Abstract
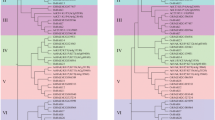
The high-affinity potassium transporter (HKT) genes have crucial roles in the regulation of sodium and potassium transportation in many species, however little is known about maize HKT genes. In this study, we obtained two alternative splicing transcripts of ZmHKT1;1 gene. One was named as ZmHKT1;1a which has the intact coding sequence, and the other was named as ZmHKT1;1b which has a deficiency of the third exon and a retention of the second intron. The phylogenic tree analysis showed that both translation products of ZmHKT1;1a and ZmHKT1;1b belong to group I HKT proteins which prefer for Na+ transport than other cations. ZmHKT1;1a and ZmHKT1;1b showed different response to stress treatment in maize. Overexpressing ZmHKT1;1a or ZmHKT1;1b in transgenic tobacco plants conferred high salt tolerance by increasing root length and fresh weight of plants. When treated with high concentration of salt, transgenic tobacco plants manifested a trend of reduced Na+ content and increased K+ content in both shoot and root, suggesting that ZmHKT1;1 may involve in Na+ unloading and indirectly affect other transporter activity. It was also found that overexpression of ZmHKT1;1a and ZmHKT1;1b caused different expression of stress-related genes. The results in this study indicate that two alternative splicing variants of ZmHKT1;1 might be useful for the development of salt-tolerant transgenic crops.







Similar content being viewed by others
References
Almeida P, Katschnig D, de Boer AH (2013) HKT transporters—state of the art. Int J Mol Sci 14(10):20359–20385
Apse MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by over expression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 85:1256–1258
Corratgé-Faillie C, Jabnoune M, Zimmermann S, Véry A-A, Fizames C, Sentenac H (2010) Potassium and sodium transport in non-animal cells: the Trk/Ktr/HKT transporter family. Cell Mol Life Sci 67(15):2511–2532
Cotsaftis O, Plett D, Shirley N, Tester M, Hrmova M (2012) A two-staged model of Na+ exclusion in rice explained by 3D modeling of HKT transporters and alternative splicing. PLoS ONE 7(7):e39865
Diatloff E, Kumar R, Schachtman DP (1998) Site directed mutagenesis reduces the Na+ affinity of HKT1, an Na+ energized high affinity K+ transporter. FEBS Lett 432:31–36
Garciadeblas B, Senn ME, Banuelos MA, Rodriguez-Navarro A (2003) Sodium transport and HKT transporters: the rice model. Plant J 34(6):788–801
Golldack D, Su H, Quigley F, Kamasani UR, Munoz-Garay C, Balderas E, Popova OV, Bennett J, Bohnert HJ, Pantoja O (2002) Characterization of a HKT-type transporter in rice as a general alkali cation transporter. Plant J 31(4):529–542
Graveley BR (2001) Alternative splicing: increasing diversity in the proteomic world. Trends Genet 17:100–107
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Annu Rev Plant Physiol Plant Mol Biol 51:463–499
Hauser F, Horie T (2010) A conserved primary salt tolerance mechanism mediated by HKT transporters: a mechanism for sodium exclusion and maintenance of high K+/Na+ ratio in leaves during salinity stress. Plant Cell Environ 33(4):552–565
He C, Yan J, Shen G, Fu L, Holaday AS, Auld D, Blumwald E, Zhang H (2005) Expression of an Arabidopsis vacuolar sodium/proton antiporter gene in cotton improves photosynthetic performance under salt conditions and increases fiber yield in the field. Plant Cell Physiol 46:1848–1854
Horie T, Hauser F, Schroeder JI (2009) HKT transporter-mediated salinity resistance mechanisms in Arabidopsis and monocot crop plants. Trends Plant Sci 14(12):660–668
Horsch R, Fry J, Hoffman N, Eichholz D, Rogers S, Fraley R (1985) A simple and general method for transferring genes into plants. Science 227(4691):1229–1231
Huang S, Spielmeyer W, Lagudah ES, James RA, Platten JD, Dennis ES, Munns R (2006) A sodium transporter (HKT7) is a candidate for Nax1, a gene for salt tolerance in durum wheat. Plant Physiol 142(4):1718–1727
Laurie S, Feeney KA, Maathuis FJ, Heard PJ, Brown SJ, Leigh RA (2002) A role for HKT1 in sodium uptake by wheat roots. Plant J 32(2):139–149
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25(4):402–408
Maser P, Hosoo Y, Goshima S, Horie T, Eckelman B, Yamada K, Yoshida K, Bakker EP, Shinmyo A, Oiki S, Schroeder JI, Uozumi N (2002) Glycine residues in potassium channel-like selectivity filters determine potassium selectivity in four-loop-per-subunit HKT transporters from plants. Proc Natl Acad Sci USA 99(9):6428–6433
Mian A, Oomen RJ, Isayenkov S, Sentenac H, Maathuis FJ, Very AA (2011) Over-expression of an Na+-and K+-permeable HKT transporter in barley improves salt tolerance. Plant J 68(3):468–479
Mishra S, Alavilli H, Lee B, Panda SK, Sahoo L (2015) Cloning and characterization of a novel vacuolar Na+/H+ antiporter gene (VuNHX1) from drought hardy legume, cowpea for salt tolerance. Plant Cell Tissue Organ Cult 120:19–33
Moller IS, Tester M (2007) Salinity tolerance of Arabidopsis: a good model for cereals? Trends Plant Sci 12(12):534–540
Moller IS, Gilliham M, Jha D, Mayo GM, Roy SJ, Coates JC, Haseloff J, Tester M (2009) Shoot Na+ exclusion and increased salinity tolerance engineered by cell type-specific alteration of Na+ transport in Arabidopsis. Plant Cell 21(7):2163–2178
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Munns R, James RA, Xu B, Athman A, Conn SJ, Jordans C, Byrt CS, Hare RA, Tyerman SD, Tester M, Plett D, Gilliham M (2012) Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat Biotechnol 30(4):360–364
Patel MK, Joshi M, Mishra A, Jha B (2015) Ectopic expression of SbNHX1 gene in transgenic castor (Ricinus communis L.) enhances salt stress by modulating physiological process. Plant Cell, Tissue Organ Cult 122:477–490
Platten JD, Cotsaftis O, Berthomieu P, Bohnert H, Davenport RJ, Fairbairn DJ, Horie T, Leigh RA, Lin H-X, Luan S (2006) Nomenclature for HKT transporters, key determinants of plant salinity tolerance. Trends Plant Sci 11(8):372–374
Plett D, Safwat G, Gilliham M, Skrumsager Moller I, Roy S, Shirley N, Jacobs A, Johnson A, Tester M (2010) Improved salinity tolerance of rice through cell type-specific expression of AtHKT1;1. PLoS ONE 5(9):e12571
Ren ZH, Gao JP, Li LG, Cai XL, Huang W, Chao DY, Zhu MZ, Wang ZY, Luan S, Lin HX (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37(10):1141–1146
Rubio F, Gassmann W, Schroeder JI (1995) Sodium-driven potassium uptake by the plant potassium transporter HKT1 and mutations conferring salt tolerance. Science 270:1660–1663
Rubio F, Schwarz M, Gassmann W, Schroeder JI (1999) Genetic selection of mutations in the high affinity K+ transporter HKT1 that define functions of a loop site for reduced Na+ permeability and increased Na+ tolerance. J Biol Chem 274(11):6839–6847
Schachtman DP, Schroeder JI (1994) Structure and transport mechanism of a high-affinity potassium uptake transporter from higher plants. Nature 370:655–658
Shukla PS, Agarwal PK, Jha B (2012) Improved salinity tolerance of Arachis hypogaea (L.) by the interaction of halotolerant plant-growth-promoting rhizobacteria. J Plant Growth Regul 31(2):195–206
Wang TT, Ren ZJ, Liu ZQ, Feng X, Guo RQ, Li BG, Li LG, Jing HC (2014) SbHKT1;4, a member of the high-affinity potassium transporter gene family from Sorghum bicolor, functions to maintain optimal Na+/K+ balance under Na+ stress. J Integr Plant Biol 56(3):315–332
Acknowledgments
This work is financially supported by the National Basic Research Program of China (2014CB138202) and the Agricultural Science and Technology Innovation Program (ASTIP) of CAAS.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ren, Z., Liu, Y., Kang, D. et al. Two alternative splicing variants of maize HKT1;1 confer salt tolerance in transgenic tobacco plants. Plant Cell Tiss Organ Cult 123, 569–578 (2015). https://doi.org/10.1007/s11240-015-0861-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-015-0861-9