki T, et al. Prostaglandin E(2) regulates murine hematopoietic stem/progenitor cells directly via EP4 receptor and indirectly through mesenchymal progenitor cells. Blood. 2013;121(11):1995–2007.
Kawano Y, Fukui C, Shinohara M, Wakahashi K, Ishii S, Suzuki T, et al. G-CSF-induced sympathetic tone provokes fever and primes antimobilizing functions of neutrophils via PGE2. Blood. 2017;129(5):587–97.
Kornberg A. Amino acids in the production of granulocytes in rats. J Biol Chem. 1946;164:203–12.
Kornberg A, Daft FS, Sebrell WH. Granulocytopenia and anemia in rats fed diets of low casein content. Science. 1946;103(2682):646–8.
Taya Y, Ota Y, Wilkinson AC, Kanazawa A, Watarai H, Kasai M, et al. Depleting dietary valine permits nonmyeloablative mouse hematopoietic stem cell transplantation. Science. 2016;354(6316):1152–5.
Oburoglu L, Tardito S, Fritz V, de Barros SC, Merida P, Craveiro M, et al. Glucose and glutamine metabolism regulate human hematopoietic stem cell lineage specification. Cell Stem Cell. 2014;15(2):169–84.
Egler RA, Ahuja SP, Matloub Y. l-asparaginase in the treatment of patients with acute lymphoblastic leukemia. J Pharmacol Pharmacother. 2016;7(2):62–71.
Hedstrom L. IMP dehydrogenase: structure, mechanism, and inhibition. Chem Rev. 2009;109(7):2903–28.
Jackson RC, Weber G, Morris HP. IMP dehydrogenase, an enzyme linked with proliferation and malignancy. Nature. 1975;256(5515):331–3.
Karigane D, Kobayashi H, Morikawa T, Ootomo Y, Sakai M, Nagamatsu G, et al. p38alpha activates purine metabolism to initiate hematopoietic stem/progenitor cell cycling in response to stress. Cell Stem Cell. 2016;19(2):192–204.
Sykes DB, Kfoury YS, Mercier FE, Wawer MJ, Law JM, Haynes MK, et al. Inhibition of dihydroorotate dehydrogenase overcomes differentiation blockade in acute myeloid leukemia. Cell. 2016;167(1):171–186.e15.
Ward PS, Patel J, Wise DR, Abdel-Wahab O, Bennett BD, Coller HA, et al. The common feature of leukemia-associated IDH1 and IDH2 mutations is a neomorphic enzyme activity converting alpha-ketoglutarate to 2-hydroxyglutarate. Cancer Cell. 2010;17(3):225–34.
Figueroa ME, Abdel-Wahab O, Lu C, Ward PS, Patel J, Shih A, et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell. 2010;18(6):553–67.
Cancer Genome Atlas Research Network, Ley TJ, Miller C, Ding L, Raphael BJ, Mungall AJ, et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368(22):2059–74.
Corces-Zimmerman MR, Hong WJ, Weissman IL, Medeiros BC, Majeti R. Preleukemic mutations in human acute myeloid leukemia affect epigenetic regulators and persist in remission. Proc Natl Acad Sci USA. 2014;111(7):2548–53.
Xu W, Yang H, Liu Y, Yang Y, Wang P, Kim SH, et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of alpha-ketoglutarate-dependent dioxygenases. Cancer Cell. 2011;19(1):17–30.
Medeiros BC, Fathi AT, DiNardo CD, Pollyea DA, Chan SM, Swords R. Isocitrate dehydrogenase mutations in myeloid malignancies. Leukemia. 2017;31:272–81.
Chiba S. Dysregulation of TET2 in hematologic malignancies. Int J Hematol. 2017;105(1):17–22.
Bose P, Vachhani P, Cortes JE. Treatment of relapsed/refractory acute myeloid leukemia. Curr Treat Options Oncol. 2017;18(3):17.
Intlekofer AM, Dematteo RG, Venneti S, Finley LW, Lu C, Judkins AR, et al. Hypoxia induces production of l-2-hydroxyglutarate. Cell Metab. 2015;22(2):304–11.
Intlekofer AM, Wang B, Liu H, Shah H, Carmona-Fontaine C, Rustenburg AS, et al. l-2-Hydroxyglutarate production arises from noncanonical enzyme function at acidic pH. Nat Chem Biol. 2017;13(5):494–500.
Xiao M, Yang H, Xu W, Ma S, Lin H, Zhu H, et al. Inhibition of alpha-KG-dependent histone and DNA demethylases by fumarate and succinate that are accumulated in mutations of FH and SDH tumor suppressors. Genes Dev. 2012;26(12):1326–38.
Acknowledgements
We thank all members of the Takubo laboratory for indispensable support and E. Lamar for preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
K.T. was supported in part by KAKENHI Grants from MEXT/JSPS (26115005, 15H04861, 16K15507, 26115001, 15K21751), a Grant of the National Center for Global Health and Medicine (26-001), AMED-CREST, an AMED Grant for Realization of Regenerative Medicine and Grants from the Tokyo Biochemical Research Foundation, the Uehara Memorial Foundation, the Japan Leukemia Research Fund, the Japan Rheumatism Foundation, the Japan Foundation for Applied Enzymology, and the Kanae Foundation for the Promotion of Medical Science. D.K. was supported in part by KAKENHI Grant from MEXT/JSPS (17K16199).
About this article
Cite this article
Karigane, D., Takubo, K. Metabolic regulation of hematopoietic and leukemic stem/progenitor cells under homeostatic and stress conditions. Int J Hematol 106, 18–26 (2017). https://doi.org/10.1007/s12185-017-2261-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12185-017-2261-x