Abstract
Cancer is caused by the accumulation of inherited and/or acquired alterations in specific genes. The recent decline in the cost of DNA sequencing has allowed tumor sequencing to be conducted on a large scale, which, in turn, has led to an unprecedented understanding of the genetic events that drive neoplasia. This understanding, when integrated with meticulous histologic analyses and with clinical findings, has direct clinical implications. The recent sequencing of all of the major types of cystic and noncystic neoplasms of the pancreas has revealed opportunities for molecular diagnoses and for personalized treatment. This review summarizes the results from these recent studies focusing on the clinical relevance of genomic data.
Similar content being viewed by others
Introduction
It is becoming increasingly clear that cancer is a disease caused by mutations and other alterations to specific genes. This growing perception, combined with the introduction of next-generation sequencing, has led to an unprecedented understanding of the pathogenesis of cancer, and to the opportunity for personalized diagnoses and treatment [1].
In what can be described only as a revolution, the exomes and/or genomes of all of the commonest types of human cancer have been sequenced. In 2007, Wood et al. [2] were the first to sequence the exome of a solid tumor when they sequenced the exomes of a series of well-characterized human breast and colorectal cancers. On the basis in large part of the success of these early sequencing efforts, the International Cancer Genome Consortium and The Cancer Genome Atlas separately launched large multi-institutional efforts to sequence the genomes of 50 major cancer types [3]. These and other studies have revealed that most cancer types are characterized by a small number of genes that are targeted in most tumors of those types, and by a larger number of genes that are altered in only a small percentage of tumors of those types. Vogelstein et al. [1] designated these as “mountains” and “hills” in the cancer genomic landscapes, and unified the mountains and the hills by identifying the common cellular pathways that they target. Vogelstein et al. also made the important distinction between “driver” mutations (those that are functionally significant and promote neoplastic growth) and “passenger” mutations (somatic mutations that are present in a neoplasm but which do not have functional significance). The cancer sequencing revolution has struck tumors of the pancreas, and the results have been dramatic.
All of the major types of tumors of the pancreas, including all of the cystic and all of the solid tumors, have been sequenced [4–11]. The results of these sequencing studies have enormously improved our understanding of pancreatic neoplasia, they have created a foundation for novel research [8, 12–14], and they have significant clinical applications [15–17] (Table 1).
In this review we summarize the salient genetic alterations of the most important pancreatic neoplasms and we emphasize possible clinical applications. A detailed understanding of the gross and microscopic pathologic features of a tumor is important because the pathologic features will guide how we study a disease and because a good understanding of pathologic features can form a foundation for clinical applications. Because of the focus of this review is on genetic changes in pancreatic neoplasms, we will only briefly summarize the individual gross and histopathologic features of each entity.
Pancreatic ductal adenocarcinoma
Pancreatic cancer is the fourth leading cause of cancer death, and it is projected to become the second leading cause of cancer death in the USA by 2030 [18]. In spite of significant cost-intensive research efforts, the 5-year survival rate for patients with pancreatic ductal adenocarcinoma (PDA) remains approximately 5 %, a statistic that has virtually not changed in the past 50 years. PDA is often diagnosed in an advanced stage with primary tumors invading surrounding structures and/or with distant metastases, leaving only palliative treatment options [19].
Resected PDAs are usually solid, firm, diffusely infiltrative and white-yellow [4]. Microscopically, PDAs, by definition, are composed of infiltrating neoplastic glands. Very characteristically, these neoplastic glands elicit an intense desmoplastic reaction, such that much of the tumor is actually composed of nonneoplastic fibroblasts and inflammatory cells [20]. It is important to appreciate the desmoplastic stroma when conducting genetic analyses because if one does not dissect the stroma away from the neoplastic cells, one will end up with samples with a low neoplastic cellularity, making analyses difficult to interpret. As is true for other epithelial neoplasms, invasive PDA appears to arise from histologically well-characterized noninvasive precursor lesions. These include microscopic precursor lesions called pancreatic intraepithelial neoplasia (PanIN) lesions, and larger cystic precursor lesions called intraductal papillary mucinous neoplasms (IPMNs) and mucinous cystic neoplasms (MCNs) [21, 22]. These precursor lesions are important because they represent an opportunity to detect and treat pancreatic neoplasia before it progresses to an incurable invasive carcinoma [23].
The genetic changes in invasive PDAs are complex, with genomes harboring complex karyotypes and multiple copy number alterations often spanning very large genomic regions [6, 24, 25]. At the individual gene level, DNA changes include activating mutations in oncogenes, inactivating mutations in tumor suppressor and caretaker genes, epigenetic modifications, and altered telomere length. The first large sequencing study of invasive PDA, by Jones et al. [6] in 2008, confirmed that an oncogene, KRAS, and three tumor suppressor genes, CDKN2A (also known as P16), TP53, and SMAD4 (also known as DPC4), are targeted in most of these cancers [26]. The other genes targeted less often included MLL3, TGFBR2, ARID1A, and ATM. Other sequencing studies have followed up on the original study by Jones et al., and these have validated the genes and pathways identified by Jones et al. as targeted in PDA, and have suggested other genes that are targeted at lower prevalence. For example, the International Cancer Genome Consortium project suggested that genes in axon guidance pathways (e.g., SLIT/ROBO) may also be targeted in PDAs [25, 27].
One of the major driver mutations in PDA first described by Almoguera et al. [28] in 1988 is KRAS on chromosome arm 12p, which is activated by point mutations in more than 90 % of PDAs. KRAS mutations usually target a “hot spot” at codon 12, but they may also affect codons 13 and 61 [4–6, 29, 30]. Akin to other proteins of the Ras family, KRAS encodes a GTPase that works as an activation/inactivation switch that controls a number of downstream pathways that affect proliferation, differentiation, and cell survival [31–34]. A third of PDAs without a KRAS mutation harbor BRAF mutations, highlighting the importance of Ras signaling for PDA formation [35]. Many of the remaining KRAS wild-type PDAs appear to be microsatellite unstable, and will be discussed later [36].
The most frequently inactivated tumor suppressor gene in PDA is CDKN2A on chromosome arm 9p [37]. CDKN2A is inactivated in 95 % of PDAs: in 40 % by intragenic mutations coupled with loss of the second allele (loss of heterozygosity, LOH), in 40 % by homozygous deletion, and in 15 % by epigenetic silencing [30, 38, 39]. The protein p16 plays a crucial role in cell cycle control as it slows the transition from G1 phase to S phase. Hence, loss of expression of p16 in pancreatic cancers leads to a decreased inhibition of the cell cycle and therefore promotes unrestrained proliferation of the neoplastic cells.
The tumor suppressor gene TP53 on chromosome arm 17p is inactivated in approximately 75 % of PDAs [40]. The protein p53 has a number of important functions that help to prevent neoplastic transformation. The gene is inactivated in PDA primarily through intragenic mutations coupled with LOH of the wild-type allele [30, 41]. A third tumor suppressor gene, SMAD4 (on chromosome arm 18q), is inactivated in 55 % of PDAs. SMAD4 is inactivated by either homozygous deletion or the combination of an intragenic mutation coupled with LOH [42]. The gene product of SMAD4 is a critical component in the transforming growth factor β pathway, regulating among other things the proliferation and production of the extracellular matrix [6, 43].
Many of the genetic changes found in invasive PDA are also present in PanIN lesions. For example, almost all PanIN lesions, even the lowest-grade PanIN-1 lesions, harbor somatic KRAS mutations [44]. In addition, the other genes targeted in invasive PDA, CDKN2A, TP53, and SMAD4, have all been found to be mutated in PanIN lesions, particularly high-grade PanIN lesions [45, 46]. These findings help establish that PanIN lesions can indeed be precursors to invasive PDA. Supporting this conclusion, is the observation in genetically engineered mouse models that activation of Kras in the pancreas produces mouse PanIN lesions [33].
Although most pancreatic cancers appear to be sporadic, a significant fraction have a strong familial component, and some of these familial cancers have unique genetic drivers. Approximately 5–10 % of PDAs arise on the background of a genetic predisposition [47]. The lifetime risk of PDA grows with the number of relatives an individual has with PDA. The risk is doubled in individuals with a single affected first-degree relative [48], culminating in a 32-fold greater risk for individuals with three or more affected first-degree relatives [49]. Although it is clear that there is a strong familial component to pancreatic cancer, a causative germline mutation in a known pancreatic cancer susceptibility gene can be identified in only approximately 20 % of familial cases [50, 51]. The genes that are known to predispose to pancreatic cancer (Table 1) include genes causing hereditary breast–ovarian cancer (caused by germline mutations in BRCA1, BRAC2, or PALB2; with a relative risk of 3.5–10 for BRCA2), familial atypical multiple mole melanoma syndrome (caused by germline mutations in p16/CDKN2A; with a relative risk of 13–22), Peutz–Jeghers syndrome (caused by germline mutations in STK11; with a relative risk of 132), hereditary pancreatitis (caused by germline mutations in PRSS1 or SPINK1; with a relative risk of 50–80), Lynch syndrome (caused by germline mutations in MLH1, MSH2, etc.; with a relative risk of 8.6) [47, 51, 52], and ataxia–telangiectasia (caused by germline mutations in ATM) [7, 53, 54].
The elucidation of the genetic alterations that drive PDA and its variants has had direct clinical implications.
First, genetic alterations can form the basis for screening and early detection. Studies of genetic evolution have been used to estimate that it takes at least 10 years from a first mutation in a pancreatic ductal epithelial cell until the development of a primary invasive cancer [55]. This suggests a reasonably large window for the early detection of curable pancreatic neoplasia [56]. Pancreatic cancer is, however, a relatively rare disease, and screening efforts will have to be focused on populations with an elevated risk of developing the disease [23]. Again, our genetic understanding can help here as well, as individuals with a strong family history of PDA and individuals who carry a known genetic predisposition, because of their quantifiably increased risk, would be the first to benefit from early detection efforts [57, 58]. An understanding of the genetics of PDA and its precursors can also form the basis for the actual tests used to screen for early disease. For example, Kanda et al. [13, 14] demonstrated that molecular screening is possible in secretin-stimulated pancreatic juice samples harvested through noninvasive endoscopy. They found TP53 mutations in pancreatic juice samples collected with the help of endoscopic ultrasonography from patients with histologically proven high-grade dysplasia or PDA [13] as well as GNAS mutations in patients with radiologically suspected IPMNs [14]. These promising methods have to be validated in a larger prospective trial before their introduction in clinical use.
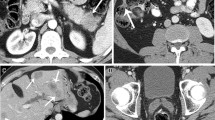
Second, some genetic alternations found in PDA have prognostic implications and may therefore serve as biomarkers. Blackford et al. [59] analyzed sequence data on 39 genes frequently mutated in PDA in a well-characterized series of 90 tumors and correlated the results with patient outcome. They found that loss of SMAD4 in a pancreatic cancer was associated with a shorter overall survival (11.5 months compared with 14.2 months in patients whose tumors had intact SMAD4). Taking these findings one step further, Iacobuzio-Donahue et al. [60] showed that PDAs with inactivated SMAD4 are more likely to metastasize widely than are PDAs with intact SMAD4. Taken together, these findings suggest that SMAD4 status could guide therapeutic approaches in patients with borderline resectable PDA. Patients with tumors that have SMAD4 inactivation may benefit from systemic therapy, as their tumors are more likely to be metastatic (Fig. 1b).
Examples of how genetic biomarkers may contribute to histologic diagnosis and/or clinical outcome prediction. a A colloid adenocarcinoma (hematoxylin and eosin stain; original magnification ×10). These tumors are strongly associated with GNAS codon 201 mutations and an improved outcome compared with tubular ductal adenocarcinomas, which are almost always GNAS wild-type tumors. b A pancreatic ductal adenocarcinoma with loss of SMAD4 (Immunostain for Smad4; original magnification ×40) as often mediated through a somatic mutation. This genetic alteration is usually associated with a particularly poor outcome with rapid widespread metastatic disease
Third, genetic changes may form the basis for individualized therapies. For example, in vitro studies have suggested that genetic testing may even be useful to preselect patients for treatment with chemotherapy agents [61]. Tumors harboring Fanconi anemia pathway gene mutations (such as BRCA1, BRCA2, and PALB2 mutations) and tumors harboring ATM mutations are examples of potential individualized treatment targets. It has been shown that this subgroup of PDAs is more amenable to DNA cross-linking drugs and to poly(ADP-ribose) polymerase inhibitors [54, 62–66]. In addition, PDAs with microsatellite instability, on the basis of experience with colorectal cancers with microsatellite instability, are less likely to be sensitive to 5-fluorouracil than are microsatellite-stable pancreatic cancers [67]. Primary tumor xenografts may help to test the effect of multiple therapeutic agents on a patient’s tumor in order to individualize the drug regimen for that patient, thereby avoiding unnecessary side effects from ineffective drugs administered [68].
Strategies to effectively block oncogenic KRAS signaling are particularly attractive approaches to therapy since virtually all PDAs harbor KRAS mutations. Unfortunately, these efforts have been largely ineffective. Nevertheless, expert groups such as the RAS Initiative led by Frank McCormick at Frederick National Laboratory for Cancer Research have dedicated their work to finding ways to block this apparently “undruggable” pathway.
Cystic lesions of the pancreas
Intraductal papillary mucinous neoplasms
IPMNs are mucin-producing epithelial lesions that arise in the larger pancreatic ducts [69]. IPMNs are, by definition, larger than 1 cm and can be solitary or multifocal [70–72]. IPMNs can involve the main pancreatic duct or they may arise in a branch off from the main duct. Some are combined or mixed, involving both the main duct and branch ducts [73]. IPMNs localized in the main pancreatic duct are more frequently associated with an invasive adenocarcinoma than are branch-duct-type IPMNs [74, 75]. IPMNs can be categorized histologically on the basis of the direction of differentiation of the neoplastic cells into intestinal, gastric-foveolar, pancreatobiliary, and oncocytic subtypes. The epithelial subtype has been shown to correlate with the likelihood that the IPMN harbors high-grade dysplasia or an associated invasive carcinoma [76, 77].
The main clinical problem in treating patients with an IPMN is determining if and when the lesion should be resected. IPMNs that are likely to have high-grade dysplasia or a resectable associated invasive carcinoma should be resected, but many IPMNs will have only low-grade dysplasia and their clinical course can be safely followed without surgery [75, 78, 79].
Previous studies have shown that IPMNs harbor specific genetic alterations, including copy number changes [80] and intragenic mutations [81]. Recent comprehensive sequencing studies have shown that IPMNs harbor an average of 26 nonsynonymous coding mutations, with KRAS as the most frequently mutated gene (approximately 80 %) [8, 9, 82]. Although KRAS is also a hallmark of PDA, two other genetic alterations are highly prevalent in IPMNs (Table 1). In contrast to “common” PDAs (arising in the absence of an associated IPMN), GNAS codon 201 mutations are present in two thirds of IPMNs and inactivating RNF43 mutations can also be seen [82, 83]. Somatic mutations in GNAS lead to an activation of an alpha subunit of a G protein causing increased adenyl cyclase activity and high cyclic AMP production. GNAS mutations appear to be most prevalent in intestinal-type IPMNs and their associated invasive colloid adenocarcinomas [84]. Patients with colloid cancers usually have a more favorable outcome compared with those with a tubular PDA (Fig. 1a). Inactivating mutations in the tumor suppressor gene RNF43 are also found in a subset of IPMNs. The associated protein, a ubiquitin E3 ligase, has been implicated as an inhibitor of Wnt signaling [8]. The four most frequently involved genes in PDA play a role in IPMNs too. As mentioned, somatic KRAS mutations occur in a high proportion of IPMNs, with higher incidence in gastric and pancreatobiliary subtypes and the associated tubular adenocarcinomas [85–88]. Somatic TP53 mutations are almost exclusively found in areas of high-grade dysplasia or invasion, which indicates that this mutation is a relatively late event in carcinogenesis [88–91]. Most noninvasive IPMNs show intact expression of the tumor suppressor gene SMAD4, whereas SMAD4 loss can be seen in invasive carcinomas arising in association with IPMNs, again suggesting that SMAD4 is targeted late in neoplastic progression [92, 93]. Similarly, loss of p16 is very common in invasive carcinoma arising in association with an IPMN, whereas p16 is preserved in most lower-grade IPMNs [93]. Other mutations that can be found in IPMNs include mutations of PIK3CA (approximately 10 %) [88], STK11 (also known as LKB1) (5 % of sporadic IPMNs) [94] and BRAF (2.7 % of IPMNs) [95, 96].
Mucinous cystic neoplasms
MCNs are cyst-forming mucin-secreting epithelial neoplasms with a distinctive ovarian-type stroma [4]. They are almost always found in the tail of the pancreas, and are much commoner in women than in men. In contrast to IPMNs, the cysts of MCNs do not communicate with the pancreatic duct system. MCNs, like IPMNs and PanIN lesions, can be a precursor to invasive adenocarcinoma [21, 72, 97]. Microscopically, MCNs are characterized by a neoplastic mucin-rich columnar epithelium with varying degrees of dysplasia sitting on an ovarian-type stroma composed of spindle-shaped cells [72, 98]. Owing to their predominance in women, their distal location in the pancreas, and their ovarian-type stroma, it has been speculated that MCNs do not originate from pancreatic tissue, but instead arise from cells left behind by the ovaries as they descend past the pancreas embryologically.
Sequencing of the exomes of a series of histologically well-characterized MCNs detected 16 mutations on average in each tumor [8]. Mutations in KRAS are common (approximately 80 %) [99–101], and 40 % of MCNs harbor an inactivating mutation in RNF43. CDKN2A [100], TP53 [101, 102], and SMAD4 [92, 103] can also be targeted in MCNs, especially those with high-grade dysplasia or an associated invasive carcinoma. In contrast to IPMNs, GNAS does not appear to be targeted in MCNs [8, 104].
Serous cystadenomas
Serous cystadenomas (SCAs) are cystic neoplasms that are almost always benign and only extremely rarely behave in a malignant manner [4]. Grossly, most SCAs appear as multilocular or unilocular microcystic lesions. They sometimes have a characteristic central star-shaped scar with calcifications [33]. Microscopically, the well-demarcated cysts have a single layer of cuboidal glycogen-rich epithelium.
Whole-exome sequencing revealed an average of ten nonsynonymous mutations per SCA [8]. All SCAs sequenced harbored at least one region with LOH, predominantly located at chromosome arm 3p. The tumor suppressor gene VHL is located on 3p, and sequencing also revealed intragenic mutations in VHL, helping to establish VHL as a driver of this tumor type [105–107]. The gene product of VHL has an E3 ubiquitin ligase activity responsible for “marking” proteins for subsequent degradation. Patients with von Hippel–Lindau syndrome are predisposed to develop SCAs, as well as a variety of other clear cell neoplasms. The syndrome is caused by a germline (autosomal dominant) mutation of VHL with inactivating alteration or loss of the other functioning allele, further highlighting the critical role of this gene in SCA formation [108]. VHL alterations (intragenic mutations or LOH at 3p) can be used to help identify a cystic tumor as an SCA, and since virtually all SCAs are benign, these patients, if asymptomatic, can be safely followed up and unnecessary surgery can be avoided.
Solid-pseudopapillary neoplasms
Solid-pseudopapillary neoplasms (SPNs) are rare exocrine pancreatic neoplasms predominantly seen in younger female patients [4]. Grossly, they have solid and cystic areas, and the cystic areas are often filled with hemorrhagic debris. SPNs are malignant; 15 % of patients have distant metastases at the time of diagnosis [109–111]. Histologically, the neoplastic cells are characteristically poorly cohesive. They have uniform nuclei, sometimes with a nuclear groove. Other histologic features that can help establish the diagnosis include hyaline globules, foam cells, and cholesterol clefts [112]. Sequencing of SPNs has revealed that they harbor only a few DNA alterations [8, 113]: LOH is rare in SPNs, and they harbor only three somatic mutations on average. A missense mutation in CTNNB1, encoding β-catenin, is found in virtually all SPNs [8, 114]. β-Catenin is a component of the cadherin protein complex with critical function in cell–cell adhesion and gene transcription within Wnt signaling. CTNNB1 mutations inhibit ubiquitination and degradation of β-catenin, leading to an accumulation of β-catenin in the nucleus. In the nucleus, β-catenin induces the expression of its target proteins [115].
Clinical implications
Cystic lesions of the pancreas are increasingly diagnosed owing to an ever-increasing use of modern cross-sectional imaging. At autopsy as many as 25 % of individuals without any pancreas-linked symptoms or diseases have cystic changes in their pancreas [116]. Most of these sometimes minute cysts can now be visualized by high-resolution computed tomography, magnetic resonance imaging, or endoscopic ultrasonography [117–119]. This “epidemic” of pancreatic cysts is, on the one hand, an unprecedented chance to treat pancreatic neoplasia before it progresses to an invasive carcinoma. On the other hand, there is a risk of harmful overtreatment if, for example, a patient undergoes unnecessary surgery for an asymptomatic entirely benign pancreatic cyst. Therefore, it is essential that clinicians have sensitive and specific tools to distinguish benign neoplastic cysts (e.g., SCAs) from cysts with the potential to progress to metastatic cancer (e.g., IPMNs, MCNs, or SPNs). Advances in our understanding of the genetics of cystic neoplasms of the pancreas suggest a new approach to characterizing pancreatic cysts. Fluid can be aspirated from pancreatic cysts at the time of endoscopic ultrasonography. This fluid will contain molecular material shed from the neoplastic cells and therefore will contain the specific DNA alterations that can be used to characterize the cyst type (i.e., IPMN, KRAS, GNAS, and RNF43; MCN, KRAS and RNF43; SCA, VHL and LOH of chromosome 3; SPN, CTNNB1). Additionally, cyst fluid may be evaluated for microRNAs or for mucins. When integrated with clinical findings, such molecular approaches have the potential of fundamentally changing the way cystic neoplasms of the pancreas are managed [120–124].
Pancreatic neuroendocrine tumors
Pancreatic neuroendocrine tumors (PanNETs) are the second most frequently diagnosed primary tumor of the pancreas [125]. Compared with PDA, however, the prognosis is usually much better, with approximately 40 % of affected patients surviving 10 years after diagnosis [126–128]. PanNETs are heterogeneous and clinically can be divided into functioning (syndromic) or nonfunctioning (nonsyndromic) tumors. Among all functioning PanNETs, insulinomas are the commonest subgroup (30–45 %) [127], followed by gastrinomas (16–30 %), VIPomas, glucagonomas (less than 10 % each), and somatostatinomas (less than 5 %) [129]. Most PanNETs occur sporadically, although up to 10 % arise in individuals with a cancer-predisposition syndrome, such as von Hippel–Lindau syndrome (caused by inherited mutations in VHL), tuberous sclerosis (caused by inherited mutations in TSC1 or TSC2), or multiple endocrine neoplasia 1 (caused by inherited mutations in MEN1) [130].
PanNETs are grossly distinct from PDAs. They usually form well-demarcated soft, solid, or rarely cystic neoplasms [4]. Histologically, PanNETs are characterized by epithelial cells with round to oval nuclei containing a “salt and pepper” type chromatin. PanNETs are graded by their proliferative activity as assessed by Ki-67 immunohistochemical labeling. Grade 1 PanNETs have a Ki-67 labeling index of 0–2 %, grade 2 PanNETs have a Ki-67 labeling index of 3–20 %, and grade 3 PanNETs (designated neuroendocrine carcinomas) have a Ki-67 labeling index of more than 20 % [4].
Jiao et al. [10] sought to elucidate the genetics of PanNETs. They sequenced the exomes of ten nonfamilial PanNETs and validated their findings in a series of 58 independent surgically resected PanNETs. They detected a mean of 16 mutations per tumor, and the genes affected included MEN1, DAXX, ATRX, and mechanistic target of rapamycin (mTOR) pathway genes (TSC2, PTEN, and PIK3CA). Intragenic mutations in MEN1 were most frequent and were detected in 44 % of the PanNETs, and 70 % of the PanNETs had LOH in this region. DAXX (25 %) and ATRX (18 %) were also targeted in PanNETs [10]. DAXX and ATRX code for proteins that function in a chromatin remodeling complex. These proteins have a pivotal impact in telomere preservation by incorporating histone variant H3.3 at the telomeric ends of chromosomes [131, 132]. PanNETs with DAXX/ATRX mutations almost always have the “alternative lengthening of telomeres” (ALT+) phenotype [133, 134]. Other genes targeted in PanNETs include genes coding for members of the phosphatidylinositol 3′-kinase (PI3K)–AKT–mTOR pathway. In approximately 16 % of PanNETs, an alterations in PTEN, TSC2, or PIK3CA was found [10]. The PI3K–AKT–mTOR pathway is an important regulator of energy metabolism, cell growth, and cell proliferation. The tumor suppressor gene PTEN encodes a phosphatase that inhibits an unrestrained activation of the mTOR pathway, therefore protecting a cell from uncontrolled proliferation [135]. PIK3CA, on the other hand, is thought be involved in the tumorigenesis of many forms of cancers by coordinating a wide range of cellular activities [136]. In a recent study, Ohki et al. [137] reported that PanNETs often show LOH in the PHLDA3 gene, a genetic change which seems to be associated with a negative outcome and disease progression. Previously, it was shown that the gene product of PHLDA3 acts as a p53-regulated tumor suppressor in PanNETs through an inactivation of the PI3K–AKT–mTOR pathway [138].
The genes targeted in PanNETs suggest a personalized approach to treatment [139, 140]. Since 16 % of PanNETs harbor mutations in genes involved in PI3K–Akt–mTOR signaling, drugs that target the mTOR pathway should be effective in PanNETs with one of these mutations. A variety of mTOR inhibitors have been developed, and everolimus has been successfully used to treat patients with advanced PanNETs; extending the relapse-free survival from 4.6 months with placebo to 11 months in the treated cohort [141–143]. Another promising therapeutic option may be the inhibition of the angiogenesis of the tumor. Here sunitinib, a receptor tyrosine kinase inhibitor, was approved by the Food and Drug Administration for treatment of advanced PanNETs after successful phase III trials [144].
Acinar cell carcinoma
Acinar cell carcinomas (ACCs) are rare but important tumors. The outcome for patients with an ACC is almost as unfavorable as it is for patients with a PDA, and some (15 %) patients develop a dramatic syndrome of metastatic fat necrosis caused by lipase release from the tumor. Grossly, ACCs are often large (up to 10 cm) and relatively soft compared with PDAs [4]. Histologically, they have very little stroma, and the neoplastic cells form small acinar structures or sheets [15, 17]. The nuclei of the neoplastic cells often have single prominent nucleoli.
In 2014 Jiao et al. [11] sequenced the exomes of pancreatic neoplasms with acinar differentiation, and they found that ACCs have dramatic chromosomal instability and genetic heterogeneity. They detected an average of 64 somatic mutations per neoplasm (and this is not considering three outliers), which is higher than for PDAs. Genes implicated in other pancreatic neoplasms were altered in some of the ACCs, including SMAD4 mutations in 25 %, TP53 mutations in 15 %, and GNAS mutations in 10 %, as well as mutations in RNF43 and MEN1 (4 %) [11]. A small number of patients harbored somatic mutations in genes associated with familial pancreatic cancer syndromes. More recent genomic profiling of a large number of ACCs revealed recurrent and potentially actionable genomic alterations, including BRAF and RAF1 fusions, and mutually exclusive DNA repair deficiencies [145].
Marked chromosomal instability may explain the general resistance of most ACCs to traditional systemic therapy regimens [11, 17]. In spite of a vast genetic heterogeneity in ACC, the whole-exome sequencing did identify genetic mutations in 40 % of the cancers that might be amenable for targeted therapies. These include, for example, BRCA2, PALB2, ATM, BAP1, BRAF, and JAK1 mutations [11]. In particular, ACCs with an ATM mutation might be predicted to respond to poly(ADP-ribose) polymerase inhibitors and inhibitors of DNA-dependent protein kinase catalytic subunit [63, 64]. Novel strategies may also be use to try to treat RAF-gene-fusion-positive ACC with mitogen-activated protein/extracellular-signal-regulated kinase kinase inhibitors [145] as previously described in melanomas with BRAF fusions [146].
Pancreatoblastoma
Pancreatoblastomas often arise in the first decade of life, although cases in adults have been described [147]. Pancreatoblastomas have been reported in newborn children with the Beckwith–Wiedemann syndrome. Pancreatoblastomas are aggressive neoplasms with a median overall survival of only 15 months [148]. Histologically, pancreatoblastomas can show multiple directions of differentiation, including acinar, ductal, mesenchymal, neuroendocrine, and primitive embryonic differentiation [4]. By definition, an acinar component is present [147], as are characteristic squamous nests [149].
The hallmark DNA alteration in pancreatoblastomas is LOH at a highly imprinted region on chromosome arm 11p [149]. Most pancreatoblastomas harbor mutations in the Wnt signaling pathway, including inactivating mutations in APC and activating mutations in CTNNB1 [149]. The genetic mutations frequently seen in PDA are rare (KRAS) or absent (TP53, CDKN2A, or SMAD4) [150]. Pancreatoblastomas may overlap in some morphological, immunohistochemical, and clinical features with ACCs. In contrast microsatellite instability is very uncommon in pancreatoblastomas [148, 149].
Owing to the rareness of pancreatoblastomas, no single study could reliably investigate a relation between genetic alternations and factors such as patient survival. No targeted therapies have been shown to be effective to date. Various adjuvant chemotherapy regimens have been applied, and ambiguous results have been published as case reports [151]. As for most pancreatic malignancies, surgical resection is still the gold standard.
Conclusions
All of the major tumor types that arise in the pancreas have been sequenced in recent years. These sequencing studies have provided insight into key mechanisms of pathogenesis and have revealed a number of novel approaches to improving patient care. There are multiple challenges before personalized management based on the genetic signatures of a patient’s tumor will become common practice, but we believe that hope is on the horizon. Well-organized work at multi-institutional and international levels will decrease the “bench-to-bedside” time to clinically use the valuable new data as soon as possible.
Abbreviations
- ACC:
-
Acinar cell carcinoma
- IPMN:
-
Intraductal papillary mucinous neoplasm
- LOH:
-
Loss of heterozygosity
- MCN:
-
Mucinous cystic neoplasm
- mTOR:
-
Mechanistic target of rapamycin
- PanIN:
-
Pancreatic intraepithelial neoplasia
- PanNET:
-
Pancreatic neuroendocrine tumor
- PDA:
-
Pancreatic ductal adenocarcinoma
- PI3K:
-
Phosphatidylinositol 3′-kinase
- SCA:
-
Serous cystadenoma
- SPN:
-
Solid-pseudopapillary neoplasm
- STK11/LKB1:
-
serine-threonine kinase 11/ liver kinase B1
- TGFBR2:
-
transforming growth factor, beta receptor II
- TP53:
-
tumor protein 53
- VEGF:
-
vascular endothelial growth factor
- VHL:
-
von Hippel Lindau
- WHO:
-
World Health Organization
- ZIM2:
-
zinc finger, imprinted 2
References
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA, Kinzler KW. Cancer genome landscapes. Science. 2013;339:1546–58.
Wood L, Parsons DW, Jones S, Lin J, Sjöblom T, Leary RJ, et al. The genomic landscapes of human breast and colorectal cancers. Science. 2007;318:1108–13.
International Cancer Genome Consortium, Hudson TJ, Anderson W, Artez A, Barker AD, Bell C, et al. International network of cancer genome projects. Nature. 2010;464:993–8.
Bosman FT, Carneiro F, Hruban RH, Theise ND, editors. WHO classification of tumours of the digestive system. 4th ed. Lyon: International Agency for Research on Cancer; 2010.
Liang WS, Craig DW, Carpten J, Borad MJ, Demeure MJ, Weiss GJ, et al. Genome-wide characterization of pancreatic adenocarcinoma patients using next generation sequencing. PLoS One. 2012;7:e43192.
Jones S, Zhang X, Parsons DW, Lin JC-H, Leary RJ, Angenendt P, et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science. 2008;321:1801–6.
Jones S, Hruban RH, Kamiyama M, Borges M, Zhang X, Parsons DW, et al. Exomic sequencing identifies PALB2 as a pancreatic cancer susceptibility gene. Science. 2009;324:217.
Wu J, Jiao Y, Dal Molin M, Maitra A, de Wilde RF, wood LD, et al. Whole-exome sequencing of neoplastic cysts of the pancreas reveals recurrent mutations in components of ubiquitin-dependent pathways. Proc Natl Acad Sci U S A. 2011;108:21188–93.
Furukawa T, Kuboki Y, Tanji E, Yoshida S, Hatori T, Yamamoto M, et al. Whole-exome sequencing uncovers frequent GNAS mutations in intraductal papillary mucinous neoplasms of the pancreas. Sci Rep. 2011;1:161.
Jiao Y, Shi C, Edil BH, de Wilde RF, Klimstra DS, Maitra A, et al. DAXX/ATRX, MEN1, and mTOR pathway genes are frequently altered in pancreatic neuroendocrine tumors. Science. 2011;331:1199–203.
Jiao Y, Yonescu R, Offerhaus GJA, Klimstra DS, Maitra A, Eshleman JR, et al. Whole-exome sequencing of pancreatic neoplasms with acinar differentiation. J Pathol. 2014;232:428–35.
Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014;6:224ra24.
Kanda M, Sadakari Y, Borges M, Topazian M, Farrell J, Syngal S, et al. Mutant TP53 in duodenal samples of pancreatic juice from patients with pancreatic cancer or high-grade dysplasia. Clin Gastroenterol Hepatol. 2013;11:719–30.e5.
Kanda M, Knight S, Topazian M, Syngal S, Farrell J, Lee J, et al. Mutant GNAS detected in duodenal collections of secretin-stimulated pancreatic juice indicates the presence or emergence of pancreatic cysts. Gut. 2013;62:1024–33.
Klimstra DS, Heffess CS, Oertel JE, Rosai J. Acinar cell carcinoma of the pancreas. A clinicopathologic study of 28 cases. Am J Surg Pathol. 1992;16:815–37.
Okano K, Suzuki Y. Strategies for early detection of resectable pancreatic cancer. World J Gastroenterol. 2014;20:11230–40.
La Rosa S, Adsay V, Albarello L, Asioli S, Casnedi S, Franzi F, et al. Clinicopathologic study of 62 acinar cell carcinomas of the pancreas: insights into the morphology and immunophenotype and search for prognostic markers. Am J Surg Pathol. 2012;36:1782–95.
Siegel R, Ma J, Zou Z, Jemal A. Cancer statistics, 2014. CA Cancer J Clin. 2014;64:9–29.
Wolfgang CL, Herman JM, Laheru DA, Klein AP, Erdek MA, Fishman EK, et al. Recent progress in pancreatic cancer. CA Cancer J Clin. 2013;63:318–48.
Reid MD, Bagci P, Adsay NV. Histopathologic assessment of pancreatic cancer: does one size fit all? J Surg Oncol. 2013;107:67–77.
Matthaei H, Schulick RD, Hruban RH, Maitra A. Cystic precursors to invasive pancreatic cancer. Nat Rev Gastroenterol Hepatol. 2011;8:141–50.
Maitra A, Fukushima N, Takaori K, Hruban RH. Precursors to invasive pancreatic cancer. Adv Anat Pathol. 2005;12:81–91.
Lennon AM, Wolfgang CL, Canto MI, Klein AP, Herman JM, Goggins M, et al. The early detection of pancreatic cancer: what will it take to diagnose and treat curable pancreatic neoplasia? Cancer Res. 2014;74:3381–9.
Kowalski J, Morsberger LA, Blackford A, Hawkins A, Yeo CJ, Hruban RH, et al. Chromosomal abnormalities of adenocarcinoma of the pancreas: identifying early and late changes. Cancer Genet Cytogenet. 2007;178:26–35.
Biankin AV, Waddell N, Kassahn KS, Gingras M-C, Muthuswamy LB, Johns AL, et al. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature. 2012;491:399–405.
Wood LD, Adsay NV, Hruban RH. Molecular surgical pathology. In: Cheng L, Eble JN, editors. New York: Springer; 2013. p. 17–42. doi:10.1007/978-1-4614-4900-3_2.
Mehlen P, Delloye-Bourgeois C, Chédotal A. Novel roles for Slits and netrins: axon guidance cues as anticancer targets? Nat Rev Cancer. 2011;11:188–97.
Almoguera C, Shibata D, Forrester K, Martin J, Arnheim N, Perucho M. Most human carcinomas of the exocrine pancreas contain mutant c-K-ras genes. Cell. 1988;53:549–54.
Tseng JF, Warshaw AL, Sahani DV, Lauwers GY, Rattner DW, Fernández-del Castillo C. Serous cystadenoma of the pancreas: tumor growth rates and recommendations for treatment. Ann Surg. 2005;242:413–9; discussion 419–21.
Hruban RH, van Mansfeld AD, Offerhaus GJ, van Weering DH, Allison DC, Goodman SN, et al. K-ras oncogene activation in adenocarcinoma of the human pancreas. A study of 82 carcinomas using a combination of mutant-enriched polymerase chain reaction analysis and allele-specific oligonucleotide hybridization. Am J Pathol. 1993;143:545–54.
Vogelstein B, Kinzler KW. Cancer genes and the pathways they control. Nat Med. 2004;10:789–99.
Vakiani E, Solit DB. KRAS and BRAF: drug targets and predictive biomarkers. J Pathol. 2011;223:219–29.
Wood L, Hruban RH. Pathology and molecular genetics of pancreatic neoplasms. Cancer J. 2012;18:492–501.
Hingorani SR, Petricoin EF, Maitra A, Rajapakse V, King C, Jacobetz MA, et al. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell. 2003;4:437–50.
Calhoun ES, Jones JB, Ashfaq R, Adsay V, Baker SJ, Valentine V, et al. BRAF and FBXW7 (CDC4, FBW7, AGO, SEL10) mutations in distinct subsets of pancreatic cancer: potential therapeutic targets. Am J Pathol. 2003;163:1255–60.
Goggins M, Offerhaus GJ, Hilgers W, Griffin CA, Shekher M, Tang D, et al. Pancreatic adenocarcinomas with DNA replication errors (RER+) are associated with wild-type K-ras and characteristic histopathology. Poor differentiation, a syncytial growth pattern, and pushing borders suggest RER+. Am J Pathol. 1998;152:1501–7.
Caldas C, Hahn SA, da Costa LT, Redston MS, Schutte M, Seymour AB, et al. Frequent somatic mutations and homozygous deletions of the p16 (MTS1) gene in pancreatic adenocarcinoma. Nat Genet. 1994;8:27–32.
Attri J, Srinivasan R, Majumdar S, Radotra BD, Wig J. Alterations of tumor suppressor gene p16INK4a in pancreatic ductal carcinoma. BMC Gastroenterol. 2005;5:22.
Li J, Poi MJ, Tsai M-D. Regulatory mechanisms of tumor suppressor P16(INK4A) and their relevance to cancer. Biochemistry. 2011;50:5566–82.
Rozenblum E, Schutte M, Goggins M, Hahn SA, Panzer S, Zahurak M, et al. Tumor-suppressive pathways in pancreatic carcinoma. Cancer Res. 1997;57:1731–4.
Scarpa A, Capelli P, Mukai K, Zamboni G, Oda T, Iacono C, et al. Pancreatic adenocarcinomas frequently show p53 gene mutations. Am J Pathol. 1993;142:1534–43.
Hahn SA, Schutte M, Hoque AT, Moskaluk CA, da Costa LT, Rozenblum E, et al. DPC4, a candidate tumor suppressor gene at human chromosome 18q21.1. Science. 1996;271:350–3.
Iacobuzio-Donahue CA, Song J, Parmiagiani G, Yeo CJ, Hruban RH, Kern SE. Missense mutations of MADH4: characterization of the mutational hot spot and functional consequences in human tumors. Clin Cancer Res. 2004;10:1597–604.
Kanda M, Matthaei H, Wu J, Hong S-M, Yu J, Borges M, et al. Presence of somatic mutations in most early-stage pancreatic intraepithelial neoplasia. Gastroenterology. 2012;142:730–9.
Lüttges J, Galehdari H, Bröcker V, Schwarte-Waldhoff I, Henne-Bruns D, Klöppel G, et al. Allelic loss is often the first hit in the biallelic inactivation of the p53 and DPC4 genes during pancreatic carcinogenesis. Am J Pathol. 2001;158:1677–83.
Wilentz RE, Iacobuzio-Donahue CA, Argani P, McCarthy DM, Parsons JL, Yeo CJ, et al. Loss of expression of Dpc4 in pancreatic intraepithelial neoplasia: evidence that DPC4 inactivation occurs late in neoplastic progression. Cancer Res. 2000;60:2002–6.
Shi C, Hruban RH, Klein AP. Familial pancreatic cancer. Arch Pathol Lab Med. 2009;133:365–74.
Amundadottir LT, Thorvaldsson S, Gudbjartsson DF, Sulem P, Kristjansson K, Arnason S, et al. Cancer as a complex phenotype: pattern of cancer distribution within and beyond the nuclear family. PLoS Med. 2004;1:e65.
Klein AP, Brune KA, Petersen GM, Goggins M, Tersmette AC, Offerhaus GJA, et al. Prospective risk of pancreatic cancer in familial pancreatic cancer kindreds. Cancer Res. 2004;64:2634–8.
Petersen GM, Hruban RH. Familial pancreatic cancer: where are we in 2003? J Natl Cancer Inst. 2003;95:180–1.
Hruban RH, Canto MI, Goggins M, Schulick R, Klein AP. Update on familial pancreatic cancer. Adv Surg. 2010;44:293–311.
Habbe N, Langer P, Sina-Frey M, Bartsch DK. Familial pancreatic cancer syndromes. Endocrinol Metab Clin N Am. 2006;35:417–30.
Slater EP, Langer P, Niemczyk E, Strauch K, Butler J, Habbe N, et al. PALB2 mutations in European familial pancreatic cancer families. Clin Genet. 2010;78:490–4.
Roberts NJ, Jiao Y, Yu J, Kopelovich L, Petersen GM, Bondy ML, et al. ATM mutations in patients with hereditary pancreatic cancer. Cancer Discov. 2012;2:41–6.
Yachida S, Jones S, Bozic I, Antal T, Leary R, Fu B, et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature. 2010;467:1114–7.
Templeton AW, Brentnall TA. Screening and surgical outcomes of familial pancreatic cancer. Surg Clin N Am. 2013;93:629–45.
Wang W, Chen S, Brune KA, Hruban RH, Parmigiani G, Klein AP. PancPRO: risk assessment for individuals with a family history of pancreatic cancer. J Clin Oncol. 2007;25:1417–22.
Norris AL, Roberts NJ, Jones S, Wheelan SJ, Papadopoulos N, Vogelstein B, et al. Familial and sporadic pancreatic cancer share the same molecular pathogenesis. Fam Cancer. 2014. doi:10.1007/s10689-014-9755-y.
Blackford A, Serrano OK, Wolfgang CL, Parmigiani G, Jones S, Zhang X, et al. SMAD4 gene mutations are associated with poor prognosis in pancreatic cancer. Clin Cancer Res. 2009;15:4674–9.
Iacobuzio-Donahue CA, Fu B, Yachida S, Luo M, Abe H, Henderson CM, et al. DPC4 gene status of the primary carcinoma correlates with patterns of failure in patients with pancreatic cancer. J Clin Oncol. 2009;27:1806–13.
Cui Y, Brosnan JA, Blackford AL, Sur S, Hruban RH, Kinzler KW, et al. Genetically defined subsets of human pancreatic cancer show unique in vitro chemosensitivity. Clin Cancer Res. 2012;18:6519–30.
Villarroel MC, Rajeshkumar NV, Garrido-Laguna I, De Jesus-Acosta A, Jones S, Maitra A, et al. Personalizing cancer treatment in the age of global genomic analyses: PALB2 gene mutations and the response to DNA damaging agents in pancreatic cancer. Mol Cancer Ther. 2011;10:3–8.
Williamson CT, Kubota E, Hamill JD, Klimowicz A, Ye R, Muzik H, et al. Enhanced cytotoxicity of PARP inhibition in mantle cell lymphoma harbouring mutations in both ATM and p53. EMBO Mol Med. 2012;4:515–27.
Riabinska A, Daheim M, Herter-Sprie GS, Winkler J, Fritz C, Hallek M, et al. Therapeutic targeting of a robust non-oncogene addiction to PRKDC in ATM-defective tumors. Sci Transl Med. 2013;5:189ra78.
Bryant HE, Schultz N, Thomas HD, Parker KM, Flower D, Lopez E, et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature. 2005;434:913–7.
Fogelman DR, Wolff RA, Kopetz S, Javle M, Bradley C, Mok I, et al. Evidence for the efficacy of iniparib, a PARP-1 inhibitor, in BRCA2-associated pancreatic cancer. Anticancer Res. 2011;31:1417–20.
Collura A, Lagrange A, Svrcek M, Marisa L, Buhard O, Guilloux A, et al. Patients with colorectal tumors with microsatellite instability and large deletions in HSP110 T17 have improved response to 5-fluorouracil–based chemotherapy. Gastroenterology. 2014;146:401–11.
Kamiyama H, Rauenzahn S, Shim JS, Karikari CA, Feldmann G, Hua L, et al. Personalized chemotherapy profiling using cancer cell lines from selectable mice. Clin Cancer Res. 2013;19:1139–46.
Matthaei H, Norris AL, Tsiatis AC, Olino K, Hong S-M, Dal Molin M, et al. Clinicopathological characteristics and molecular analyses of multifocal intraductal papillary mucinous neoplasms of the pancreas. Ann Surg. 2012;255:326–33.
Hruban RH, Takaori K, Klimstra DS, Adsay NV, Albores-Saavedra J, Biankin AV, et al. An illustrated consensus on the classification of pancreatic intraepithelial neoplasia and intraductal papillary mucinous neoplasms. Am J Surg Pathol. 2004;28:977–87.
Schmitz-Winnenthal FH, Z’graggen K, Volk C, Schmied BM, Büchler MW. Intraductal papillary mucinous tumors of the pancreas. Curr Gastroenterol Rep. 2003;5:133–40.
Basturk O, Coban I, Adsay NV. Pancreatic cysts: pathologic classification, differential diagnosis, and clinical implications. Arch Pathol Lab Med. 2009;133:423–38.
Tanaka M, Kobayashi K, Mizumoto K, Yamaguchi K. Clinical aspects of intraductal papillary mucinous neoplasm of the pancreas. J Gastroenterol. 2005;40:669–75.
Sugiyama M, Izumisato Y, Abe N, Masaki T, Mori T, Atomi Y. Predictive factors for malignancy in intraductal papillary-mucinous tumours of the pancreas. Br J Surg. 2003;90:1244–9.
Sohn TA, Yeo CJ, Cameron JL, Hruban RH, Fukushima N, Campbell KA, et al. Intraductal papillary mucinous neoplasms of the pancreas: an updated experience. Ann Surg. 2004;239:788–97; discussion 797–9.
Paini M, Crippa S, Partelli S, Scopelliti F, Tamburrino D, Baldoni A, et al. Molecular pathology of intraductal papillary mucinous neoplasms of the pancreas. World J Gastroenterol. 2014;20:10008–23.
Adsay NV, Merati K, Basturk O, Iacobuzio-Donahue C, Levi E, Cheng JD, et al. Pathologically and biologically distinct types of epithelium in intraductal papillary mucinous neoplasms: delineation of an “intestinal” pathway of carcinogenesis in the pancreas. Am J Surg Pathol. 2004;28:839–48.
Tanaka M, Fernández-del Castillo C, Adsay V, Chari S, Falconi M, Jang J-Y, et al. International consensus guidelines for the management of IPMN and MCN of the pancreas. Pancreatology. 2012;2012(12):183–97.
Fritz S, Klauss M, Bergmann F, Hackert T, Hartwig W, Strobel O, et al. Small (Sendai negative) branch-duct IPMNs: not harmless. Ann Surg. 2012;256:313–20.
Fritz S, Fernández-del Castillo C, Mino-Kenudson M, Crippa S, Deshpande V, Lauwers GY, et al. Global genomic analysis of intraductal papillary mucinous neoplasms of the pancreas reveals significant molecular differences compared to ductal adenocarcinoma. Ann Surg. 2009;249:440–7.
Schönleben F, Qiu W, Bruckman KC, Ciau NT, Li X, Lauerman MH, et al. BRAF and KRAS gene mutations in intraductal papillary mucinous neoplasm/carcinoma (IPMN/IPMC) of the pancreas. Cancer Lett. 2007;249:242–8.
Wu J, Matthaei H, Maitra A, Dal Molin M, Wood LD, Eshleman JR, et al. Recurrent GNAS mutations define an unexpected pathway for pancreatic cyst development. Sci Transl Med. 2011;3:92ra66.
Amato E, Molin MD, Mafficini A, Yu J, Malleo G, Rusev B, et al. Targeted next-generation sequencing of cancer genes dissects the molecular profiles of intraductal papillary neoplasms of the pancreas. J Pathol. 2014;233:217–27.
Dal Molin M, Matthaei H, Wu J, Blackford A, Debeljak M, Rezaee N, et al. Clinicopathological correlates of activating GNAS mutations in intraductal papillary mucinous neoplasm (IPMN) of the pancreas. Ann Surg Oncol. 2013;20:3802–8.
Kim ST, Lim DH, Jang K-T, Lim T, Lee J, Choi Y-L, et al. Impact of KRAS mutations on clinical outcomes in pancreatic cancer patients treated with first-line gemcitabine-based chemotherapy. Mol Cancer Ther. 2011;10:1993–9.
Rachakonda PS, Bauer AS, Xie H, Campa D, Rizzato C, Canzian F, et al. Somatic mutations in exocrine pancreatic tumors: association with patient survival. PLoS One. 2013;8:e60870.
Yoshizawa K, Nagai H, Sakurai S, Hironaka M, Morinaga S, Saitoh K, et al. Clonality and K-ras mutation analyses of epithelia in intraductal papillary mucinous tumor and mucinous cystic tumor of the pancreas. Virchows Arch. 2002;441:437–43.
Lubezky N, Ben-Haim M, Marmor S, Brazowsky E, Rechavi G, Klausner JM, et al. High-throughput mutation profiling in intraductal papillary mucinous neoplasm (IPMN). J Gastrointest Surg. 2011;15:503–11.
Wada K. p16 and p53 gene alterations and accumulations in the malignant evolution of intraductal papillary-mucinous tumors of the pancreas. J Hepatobiliary Pancreat Surg. 2002;9:76–85.
Kawahira H, Kobayashi S, Kaneko K, Asano T, Ochiai T. p53 protein expression in intraductal papillary mucinous tumors (IPMT) of the pancreas as an indicator of tumor malignancy. Hepatogastroenterology. 2000;47:973–7.
Abe K, Suda K, Arakawa A, Yamasaki S, Sonoue H, Mitani K, et al. Different patterns of p16INK4A and p53 protein expressions in intraductal papillary-mucinous neoplasms and pancreatic intraepithelial neoplasia. Pancreas. 2007;34:85–91.
Iacobuzio-Donahue CA, Klimstra DS, Adsay NV, Wilentz RE, Argani P, Sohn TA, et al. Dpc-4 protein is expressed in virtually all human intraductal papillary mucinous neoplasms of the pancreas: comparison with conventional ductal adenocarcinomas. Am J Pathol. 2000;157:755–61.
Biankin AV, Biankin SA, Kench JG, Morey AL, Lee C-S, Head DR, et al. Aberrant p16INK4A and DPC4/Smad4 expression in intraductal papillary mucinous tumours of the pancreas is associated with invasive ductal adenocarcinoma. Gut. 2002;50:861–8.
Sahin F, Maitra A, Argani P, Sato N, Maehara N, Montgomery E, et al. Loss of Stk11/Lkb1 expression in pancreatic and biliary neoplasms. Mod Pathol. 2003;16:686–91.
Schönleben F, Allendorf JD, Qiu W, Li X, Ho DJ, Ciau NT, et al. Mutational analyses of multiple oncogenic pathways in intraductal papillary mucinous neoplasms of the pancreas. Pancreas. 2008;36:168–72.
Chadwick B, Willmore-Payne C, Tripp S, Layfield LJ, Hirschowitz S, Holden J. Histologic, immunohistochemical, and molecular classification of 52 IPMNs of the pancreas. Appl Immunohistochem Mol Morphol. 2009;17:31–9.
Hruban RH, Maitra A, Kern SE, Goggins M. Precursors to pancreatic cancer. Gastroenterol Clin N Am. 2007;36:831–49.
Wilentz RE, Albores-Saavedra J, Hruban RH. Mucinous cystic neoplasms of the pancreas. Semin Diagn Pathol. 2000;17:31–42.
Ghiorzo P. Genetic predisposition to pancreatic cancer. World J Gastroenterol. 2014;20:10778–89.
Kim SG, Wu T-T, Lee JH, Yun YK, Issa J-P, Hamilton SR, et al. Comparison of epigenetic and genetic alterations in mucinous cystic neoplasm and serous microcystic adenoma of pancreas. Mod Pathol. 2003;16:1086–94.
Jimenez RE, Warshaw AL, Z’graggen K, Hartwig W, Taylor DZ, Compton CC, et al. Sequential accumulation of K-ras mutations and p53 overexpression in the progression of pancreatic mucinous cystic neoplasms to malignancy. Ann Surg. 1999;230:501–9; discussion 509–11.
Sorio C, Capelli P, Lissandrini D, Moore PS, Balzarini P, Falconi M, et al. Mucinous cystic carcinoma of the pancreas: a unique cell line and xenograft model of a preinvasive lesion. Virchows Arch. 2005;446:239–45.
Lüttges J, Feyerabend B, Buchelt T, Pacena M, Klöppel G. The mucin profile of noninvasive and invasive mucinous cystic neoplasms of the pancreas. Am J Surg Pathol. 2002;26:466–71.
Singhi AD, Nikiforova MN, Fasanella KE, McGrath KM, Pai RK, Ohori NP, et al. Preoperative GNAS and KRAS testing in the diagnosis of pancreatic mucinous cysts. Clin Cancer Res. 2014;20:4381–9.
Moore PS, Zamboni G, Brighenti A, Lissandrini D, Antonello D, Capelli P, et al. Molecular characterization of pancreatic serous microcystic adenomas: evidence for a tumor suppressor gene on chromosome 10q. Am J Pathol. 2001;158:317–21.
Moore PS, Missiaglia E, Antonello D, Zamò A, Zamboni G, Corleto V, et al. Role of disease-causing genes in sporadic pancreatic endocrine tumors: MEN1 and VHL. Genes Chromosomes Cancer. 2001;32:177–81.
Mohr VH, Vortmeyer AO, Zhuang Z, Libutti SK, Walther MM, Choyke PL, et al. Histopathology and molecular genetics of multiple cysts and microcystic (serous) adenomas of the pancreas in von Hippel-Lindau patients. Am J Pathol. 2000;157:1615–21.
Haddad NMN, Cavallerano JD, Silva PS. Von Hippel-Lindau disease: a genetic and clinical review. Semin Ophthalmol. 2013;28:377–86.
Yang F, Fu D-L, Jin C, Long J, Yu X-J, Xu J, et al. Clinical experiences of solid pseudopapillary tumors of the pancreas in China. J Gastroenterol Hepatol. 2008;23:1847–51.
Kang CM, Kim KS, Choi JS, Kim H, Lee WJ, Kim BR. Solid pseudopapillary tumor of the pancreas suggesting malignant potential. Pancreas. 2006;32:276–80.
Tang LH, Aydin H, Brennan MF, Klimstra DS. Clinically aggressive solid pseudopapillary tumors of the pancreas: a report of two cases with components of undifferentiated carcinoma and a comparative clinicopathologic analysis of 34 conventional cases. Am J Surg Pathol. 2005;29:512–9.
Meriden Z, Shi C, Edil BH, Ellison T, Wolfgang CL, Cornish TC, et al. Hyaline globules in neuroendocrine and solid-pseudopapillary neoplasms of the pancreas: a clue to the diagnosis. Am J Surg Pathol. 2011;35:981–8.
Reid MD, Saka B, Balci S, Goldblum AS, Adsay NV. Molecular genetics of pancreatic neoplasms and their morphologic correlates: an update on recent advances and potential diagnostic applications. Am J Clin Pathol. 2014;141:168–80.
Abraham SC, Wu T-T, Hruban RH, Lee JH, Yeo CJ, Conlon K, et al. Genetic and immunohistochemical analysis of pancreatic acinar cell carcinoma: frequent allelic loss on chromosome 11p and alterations in the APC/β-catenin pathway. Am J Pathol. 2002;160:953–62.
Kim M-J, Jang S-J, Yu E. Loss of E-cadherin and cytoplasmic-nuclear expression of β-catenin are the most useful immunoprofiles in the diagnosis of solid-pseudopapillary neoplasm of the pancreas. Hum Pathol. 2008;39:251–8.
Kimura W, Nagai H, Kuroda A, Muto T, Esaki Y. Analysis of small cystic lesions of the pancreas. Int J Pancreatol. 1995;18:197–206.
Laffan TA, Horton KM, Klein AP, Berlanstein B, Siegelman SS, Kawamoto S, et al. Prevalence of unsuspected pancreatic cysts on MDCT. AJR Am J Roentgenol. 2008;191:802–7.
Lee KS, Sekhar A, Rofsky NM, Pedrosa I. Prevalence of incidental pancreatic cysts in the adult population on MR imaging. Am J Gastroenterol. 2010;105:2079–84.
Spinelli KS, Fromwiller TE, Daniel RA, Kiely JM, Nakeeb A, Komorowski RA, et al. Cystic pancreatic neoplasms: observe or operate. Ann Surg. 2004;239:651–7; discussion 657–9.
Matthaei H, Wylie D, Lloyd MB, Dal Molin M, Kemppainen J, Mayo SC, et al. miRNA biomarkers in cyst fluid augment the diagnosis and management of pancreatic cysts. Clin Cancer Res. 2012;18:4713–24.
Caponi S, Funel N, Frampton AE, Mosca F, Santarpia L, Van der Velde AG, et al. The good, the bad and the ugly: a tale of miR-101, miR-21 and miR-155 in pancreatic intraductal papillary mucinous neoplasms. Ann Oncol. 2013;24:734–41.
Hollingsworth MA, Swanson BJ. Mucins in cancer: protection and control of the cell surface. Nat Rev Cancer. 2004;4:45–60.
Ringel J, Löhr M. The MUC gene family: their role in diagnosis and early detection of pancreatic cancer. Mol Cancer. 2003;2:9.
Stelow EB, Stanley MW, Bardales RH, Mallery S, Lai R, Linzie BM, et al. Intraductal papillary-mucinous neoplasm of the pancreas. The findings and limitations of cytologic samples obtained by endoscopic ultrasound-guided fine-needle aspiration. Am J Clin Pathol. 2003;120:398–404.
Yao JC, Hassan M, Phan A, Dagohoy C, Leary C, Mares JE, et al. One hundred years after “carcinoid”: epidemiology of and prognostic factors for neuroendocrine tumors in 35,825 cases in the United States. J Clin Oncol. 2008;26:3063–72.
Halfdanarson TR, Rabe KG, Rubin J, Petersen GM. Pancreatic neuroendocrine tumors (PNETs): incidence, prognosis and recent trend toward improved survival. Ann Oncol. 2008;19:1727–33.
Halfdanarson TR, Rubin J, Farnell MB, Grant CS, Petersen GM. Pancreatic endocrine neoplasms: epidemiology and prognosis of pancreatic endocrine tumors. Endocr Relat Cancer. 2008;15:409–27.
Ekeblad S, Skogseid B, Dunder K, Öberg K, Eriksson B. Prognostic factors and survival in 324 patients with pancreatic endocrine tumor treated at a single institution. Clin Cancer Res. 2008;14:7798–803.
de Wilde RF, Edil BH, Hruban RH, Maitra A. Well-differentiated pancreatic neuroendocrine tumors: from genetics to therapy. Nat Rev Gastroenterol Hepatol. 2012;9:199–208.
Jensen RT, Berna MJ, Bingham DB, Norton JA. Inherited pancreatic endocrine tumor syndromes: advances in molecular pathogenesis, diagnosis, management, and controversies. Cancer. 2008;113:1807–43.
Elsaesser SJ, Allis CD. HIRA and Daxx constitute two independent histone H3.3-containing predeposition complexes. Cold Spring Harb Symp Quant Biol. 2010;75:27–34.
Goldberg AD, Banaszynski LA, Noh K-M, Lewis PW, Elsaesser SJ, Stadler S, et al. Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell. 2010;140:678–91.
Heaphy CM, de Wilde RF, Jiao Y, Klein AP, Edil BH, Shi C, et al. Altered telomeres in tumors with ATRX and DAXX mutations. Science. 2011;333:425.
Heaphy CM, Subhawong AP, Hong S-M, Goggins MG, Montgomery EA, Gabrielson E, et al. Prevalence of the alternative lengthening of telomeres telomere maintenance mechanism in human cancer subtypes. Am J Pathol. 2011;179:1608–15.
Hay N. The Akt-mTOR tango and its relevance to cancer. Cancer Cell. 2005;8:179–83.
Samuels Y, Wang Z, Bardelli A, Silliman N, Ptak J, Szabo S, et al. High frequency of mutations of the PIK3CA gene in human cancers. Science. 2004;304:554.
Ohki R, Saito K, Chen Y, Kawase T, Hiraoka N, Saigawa R, et al. PHLDA3 is a novel tumor suppressor of pancreatic neuroendocrine tumors. Proc Natl Acad Sci U S A. 2014;111:E2404–13.
Kawase T, Ohki R, Shibata T, Tsutsumi S, Kamimura N, Inazawa J, et al. PH domain-only protein PHLDA3 is a p53-regulated repressor of Akt. Cell. 2009;136:535–50.
Fendrich V, Bartsch DK. Surgical treatment of gastrointestinal neuroendocrine tumors. Langenbecks Arch Surg. 2011;396:299–311.
Panzuto F, Boninsegna L, Fazio N, Campana D, Pia Brizzi M, Capurso G, et al. Metastatic and locally advanced pancreatic endocrine carcinomas: analysis of factors associated with disease progression. J Clin Oncol. 2011;29:2372–7.
Yao JC, Lombard-Bohas C, Baudin E, Kvols LK, Rougier P, Ruszniewski P, et al. Daily oral everolimus activity in patients with metastatic pancreatic neuroendocrine tumors after failure of cytotoxic chemotherapy: a phase II trial. J Clin Oncol. 2010;28:69–76.
Yao JC, Phan AT, Chang DZ, Wolff RA, Hess K, Gupta S, et al. Efficacy of RAD001 (everolimus) and octreotide LAR in advanced low- to intermediate-grade neuroendocrine tumors: results of a phase II study. J Clin Oncol. 2008;26:4311–8.
Yao JC, Shah MH, Ito T, Bohas CL, Wolin EM, Van Cutsem E, et al. Everolimus for advanced pancreatic neuroendocrine tumors. N Engl J Med. 2011;364:514–23.
Raymond E, Dahan L, Raoul J-L, Bang Y-J, Borbath I, Lombard-Bohas C, et al. Sunitinib malate for the treatment of pancreatic neuroendocrine tumors. N Engl J Med. 2011;364:501–13.
Chmielecki J, Hutchinson KE, Frampton GM, Chalmers ZR, Johnson A, Shi C, et al. Comprehensive genomic profiling of pancreatic acinar cell carcinomas identifies recurrent RAF fusions and frequent inactivation of DNA repair genes. Cancer Discov. 2014;4;1398–405.
Hutchinson KE, Lipson D, Stephens PJ, Otto G, Lehmann BD, Lyle PL, et al. BRAF fusions define a distinct molecular subset of melanomas with potential sensitivity to MEK inhibition. Clin Cancer Res. 2013;19:6696–702.
Klimstra DS, Wenig BM, Adair CF, Heffess CS. Pancreatoblastoma. A clinicopathologic study and review of the literature. Am J Surg Pathol. 1995;19:1371–89.
Salman B, Brat G, Yoon Y-S, Hruban RH, Singhi AD, Fishman EK, et al. The diagnosis and surgical treatment of pancreatoblastoma in adults: a case series and review of the literature. J Gastrointest Surg. 2013;17:2153–61.
Abraham SC, Wu TT, Klimstra DS, Finn LS, Lee JH, Yeo CJ, et al. Distinctive molecular genetic alterations in sporadic and familial adenomatous polyposis-associated pancreatoblastomas : frequent alterations in the APC/β-catenin pathway and chromosome 11p. Am J Pathol. 2001;159:1619–27.
Moore PS, Orlandini S, Zamboni G, Capelli P, Rigaud G, Falconi M, et al. Pancreatic tumours: molecular pathways implicated in ductal cancer are involved in ampullary but not in exocrine nonductal or endocrine tumorigenesis. Br J Cancer. 2001;84:253–62.
Cavallini A, Falconi M, Bortesi L, Crippa S, Barugola G, Butturini G. Pancreatoblastoma in adults: a review of the literature. Pancreatology. 2009;9:73–80.
Acknowledgments
This work was supported by a BONFOR research grant (O-1120051) provided to Hanno Matthaei by the University Medical Center, University of Bonn, Bonn, Germany, and by National Institutes of Health grant CA62924.
Conflict of interest
Ralph H. Hruban receives royalty payments from Myriad Genetics for the invention linking PALB2 to familial pancreatic cancer. This is managed by Johns Hopkins University.
Author information
Authors and Affiliations
Corresponding author
Additional information
H. Matthaei and A. Semaan contributed equally.
Rights and permissions
About this article
Cite this article
Matthaei, H., Semaan, A. & Hruban, R.H. The genetic classification of pancreatic neoplasia. J Gastroenterol 50, 520–532 (2015). https://doi.org/10.1007/s00535-015-1037-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-015-1037-4