Abstract
Key message
The recessive labile locus mapped on chromosome 5HL causes irregular spikelet fertility and controls floret development as well as row-type in barley.
Abstract
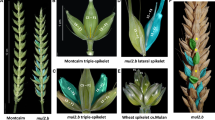
The labile-barley displays a variable number of fertile spikelets at each rachis internode (0–3 fertile spikelets/rachis internode) which is intermediate between that observed in two- or six-rowed types. Previous re-sequencing of Vrs1 in 219 labile-barley (Hordeum vulgare L. convar. labile) accessions showed that all carried a six-rowed specific allele. We therefore hypothesized that this seemingly random reduction in spikelet fertility is most likely caused by the labile (lab) locus, which we aimed to phenotypically and genetically define. Here, we report a detailed phenotypic analysis of spikelet fertility in labile-barleys in comparison to two- and six-rowed genotypes using scanning electron microscopy analysis. We found that the first visible morphological deviation occurred during the stamen primordium stage, when we regularly observed the appearance of arrested central floral primordia in labile but not in two- or six-rowed barleys. At late stamen and early awn primordium stages, lateral florets in two-rowed and only some in labile-barley showed retarded development and reduction in size compared with fully fertile lateral florets in six-rowed barley. We used two F2 mapping populations to generate whole genome genetic linkage maps and ultimately locate the lab locus as a recessive Mendelian trait to a 4.5–5.8 cM interval at approximately 80 cM on chromosome 5HL. Our results will help identifying the role of the lab gene in relation to other spikelet fertility factors in barley.



Similar content being viewed by others
Change history
21 July 2020
While continuing our quest towards the identification of the labile (lab) locus in barley, we discovered that the previously assigned map location on the long arm of chromosome 5H was wrong.
References
Abay F, Bjørnstad A (2009) Specific adaptation of barley varieties in different locations in Ethiopia. Euphytica 167:181–195
Åberg E, Wiebe GA (1945) Irregular barley, Hordeum irregulare, sp. nov. J Wash Acad Sci 35:161–164
Bjørnstad A, Abay F (2010) Multivariate patterns of diversity in Ethiopian barleys. Crop Sci 50:1579–1586
Bonnett OT (1935) A description of the development of the barley spike. J Agric Res 51:451–457
Breitenfeld C (1957) Genetische Untersuchungen zur Phylogenie und Systematik der intermedium- und labile-Gersten. Z Pflanzenzücht 38:275–312
Close TJ, Bhat PR, Lonardi S, Wu YH, Rostoks N, Ramsay L, Druka A, Stein N, Svensson JT, Wanamaker S, Bozdag S, Roose ML, Moscou MJ, Chao SAM, Varshney RK, Szucs P, Sato K, Hayes PM, Matthews DE, Kleinhofs A, Muehlbauer GJ, DeYoung J, Marshall DF, Madishetty K, Fenton RD, Condamine P, Graner A, Waugh R (2009) Development and implementation of high-throughput SNP genotyping in barley. BMC Genom 10:582. doi:10.1186/1471-2164-10-582
Comadran J, Kilian B, Russell J, Ramsay L, Stein N, Ganal M, Shaw P, Bayer M, Thomas W, Marshall D, Hedley P, Tondelli A, Pecchioni N, Francia E, Korzun V, Walther A, Waugh R (2012) Natural variation in a homolog of Antirrhinum CENTRORADIALIS contributed to spring growth habit and environmental adaptation in cultivated barley. Nat Genet 44:1388–1392
Djalali M (1970) Investigations on expressivity and penetrance of labile character of barley (Hordeum vulgare L.). Z Pflanzenzücht 63:274–322
Djalali M, Hoffman W, Plarre W (1970) Genetics and varaibility of the labile-gene in barley under different environmental conditions. Barley Genet II:201–207. WSUP, Washington
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Engledow FL (1924) Inheritance in barley III. J Genet 14:57–58, 60–62 illus
Forster BP, Franckowiak JD, Lundqvist U, Lyon J, Pitkethly I, Thomas WTB (2007) The barley phytomer. Ann Bot 100:725–733
Gawroński P, Schnurbusch T (2012) High-density mapping of the earliness per se-3A m (Eps-3A m) locus in diploid einkorn wheat and its relation to the syntenic regions in rice and Brachypodium distachyon L. Mol Breeding 30:1097–1108
Hadado TT, Rau D, Bitocchi E, Papa R (2009) Genetic diversity of barley (Hordeum vulgare L.) landraces from the central highlands of Ethiopia: comparison between the Belg and Meher growing seasons using morphological traits. Genet Resour Crop Evol 56:1131–1148
Harlan HV (1914) Some distinctions in our cultivated barleys with reference to their use in plant breeding, vol 137. Govt. print. off., U.S. Dept. Agr. Bull., illus, p 38, Washington. http://www.archive.org/details/somedistinctions00harl
Kirby E, Appleyard M (1987) Cereal development guide. NAC Cereal Unit, Stoneleigh
Komatsuda T, Pourkheirandish M, He CF, Azhaguvel P, Kanamori H, Perovic D, Stein N, Graner A, Wicker T, Tagiri A, Lundqvist U, Fujimura T, Matsuoka M, Matsumoto T, Yano M (2007) Six-rowed barley originated from a mutation in a homeodomain-leucine zipper I-class homeobox gene. Proc Natl Acad Sci USA 104:1424–1429
Koppolu R, Anwar N, Sakuma S, Tagiri A, Lundqvist U, Pourkheirandish M, Rutten T, Seiler C, Himmelbach A, Ariyadasa R, Youssef HM, Stein N, Sreenivasulu N, Komatsuda T, Schnurbusch T (2013) Six-rowed spike4 (Vrs4) controls spikelet determinacy and row-type in barley. Proc Natl Acad Sci USA 110:13198–13203
Li JZ, Sjakste TG, Röder MS, Ganal MW (2003) Development and genetic mapping of 127 new microsatellite markers in barley. Theor Appl Genet 107:1021–1027
Manikanda BN (2012) Genetic mapping and marker assisted selection: basics, practice and benefits. Springer, Berlin, pp 272–273
Mansfeld R (1950) Das morphologische System der Saatgerste, Hordeum vulgare L. s. l. Der Züchter 20:8–24
Mayer K, Martis M, Hedley P, Simková H, Liu H, Morris J, Steuernagel B, Taudien S, Roessner S, Gundlach H, Kubaláková M, Suchánková P, Murat F, Felder M, Nussbaumer T, Graner A, Salse J, Endo T, Sakai H, Tanaka T, Itoh T, Sato K, Platzer M, Matsumoto T, Scholz U, Dolezel J, Waugh R, Stein N (2011) Unlocking the barley genome by chromosomal and comparative genomics. Plant cell 23:1249–1263
Nötzel H (1952) Genetische Untersuchungen an röntgeninduzierten Gerstenmutanten. Kühn-Arch 66:72–132
Pourkheirandish M, Komatsuda T (2007) The importance of barley genetics and domestication in a global perspective. Ann Bot 100:999–1008
Ramsay L, Comadran J, Druka A, Marshall DF, Thomas WTB, Macaulay M, MacKenzie K, Simpson C, Fuller J, Bonar N, Hayes PM, Lundqvist U, Franckowiak JD, Close TJ, Muehlbauer GJ, Waugh R (2011) INTERMEDIUM-C, a modifier of lateral spikelet fertility in barley, is an ortholog of the maize domestication gene TEOSINTE BRANCHED 1. Nat Genet 43:169–172
Saisho D, Pourkheirandish M, Kanamori H, Matsumoto T, Komatsuda T (2009) Allelic variation of row type gene Vrs1 in barley and implication of the functional divergence. Breed Sci 59:621–628
Sreenivasulu N, Schnurbusch T (2012) A genetic playground for enhancing grain number in cereals. Trends Plant Sci 17:91–101
Stein N, Prasad M, Scholz U, Thiel T, Zhang H, Wolf M, Kota R, Varshney RK, Perovic D, Grosse I, Graner A (2007) A 1,000-loci transcript map of the barley genome: new anchoring points for integrative grass genomics. Theor Appl Genet 114:823–839
Takeda K, Saito W (1988) Inheritance of the percentage of missing lateral florets in ‘Irregular’ barley. Japan J Breed 38:72–80 (in Japanese with English summary p. 80)
van Ooijen JW (2006) JoinMap4, software for calculation of genetic linkage maps in experimental populations. Kyazma B.V, Wageningen
Vincze T, Posfai J, Roberts RJ (2003) NEBcutter: a program to cleave DNA with restriction enzymes. Nucleic Acids Res 31:3688–3691
von Bothmer R, Jacobsen N, Jorgensen RB (1985) Two new American species of Hordeum (Poaceae). Willdenowia 15:85–90
Youssef HM, Koppolu R, Schnurbusch T (2012) Re-sequencing of vrs1 and int-c loci shows that labile-barleys (Hordeum vulgare convar. labile) have a six-rowed genetic background. Genet Resour Crop Evol 59:1319–1328
Youssef HM, Fazeli A, Koppolu R, Rutten T, Ariyadasa R, Himmelbach A, Stein N, Schnurbusch T (2013) Does six rowed spike 2 (vrs2) play a role in regulating barley lateral spikelet fertility? VIPCA, Plant Genetics and Breeding Technologies, 18–20 February 2013, Vienna, Austria
Acknowledgments
The authors gratefully acknowledge Dr. Marion Röder for providing the SSR markers and lab facilities to test the F1 plants. We also thank Mrs. S. König and I. Walde for excellent technical support and H. Ernst for taking spike photos. We also would like to acknowledge funding from the DFG (Deutsche Forschungsgemeinschaft, FKZ SCHN 768/2-1) and the BMBF (German Federal Ministry of Education and Research, GABI-FUTURE Start Program grant number FKZ 0315071) during the course of this study.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by T. Komatsuda.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Youssef, H.M., Koppolu, R., Rutten, T. et al. Genetic mapping of the labile (lab) gene: a recessive locus causing irregular spikelet fertility in labile-barley (Hordeum vulgare convar. labile). Theor Appl Genet 127, 1123–1131 (2014). https://doi.org/10.1007/s00122-014-2284-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-014-2284-0