Abstract
Key message
Genetically stable deletion lines of Agropyron cristatum chromosome 6P in common wheat background were generated, which allowed for physical mapping of 255 6P-specific STS markers and leaf rust resistance gene(s).
Abstract
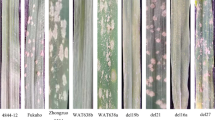
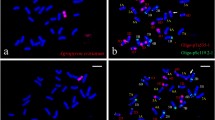
Chromosomal deletion lines are valuable tools for gene discovery and localization. The chromosome 6P of Agropyron cristatum (2n = 4x = 28, PPPP) confers many desirable agronomic traits to common wheat, such as higher grain number per spike, multiple fertile tiller number, and enhanced resistance to certain diseases. Although many elite genes from A. cristatum have been identified, their chromosomal locations were largely undetermined due to the lack of A. cristatum 6P deletion lines. In this study, various A. cristatum 6P deletion lines were developed using a wheat–A. cristatum 6P disomic addition line 4844-12 subjected to 60Co-γ irradiation as well as an Aegilops cylindrica gametocidal chromosome. Twenty-six genetically stable A. cristatum 6P deletion lines in the genetic background of common wheat were obtained, and their genetic constitutions were elucidated by genomic in situ hybridization (GISH) and sequence-tagged site (STS) markers specific to A. cristatum chromosome 6P. Moreover, 255 novel chromosome 6P-specific STS markers were physically mapped to 14 regions of chromosome 6P. Field evaluation of leaf rust resistance of various deletion lines and BC1F2 populations indicated that the A.cristatum chromosome 6P-originated leaf rust resistance gene(s) was located in the region 6PS-0.81-1.00. This study will provide not only useful tools for characterization and utilization of wheat materials with alien chromosomal segments, but also novel wheat germplasms potentially valuable in wheat breeding and improvement.







Similar content being viewed by others
References
Adamski NM, Bush MS, Simmonds J, Turner AS, Mugford SG, Jones A, Findlay K, Pedentchouk N, Wettstein-Knowles P, Uauy C (2013) The Inhibitor of wax 1 locus (Iw1) prevents formation of β-and OH-β-diketones in wheat cuticular waxes and maps to a sub-cM interval on chromosome arm 2BS. Plant J 74:989–1002
Al-Kaff N, Knight E, Bertin I, Foote T, Hart N, Griffiths S, Moore G (2008) Detailed dissection of the chromosomal region containing the Ph1 locus in wheat Triticum aestivum: with deletion mutants and expression profiling. Ann Bot 101:863–872
Ashida T, Nasuda S, Sato K, Endo TR (2007) Dissection of barley chromosome 5H in common wheat. Genes Genet Syst 82:123–133
Basnet BR, Singh RP, Ibrahim AMH, Herrera-Foessel SA, Huerta-Espino J, Lan C, Rudd JC (2014) Characterization of Yr54 and other genes associated with adult plant resistance to yellow rust and leaf rust in common wheat Quaiu 3. Mol breed 33:385–399
Belova T, Grønvold L, Kumar A, Kianian S, He XY, Lillemo M, Springer NM, Lien S, Olsen O-A, Sandve SR (2014) Utilization of deletion bins to anchor and order sequences along the wheat 7B chromosome. Theor Appl Genet 127:2029–2040
Chen XM, Line RF (1992) Inheritance of stripe rust resistance in wheat cultivars used to differentiate races of Puccinia striiformis in North America. Phytopathology 82:633–637
Chen XM, Line RF (1995) Gene action in wheat cultivars for durable, high-temperature, adult-plant resistance and interaction with race-specific, seedling resistance to Puccinia striiformis. Phytopathology 85:567–572
Dewey DR (1984) The genomic system of classification as a guide to intergeneric hybridization with the perennial Triticeae. In: Gustafson JP (ed) Gene manipulation in plant improvement. Plenum Press, New York, pp 209–279
Dong YC, Zhou RH, Xu SJ, Li LH, Cauderon Y, Wang RRC (1992) Desirable characteristics in perennial Triticeae collected in China for wheat improvement. Hereditas 116:175–178
Dundas IS, Frappell DE, Crack DM, Fisher JM (2001) Deletion mapping of a nematode resistance gene on rye chromosome 6R in wheat. Crop Sci 41:1771–1778
Endo TR, Gill BS (1996) The deletion stocks of common wheat. J Hered 87:295–307
Friebe B, Kynast RG, Gill BS (2000) Gametocidal factor-induced structural rearrangements in rye chromosomes added to common wheat. Chromosome Res 8:501–511
Fu BS, Chen Y, Li N, Ma HQ, Kong ZX, Zhang LX, Jia HY, Ma ZQ (2013) pmX: a recessive powdery mildew resistance gene at the Pm4 locus identified in wheat landrace Xiaohongpi. Theor Appl Genet 126:913–921
Gustafson G, Shaner G (1982) Influence of plant age on the expression of slow-mildewing resistance in wheat. Phytopathology 72:746–749
Han HM, Bai L, Su JJ, Zhang JP, Song LQ, Gao AN, Yang XM, Li XQ, Liu WH, Li LH (2014) Genetic rearrangements of six wheat–Agropyron cristatum 6P addition lines revealed by molecular markers. PLoS One 9:e91066
Han HM, Zhang YX, Liu WH, Hu ZM, Li LH (2015) Degenerate oligonucleotide primed-polymerase chain reaction-based chromosome painting of P genome chromosomes in Agropyron cristatum and wheat–A. cristatum addition lines. Crop Sci 55:1–8
Hautea RA, Coffman WR, Sorrells ME, Bergstrom GC (1987) Inheritance of partial resistance to powdery mildew in spring wheat. Theor Appl Genet 73:609–615
Herrera-Foessel SA, Singh RP, Huerta-Espino J, Rosewarne GM, Periyannan SK, Viccars L, Calvo-Salazar V, Lan C, Lagudah ES (2012) Lr68: a new gene conferring slow rusting resistance to leaf rust in wheat. Theor Appl Genet 124:1475–1486
Hossain KG, Jackson SA, Kianian SF (2012) Genome structure and chromosome function. In: Bass HW, Birchler JA (eds) Plant cytogenetics. Springer, pp 37–58
Huang XQ, Röder MS (2011) High-density genetic and physical bin mapping of wheat chromosome 1D reveals that the powdery mildew resistance gene Pm24 is located in a highly recombinogenic region. Genetica 139:1179–1187
Joshi GP, Nasuda S, Endo TR (2011) Dissection and cytological mapping of barley chromosome 2H in the genetic background of common wheat. Genes Genet Syst 86:231–248
Khan MH, Bukhari A, Dar ZA, Rizvi SM (2013) Status and strategies in breeding for rust resistance in wheat. Agric Sci 4:292–301
Kojima T, Habu Y, Iida S, Ogihara Y (2000) Direct isolation of differentially expressed genes from a specific chromosome region of common wheat: application of the amplified fragment length polymorphism-based mRNA fingerprinting (AMF) method in combination with a deletion line of wheat. Mol Gen Genet 263:635–641
Li LH, Dong YC, Zhou RH, Li XQ, Li P (1995) Cytogenetics and self-fertility of hybrids between Triticum aestivum L. and Agropyron cristatum (L.) Gaertn. Acta Genetica Sinica 22:109–114
Li LH, Li XQ, Li P, Dong YC, Zhao GS (1997) Establishment of wheat–Agropyron cristatum alien addition lines. I. Cytology of F3, F2BC1, BC4, and BC3F1 progenies. Acta Genetica Sinica 24:154–159
Li LH, Yang XM, Zhou Rh, Li XQ, Dong YC (1998) Establishment of wheat–Agropyron cristatum alien addition lines II. Identification of alien chromosomes and analysis of development approaches. Acta Genetica Sinica 25:538–544
Ling HQ, Zhao SC, Liu DC, Wang JY et al (2013) Draft genome of the wheat A-genome progennitorr Triticum urartu. Nature 496:87–90
Liu WH, Luan Y, Wang JC, Wang XG, Su JJ, Zhang JP, Yang ZM, Gao AN, Li LH (2010) Production and identification of wheat–Agropyron cristatum (1·4P) alien translocation lines. Genome 53:472–481
Lu YQ, Wu XY, Yao MM, Zhang JP, Liu WH, Yang XM, Li XQ, Du J, Gao AN, Li LH (2015) Genetic mapping of a putative Agropyron cristatum-derived powdery mildew resistance gene by a combination of bulked segregant analysis and single nucleotide polymorphism array. Mol Breed 35:1–13
Luan Y, Wang XG, Liu WH, Li CY, Zhang JP, Gao AN, Wang YD, Yang XM, Li LH (2010) Production and identification of wheat–Agropyron cristatum 6P translocation lines. Planta 232:501–510
McIntosh RA, Wellings CR, Park RF (1995) Wheat rusts: an atlas of resistance genes. CSIRO, East Melbourne, pp 29–82
Nasuda S, Hudakova S, Schubert I, Houben A, Endo TR (2005a) Stable barley chromosomes without centromeric repeats. Proc Natl Acad Sci USA 102:9842–9847
Nasuda S, Kikkawa Y, Ashida T, Islam AR, Sato K, Endo TR (2005b) Chromosomal assignment and deletion mapping of barley EST markers. Genes Genet Syst 80:357–366
Ochoa V, Madrid E, Said M, Rubiales D, Cabrera A (2015) Molecular and cytogenetic characterization of a common wheat–Agropyron cristatum chromosome translocation conferring resistance to leaf rust. Euphytica 201:89–95
Pu J, Wang Q, Shen YF, Zhuang LF, Li CX, Tan MF, Bie TD, Chu CG, Qi ZJ (2015) Physical mapping of chromosome 4J of Thinopyrum bessarabicum using gamma radiation-induced aberrations. Theor Appl Genet 128:1319–1328
Qayoum A, Line RF (1985) High-temperature, adult-plant resistance to stripe rust of wheat. Phytopathology 75:1121–1125
Qi LL, Gill BS (2001) High-density physical maps reveal that the dominant male-sterile gene Ms3 is located in a genomic region of low recombination in wheat and is not amenable to map-based cloning. Theor Appl Genet 103:998–1006
Qi LL, Echalier B, Chao S et al (2004) A chromosome bin map of 16,000 expressed sequence tag loci and distribution of genes among the three genomes of polyploid wheat. Genetics 168:701–712
Raupp WJ, Friebe B, Gill BS (1995) Suggested guidelines for the nomenclature and abbreviation of the genetic stocks of wheat and its relatives. Wheat Inform Serv 81:50–55
Roelfs AP, Singh RP, Saari EE (1992) Rust diseases of wheat: concepts and methods of disease management. CIMMYT, Mexico
Sakai K, Nasuda S, Sato K, Endo TR (2009) Dissection of barley chromosome 3H in common wheat and a comparison of 3H physical and genetic maps. Genes Genet Syst 84:25–34
Sakata M, Nasuda S, Endo TR (2010) Dissection of barley chromosome 4H in common wheat by the gametocidal system and cytological mapping of chromosome 4H with EST markers. Genes Genet Syst 85:19–29
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH image to imageJ: 25 years of image analysis. Nat Methods 9:671–675
Serizawa N, Nasuda S, Shi F, Endo TR, Prodanovic S, Schubert I, Künzel G (2001) Deletion-based physical mapping of barley chromosome 7H. Theor Appl Genet 103:827–834
Singh D, Mohler V, Park RF (2013) Discovery, characterisation and mapping of wheat leaf rust resistance gene Lr71. Euphytica 190:131–136
Song LQ, Jiang LL, Han HM, Gao AN, Yang XM, Li LH, Liu WH (2013) Efficient induction of wheat–Agropyron cristatum 6P translocation lines and GISH detection. PLoS One 8:e69501
Tanaka H, Tsujimoto H (2012) Positive or negative effects on dough strength in large-scale group-1 chromosome deletion lines of common wheat (Triticum aestivum L.). Euphytica 186:57–65
Tsuchida M, Fukushima T, Nasuda S, Masoudi-Nejad A, Ishikawa G, Nakamura T, Endo TR (2008) Dissection of rye chromosome 1R in common wheat. Genes Genet Syst 83:43–53
Vales MI, Riera-Lizarazu O, Rines HW, Phillips RL (2004) Transmission of maize chromosome 9 rearrangements in oat–maize radiation hybrids. Genome 47:1202–1210
Vizir IY, Mulligan BJ (1999) Genetics of gamma-irradiation-induced mutations in Arabidopsis thaliana: large chromosomal deletions can be rescued through the fertilization of diploid eggs. J Hered 90:412–417
Wu J, Yang XM, Wang H, Li HJ, Li LH, Li XQ, Liu WH (2006) The introgression of chromosome 6P specifying for increased numbers of florets and kernels from Agropyron cristatum into wheat. Theor Appl Genet 114:13–20
Xie WL, Ben-David R, Zeng B, Dinoor A, Xie CJ, Sun QX, Röder MS, Fahoum A, Fahima T (2012a) Suppressed recombination rate in 6VS/6AL translocation region carrying the Pm21 locus introgressed from Haynaldia villosa into hexaploid wheat. Mol Breed 29:399–412
Xie WL, Ben-David R, Zeng B, Distelfeld A, Röder MS, Dinoor A, Fahima T (2012b) Identification and characterization of a novel powdery mildew resistance gene PmG3M derived from wild emmer wheat, Triticum dicoccoides. Theor Appl Genet 124:911–922
Xing LF, Wang CF, Xia XC, He ZH, Chen WQ, Liu TG, Li ZF, Liu DQ (2014) Molecular mapping of leaf rust resistance gene LrFun in Romanian wheat line Fundulea 900. Mol Breed 33:931–937
Xue SL, Xu F, Tang MZ, Zhou Y, Li GQ, An X, Lin F, Xu HB, Jia HY, Zhang LX, Kong ZX, Ma ZQ (2011) Precise mapping Fhb5, a major QTL conditioning resistance to Fusarium infection in bread wheat (Triticum aestivum L.). Theor Appl Genet 123:1055–1063
Xue SL, Xu F, Li GQ, Zhou Y, Lin MS, Gao ZX, Su XH, Xu XW, Jiang G, Zhang S, Jia HY, Kong ZX, Zhang LX, Ma ZQ (2013) Fine mapping TaFLW1, a major QTL controlling flag leaf width in bread wheat (Triticum aestivum L.). Theor Appl Genet 126:1941–1949
Ye XL, Lu YQ, Liu WH, Chen GY, Han HM, Zhang JP, Yang XM, Li XQ, Gao AN, Li LH (2015) The effects of chromosome 6P on fertile tiller number of wheat as revealed in wheat–Agropyron cristatum chromosome 5A/6P translocation lines. Theor Appl Genet 128:797–811
Zhang W, Zhang RQ, Feng YQ, Bie TD, Chen PD (2013) Distribution of highly repeated DNA sequences in Haynaldia villosa and its application in the identification of alien chromatin. Chin Sci Bull 58:890–897
Zhang JP, Liu WH, Han HM, Song LQ, Bai L, Gao ZH, Zhang Y, Yang XM, Li XQ, Gao AN, Li LH (2015a) De novo transcriptome sequencing of Agropyron cristatum to identify available gene resources for the enhancement of wheat. Genomics 106:129–136
Zhang J, Zhang JP, Liu WH, Han HM, Lu YQ, Yang XM, Li XQ, Li LH (2015b) Introgression of Agropyron cristatum 6P chromosome segment into common wheat for enhanced thousand-grain weight and spike length. Theor Appl Genet 128:1827–1837
Zhao PP, Meng QF, Guo N, Zhang LY, Yan HF, Liu DQ (2013) Analysis of virulence patterns of Puccinia triticina in Henan province in 2009–2011. J Henan Agric Sci 42:91–94
Zhou Y, Ren Y, Lillemo M, Yao ZJ, Zhang PP, Xia XC, He ZH, Li ZF, Liu DQ (2014) QTL mapping of adult-plant resistance to leaf rust in a RIL population derived from a cross of wheat cultivars Shanghai 3/Catbird and Naxos. Theor Appl Genet 127(9):1873–1883
Acknowledgments
This study was supported by the National High Technology Research and Development Program of China (the 863 program, No. 2011AA100102 and No. 2011AA100101), the National Basic Research Program of China (the 973 program, No. 2011CB100104), and the National Natural Science Foundation of China (No. 31271714).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Communicated by S. S. Xu.
L. Song and Y. Lu contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Song, L., Lu, Y., Zhang, J. et al. Physical mapping of Agropyron cristatum chromosome 6P using deletion lines in common wheat background. Theor Appl Genet 129, 1023–1034 (2016). https://doi.org/10.1007/s00122-016-2680-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-016-2680-8