Abstract
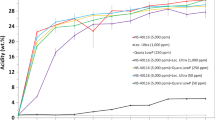
The degumming of crude soybean oil with phospholipase A1 (PLA1) and phospholipase C (PLC) was studied, and optimal conditions were obtained for each enzyme. During degumming with PLA1, more fatty acid was found in the oil than would be expected by hydrolysis of only the terminal fatty acid chains, and glycerophosphophorylcholine and glycerophosphoethanolamine were detected in the gums. These observations indicate that acyl-migration of phospholipid fatty acids occurred during PLA1 degumming. In addition, results showed that PLA1 degumming was capable of reducing the phosphorus content in the oil to levels acceptable for physical refining (<10 mg/kg). During degumming with PLC, an increase of 1,2-diacylglycerol was found, as most phosphatidylcholine and phosphatidylethanolamine were hydrolyzed by this enzyme. Treatment with either enzyme slightly decreased the oxidative stability of the oil and most metals were separated with the gums fraction.




Similar content being viewed by others
Abbreviations
- PLC:
-
Phospholipase C
- PLA1 :
-
Phospholipase A1
- PLA2 :
-
Phospholipase A2
- GPC:
-
Glycerophosphophorylcholine
- GPE:
-
Glycerophosphoethanolamine
- DAG:
-
Diacylglycerol(s)
- PC:
-
Phosphatidylcholine
- PE:
-
Phosphatidylethanolamine
- PI:
-
Phosphatidylinositol
- PA:
-
Phosphatidic acid
- FFA:
-
Free fatty acid(s)
- NHP:
-
Non-hydratable phospholipid(s)
- LPC:
-
Lyso-phosphatidylcholine
- LPE:
-
Lyso-phosphatidylethanolamine
- LPI:
-
Lyso-phosphatidylinositol
- LPA:
-
Lyso-phosphatidic acid
- NPPC:
-
p-Nitrophenylphosphorylcholine
- FA:
-
Fatty acid composition
- FAME:
-
Fatty acid methyl ester(s)
- FID:
-
Flame ionization detector
References
Wang Y, Zhao M, Song K, Wang L, Tang S, Riley WW (2010) Partial hydrolysis of soybean oil by phospholipase A1 (Lecitase Ultra). Food Chem 121:1066–1072
Yu D, Ma Y, Jiang L, Elfalleh W, Shi M, Hu L (2013) Optimization of magnetic immobilized phospholipase A1 degumming process for soybean oil using response surface methodology. Eur Food Res Technol 237:811–817
Dahlke K (1998) An enzymatic process for the physical refining of seed oils. Chem Eng Technol 21:278–281
Zamora R, Olmo C, Navarro JL, Hidalgo FJ (2004) Contribution of phospholipid pyrrolization to the color reversion produced during deodorization of poorly degummed vegetable oils. J Agri Food Chem 52:4166–4171
Dijkstra AJ, Van Opstal M (1989) The total degumming process. J Am Oil Chem Soc 66:1002–1009
Yang J-G, Wang Y-H, Yang B, Mainda G, Guo Y (2006) Degumming of vegetable oil by a new microbial lipase. Food Technol Biotechnol 44:101–104
De Maria L, Vind J, Oxenbøll K, Svendsen A, Patkar S (2007) Phospholipases and their industrial applications. Appl Microbiol Biotechnol 74:290–300
Jiang F, Wang J, Kaleem I, Dai D, Zhou X, Li C (2011) Degumming of vegetable oils by a novel phospholipase B from Pseudomonas fluorescens BIT-18. Bioresour Technol 102:8052–8056
Clausen K (2001) Enzymatic oil—degumming by a novel microbial phospholipase. Eur J Lipid Sci Technol 103:333–340
Huang S, Liang M, Xu Y, Rasool A, Li C (2014) Characteristics and vegetable oils degumming of recombinant phospholipase B. Chem Eng J 237:23–28
Jahani M, Alizadeh M, Pirozifard M, Qudsevali A (2008) Optimization of enzymatic degumming process for rice bran oil using response surface methodology. LWT Food Sci Technol 41:1892–1898
Yang B, Zhou R, Yang J-G, Wang Y-H, Wang W-F (2008) Insight into the enzymatic degumming process of soybean oil. J Am Oil Chem Soc 85:421–425
Dayton CL, Galhardo F (2007) Enzymatic degumming utilizing a mixture of PLA and PLC phospholipases. US Patent 2008/0182322 A1
Dijkstra AJ (2009) Recent developments in edible oil processing. Eur J Lipid Sci Technol 111:857–864
Flieger A, Gong S, Faigle M, Neumeister B (2000) Critical evaluation of p-nitrophenylphosphorylcholine (p-NPPC) as artificial substrate for the detection of phospholipase C. Enzym Microb Technol 26:451–458
Yang M, Zhou X, Jin Y (2008) Non-hydrated phosphatide and its quantitative examination. Chinese J Health Lab Technol 18:71–72
Smiles A, Kakuda Y, MacDonald BE (1988) Effect of degumming reagents on the recovery and nature of lecithins from crude canola, soybean and sunflower oils. J Am Oil Chem Soc 65:1151–1155
Yang B, Wang Y-H, Yang J-G (2006) Optimization of enzymatic degumming process for rapeseed oil. J Am Oil Chem Soc 83:653–658
Avalli A, Contarini G (2005) Determination of phospholipids in dairy products by SPE/HPLC/ELSD. J Chromatogr A 1071:185–190
Zhang K, Wang X, Huang J, Liu Y (2012) Purification of l-alpha glycerylphosphorylcholine by column chromatography. J Chromatogr A 1220:108–114
Pan L, Campana A, Toms M (2000) A kinetic study of phospholipid extraction by degumming process in sunflower seed oil. J Am Oil Chem Soc 77:1273–1277
Dijkstra AJ (2010) Enzymatic degumming. Eur J Lipid Sci Technol 112:1178–1189
Xu X, Fomuso LB, Akoh CC (2000) Synthesis of structured triacylglycerols by lipase-catalyzed acidolysis in a packed bed bioreactor. J Agric Food Chem 48:3–10
Kashima M, Cha G-S, Isoda Y, Hirano J, Miyazawa T (1991) The antioxidant effects of phospholipids on perilla oil. J Am Oil Chem Soc 68:119–122
Koga T, Terao J (1995) Phospholipids increase radical-scavenging activity of vitamin E in a bulk oil model system. J Agric Food Chem 43:1450–1454
Barton N (2008) A new process for degumming: the use of phospholipase C to improve yields during refining of high phosphorus vegetable oils. In: 99th AOCS Annual Meeting and Expo, vol 120. Seattle, p 53–658
Acknowledgments
The work was supported by the Public Welfare Research Funds of State Administration of Grain (201313012-03) and the Major State Basic Research Development Program of China (973 Program, 2012CB720802, 2012CB720806).
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Jiang, X., Chang, M., Wang, X. et al. A Comparative Study of Phospholipase A1 and Phospholipase C on Soybean Oil Degumming. J Am Oil Chem Soc 91, 2125–2134 (2014). https://doi.org/10.1007/s11746-014-2555-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11746-014-2555-6