Abstract
Introduction
Multidrug-resistant organisms (MDRO) commonly colonize the gut microbiota of patients with Clostridioides difficile infection (CDI). This increases the likelihood of systemic infections with these MDROs. To help guide MDRO screening and/or empiric antibiotic therapy, we derived and compared predictive indices for MDRO gut colonization in patients with CDI.
Methods
This was a multicenter, retrospective cohort study of adult patients with CDI from July 2017 to April 2018. Stool samples were screened for MDRO via growth and speciation on selective antibiotic media and confirmed using resistance gene polymerase chain reaction. A regression-based risk score for MDRO colonization was constructed. Predictive performance via area under the receiver operating characteristic curve (aROC) of this index was compared with two other simplified risk stratification approaches: (1) prior healthcare exposure and/or high-CDI risk antibiotics; (2) number of prior high-CDI risk antibiotics.
Results
50 (20.8%) of 240 included patients had MDRO colonization; 35 (14.6%) VRE, 18 (7.5%) MRSA, 2 (0.8%) CRE. Prior fluoroquinolone (aOR 2.404, 95% CI 1.095–5.279) and prior vancomycin (1.996, 95% CI 1.014–3.932) were independently associated with MDRO colonization while prior clindamycin (aOR 3.257, 95% CI 0.842–12.597) and healthcare exposure (aOR 2.138, 95% CI 0.964–4.740) were retained as explanatory variables. The regression-based risk score significantly predicted MDRO colonization (aROC 0.679, 95% CI 0.595–0.763), but was not significantly more predictive than prior healthcare exposure + prior antibiotics (aROC 0.646, 95% CI 0.565–0.727) or number of prior antibiotic exposures (aROC 0.642, 95% CI 0.554–0.730); P > 0.05 for both comparisons.
Conclusion
A simplified approach using prior healthcare exposure and receipt of prior antibiotics known to increase CDI risk identified patients at risk for MDRO gut microbiome colonization as effectively as individual patient/antibiotic risk modeling.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Screening a patient’s gut microbiome for multidrug-resistant organism (MDRO) colonization is a potentially useful antimicrobial stewardship tool in populations at high risk for gut-borne MDRO infection, such as those with hematologic malignancy, critical illness, or Clostridioides difficile infection. |
Gut MDRO screening has been historically limited as it can be considered labor intensive and cost-prohibitive for routine implementation. |
Selective screening approaches based on baseline MDRO colonization risk can improve the value of screening results and limit unnecessary testing but few gut MDRO colonization risk stratification tools exist. |
This analysis developed and compared gut MDRO colonization risk stratification approaches in a population of patients with CDI who are at increased risk of MDRO colonization and infection. |
A simplified risk-stratification approach based on prior healthcare exposure and prior high-risk antibiotic use predicted gut MDRO colonization as well as a regression-based risk score and could be used clinically to identify patients at low risk of MDRO colonization in whom screening is unnecessary. |
Introduction
Antibiotic resistance due to multidrug-resistant organisms (MDRO) is a major public health threat [1]. Increased antibiotic resistance causes delayed appropriate antibiotic therapy, which increases mortality risk, prolongs hospital stay, and increases healthcare costs in patients with serious infections [2,3,4]. Despite the known benefits of early appropriate therapy, rates of delayed appropriate therapy are estimated to be in excess of 30% among patients with bacterial infections in most healthcare institutions [3]. This is likely a result of how common MDRO infections have become, limitations of current diagnostic methods, and how challenging prescribing empiric antibiotic therapy with limited information can be. Traditional microbiologic tests often require 48–72 h to identify antibiotic-resistant infections and the resulting delayed therapy increases mortality[3,4,5]. Even rapid diagnostic tests often cannot detect resistance during the first 12–24 h of infection and in patients with sepsis, each hour matters [6]. Thus, clinicians must rely on empiric antibiotic therapy using patient- and infection-specific factors to select therapy while balancing the need for early appropriate therapy with the desire to only use broad-spectrum therapy when necessary.
Screening for MDRO colonization is a potentially useful antimicrobial stewardship tool to guide initial empiric antibiotic therapy, as it can identify patients who are at risk for MDRO infection [7]. One screening approach that has generated much interest is the use of stool cultures or rectal swabs to identify MDRO gut colonization [8]. The gastrointestinal tract serves as a portal of entry for MDRO from the environment, an arena for antimicrobial resistance transfer and selection, and is a known contributor to subsequent infection [9, 10]. Progression from gut colonization to clinical infection can occur through organism translocation into the bloodstream or through environmental contamination and subsequent re-introduction into the body [10, 11]. This association between gut microbiome colonization is best established in patients with hematologic malignancies but has also been demonstrated in hospitalized adults, critically ill and postsurgical patients, pediatrics, and patients with Clostridioides difficile infection (CDI) [7, 12,13,14,15,16,17,18,19,20]. It has been demonstrated across many MDRO including vancomycin-resistant enterococci (VRE), gram negatives such as carbapenem-resistant Enterobacterales (CRE) and MDR Pseudomonas aeruginosa, Candida spp., and methicillin-resistant Staphylococcus aureus (MRSA) [14, 15, 18, 20,21,22]. Given the link between gut MDRO colonization and infection, it is not surprising that such screening has been shown to increase appropriate empiric antibiotic therapy [7].
Despite promise as a tool for both infection control and antimicrobial stewardship, gut MDRO screening has been historically limited as it can be considered labor intensive and cost-prohibitive for routine implementation. Identifying patients where screening can provide greatest benefit is a prudent approach to assessing all patients’ MDRO risk without performing costly tests in all [23]. Selective screening approaches using baseline MDRO risk can improve the value of screening results and limit unnecessary testing [24]. However, data regarding such a selective screening approach for gut MDRO colonization are limited. Patient risk stratification tools are needed to guide use of gut MDRO screening. The objective of this analysis was to develop and compare various MDRO colonization risk stratification approaches in a population of patients with CDI who are at increased risk of MDRO colonization and infection.
Methods
Study Design and Population
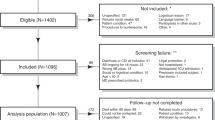
This was a multicenter, retrospective, observational cohort study of adult patients (age ≥ 18 years) diagnosed with CDI from July 2017 to April 2018 at four hospitals within CHI St. Luke’s Health in Greater Houston. Clostridioides difficile (C. difficile) infection diagnosis was based on positive C. difficile nucleic acid amplification test or enzyme-linked immunosorbent assay in patients with unexplained and new-onset diarrhea defined as three or more unformed stools in 24 h. Patients whose initial C. difficile-positive stool sample could not be collected from the hospital microbiologic labs for additional testing were excluded. This study was approved by the University of Houston (UH) Committee for the Protection of Human Subjects with waiver of informed consent granted (Institutional Review Board study 00000128) and was conducted in accordance with the principles of the Declaration of Helsinki of 1964, and its later amendments.
Patient Data Elements and Collection
Eligible patients were identified for inclusion by screening a list of patients with a positive C. difficile nucleic acid amplification test or enzyme-linked immunosorbent assay. Patient data were extracted from the medical record by trained reviewers using a structured data collection form within REDCap (Research Electronic Data Capture, Vanderbilt University) data capture tool hosted at UH [25]. Data elements included demographics, past medical history including previous healthcare exposure, comorbid conditions, medication administrations including antibiotics in the 90 days preceding CDI diagnosis, pertinent laboratory data, and clinical outcomes. The degree of patient comorbidity was quantified using the Charlson Comorbidity Index [26]. High-risk antibiotic exposures were defined on the basis of proclivity to disrupt gut microbiome and cause CDI infection and included carbapenems, 2nd–4th-generation cephalosporins, fluoroquinolones, clindamycin, ampicillin/sulbactam, and piperacillin/tazobactam. High-risk healthcare exposures were defined as prior hospitalization within the 90 days preceding CDI diagnosis or admission from a nursing home. Severity of CDI was classified in accordance with Infectious Diseases Society of America CDI guidelines [27].
Stool Sample Collection and MDRO Screening
Leftover stool from the initial CDI-positive stool samples was collected and brought to a centralized research laboratory at the University of Houston. Stool sample aliquots were then plated on antibiotic-selective media to isolate potential VRE, CRE, or MRSA (HardyCHROM, Hardy Diagnostics, Santa Maria CA) [28, 29]. Resulting colony growth was then counted and speciated using the Biolog Omnilog (http://www.biolog.com). Presence of antimicrobial resistance in resulting colonies was confirmed via polymerase chain reaction (PCR) using relevant gene targets including vanA for VRE, blaKPC for CRE, and mecA for MRSA. Patients were classified as having gut microbiome MDRO colonization if their stool sample contained any of these MDRO.
Data Analysis
The primary analysis focused on comparing the predictive performance of three approaches to stratify patients by MDRO colonization risk: (1) logistic regression-derived risk score; (2) the number of high-risk antibiotic exposures in the preceding 90 days; (3) high-risk healthcare exposure and/or any high-risk antibiotic exposure in the preceding 90 days. Predictive performance of each approach was quantified by area under the receiver operating characteristics curve (aROC) along with a 95% confidence interval. The aROC of each risk assessment approach was compared using the Hanley and McNeil Method [30]. High-risk healthcare exposure and/or any high-risk antibiotic exposure in the preceding 90 days was considered the reference category because it is the simplest and most practical risk stratification approach. The prevalence of MDRO colonization across the distribution of categories of each risk stratification approach was also examined using the χ2 test.
To construct the logistic regression-derived risk score, bivariate analysis first identified patient characteristics associated with MDRO colonization. Categorical variables were compared between those with and without MDRO colonization using the χ2 or Fisher’s exact test and continuous variables were compared using the Student’s t test or Mann–Whitney U test. Characteristics associated with MDRO colonization at a P value < 0.2 in bivariate analysis were simultaneously included as covariate in a multivariable logistic regression model and then removed in a backward, stepwise fashion. Covariates were retained in the final model if the P value for the likelihood ratio test for their removal was < 0.1. The number of covariates in each candidate regression model was limited to one covariate per 10 MDRO patients (i.e., 50 MDRO patients = 5 covariates per candidate model). Model fit was assessed with the Hosmer–Lemeshow goodness-of-fit test; models with a non-significant result were considered adequate. Multicollinearity of candidate regression models was assessed via the variance inflation factor, with values between 1 and 5 considered acceptable. The risk score was then constructed by integer rounding the final regression model β values for each risk factor (i.e., β value of 0.75 was rounded to 1).
All statistical tests were two-sided; P values ≤ 0.05 were considered statistically significant. Analyses were performed using SPSS Statistics, IBM SPSS software, version 26.0 (IBM Corp., Armonk, NY).
Results
A total of 240 patients were included. The majority were female (53.8%) and the median (IQR) age was 64.5 (52–75) years. Common comorbidities were diabetes (38.8%), gastroesophageal reflux disease (18.3%), cerebrovascular disease (17.5%), heart failure (17.1%), chronic kidney disease (17.1%), and liver disease (12.1%). The median (IQR) Charlson Comorbidity Index was 2 (1–3). The majority had a recent high-risk healthcare exposure (62.1%) including 10.8% who were admitted from a nursing home and 56.7% who were hospitalized in the 90 days preceding CDI diagnosis. The majority of patients also received antibiotic therapy in the 90 days preceding CDI diagnosis (71.7%) including 60% who received a high-CDI risk antibiotic. Fifty patients (20.8%) had a MDRO-positive stool sample including 35 (14.6%) VRE, 18 (7.5%) MRSA, and 2 (0.8%) CRE. The stool samples of 5 patients (2.1%) were positive for both VRE and MRSA.
A bivariate comparison of patient characteristics between patients with and without MDRO colonization is displayed in Table 1. Patients with MDRO colonization had a higher degree of comorbidity, were more likely to have had prior high-risk healthcare exposure, and were more likely to have received antibiotics in the past 90 days. Results of the logistic regression model are shown in Table 2. Prior fluoroquinolone (aOR 2.404, 95% CI 1.095–5.279) and prior vancomycin (1.996, 95% CI 1.014–3.932) were independently associated with MDRO colonization while prior clindamycin (aOR 3.257, 95% CI 0.842–12.597) and healthcare exposure (aOR 2.138, 95% CI 0.964–4.740) were retained as explanatory variables. Each risk factor was weighted equally in the risk score, with one point assigned for each.
The predictive performances of the regression-derived risk score, number of high-risk antibiotic exposures in the preceding 90 days, and high-risk healthcare exposure and/or any high-risk antibiotic exposure in the preceding 90 days are shown in Table 3. All three risk stratification approaches predicted MDRO colonization significantly better than random chance. Neither the regression-derived risk score (aROC 0.679, 95% CI 0.595–0.763) nor the number of high-CDI risk antibiotics in the past 90 days (aROC 0.642, 95% CI 0.554–0.730) was significantly more predictive than high-risk healthcare exposure and/or any high-risk antibiotic exposure in the preceding 90 days (aROC 0.646, 95% CI 0.565–0.727); P > 0.05 for both comparisons.
The prevalence of MDRO colonization across the distribution of categories of each risk stratification approach is displayed in Fig. 1. As the regression-derived risk score (Fig. 1a) and number of high-CDI risk antibiotic exposures increased (Fig. 1b), so did the MDRO colonization prevalence. Having a high-risk healthcare exposure or a prior high-CDI risk antibiotic exposure increased MDRO prevalence but the prevalence was highest among patients with both a high-risk healthcare exposure and a prior high-CDI risk antibiotic exposure (Fig. 1c). The regression-derived risk score was able to identify those with the highest MDRO colonization prevalence, with 68.6% of patients with a score of 3 or 4 having MDRO colonization. In contrast, the simple approach using high-risk healthcare exposure and/or any high-risk antibiotic exposure in the preceding 90 days identified patients with the lowest MDRO risk. Among patients with neither a high-risk healthcare exposure nor a prior high-CDI risk antibiotic exposure, MDRO prevalence was 7.4%.
Discussion
This study sought to compare three potential approaches to risk-stratify patients with CDI on the basis of their risk of having MDRO gut microbiome colonization. An individual patient risk score was constructed via logistic regression which was able to significantly predict MDRO colonization compared to random chance. However, this risk score incorporating specific high-risk antibiotics and high-risk healthcare exposure had similar performance characteristics compared to more two simplified approaches: (1) the number of high-risk antibiotics in the past 90 days or (2) whether a patient had a high-risk healthcare exposure and/or high-risk antibiotic exposure. Determining whether a patient had either a high-risk health exposure (nursing home or recent hospitalization), was exposed to a high-CDI risk antibiotic in the preceding 90 days (antibiotics linked to CDI), or both was the simplest and most practical approach we evaluated. Despite the relative simplicity compared to the more complicated risk scoring approaches, it was able to identify both high- and low-MDRO risk patients just as well.
The ability of the simplified risk stratification approach based on healthcare exposure and prior high-risk antibiotic exposure to predict MDRO colonization is not surprising. These two factors are the most consistently reported risk factors for MDRO colonization in patients with and without CDI infection [31,32,33,34]. This makes biologic sense as exposure to antibiotics known to disrupt the gut microbiome can create an environment ripe for colonization by more resistant pathogens and exposure to high-risk healthcare settings brings patients into contact with these resistant organisms [9, 10].
Although the performance characteristics of these risk stratification approaches only demonstrated fair discrimination, they may still provide some value to guide diagnostic tests. Because patients without a high-risk healthcare exposure who did not have a recent high-risk antibiotic exposure appeared to be at low risk of MDRO colonization (< 10%), it may be reasonable to forgo MDRO screening in those patients. Empiric antibiotic therapy can still be guided on the basis of patient clinical characteristics and infection type, but the low MDRO colonization prevalence in this group suggests MDRO screening may not be necessary, as it is likely to be a negative result. Empiric antibiotic is currently routinely selected without knowledge of gut microbiome MDRO colonization so this is not a departure from standard of care. In contrast, patients with prior healthcare exposure, prior antibiotic exposure, or both are at seemingly higher risk of MDRO colonization. In these patients, MDRO screening has two potential benefits. First, a negative MDRO screening test may allow clinicians to prescribe more narrow-spectrum antibiotic therapy in low acuity, low mortality risk infections when they may ordinarily be compelled to provide broad spectrum therapy based on the patient’s history of healthcare exposure and/or antibiotics. Second, a positive test for VRE or CRE may allow clinicians to cover these MDRO which are not typically covered with empiric antibiotic regimens. Antibiotic coverage can then be de-escalated if these MDRO are not the ultimate cause of the patient’s suspected infection while the early, appropriate therapy can reduce mortality risk in the patients who are infected with these colonizing MDRO [35, 36].
There are a number of considerations when interpreting theses analyses. The results were derived from a population of patients with CDI from a single health system and as such, it is unclear if they are generalizable to other populations of interest. The fact that all patients were diagnosed with CDI indicates they likely all had significant disruptions to their gut microbiome which likely differs from more general patient populations. Despite this, published literature strongly suggests that prior healthcare exposure and prior antibiotic exposure drive MDRO colonization, so it is reasonable that these risk stratifications would also predict MDRO colonization in other patient groups. In addition, this analysis focused on three specific MDRO: VRE, MRSA, and CRE. The results are most applicable to VRE considering the majority of MDRO-positive patients had VRE and only two patients had CRE. It is unclear whether the results would be similar had other common enteric MDRO been examined such as extended spectrum beta-lactamase (ESBL)-producing Enterobacterales. Because we did not test for this MDRO, it is possible that some of the patients classified as MDRO negative actually did have MDRO colonization. Because ESBL colonization has also been linked to healthcare exposure and prior antibiotic exposure, it is likely that including ESBL would have strengthened the ability to accurately risk-stratify patients and it is possible that the more complex regression-based approach would have performed better.
In addition, the exact CDI diagnostic method was not recorded for each patient. It is possible that some of the included patients did not have a stool toxin test during CDI diagnosis, as recommended in the most current Infectious Diseases Society of America (IDSA) and Society for Healthcare Epidemiology of America (SHEA) guidelines [27]. As such, it is possible that some of the included patients did not have true CDI. However, because all patients were required to have unexplained and new-onset diarrhea defined as three or more unformed stools in 24 h to be included, the number of patients in this study without true infection is likely very small. Because this study focused on the association between antibiotic and healthcare exposure and MDRO colonization, potential inclusion of a small number patients who were only colonized with C. difficile rather than infected should not have a major impact on findings. Finally, this was a convenience sample of modest size and no formal sample size calculations were done. Although no statistically significant difference in predictive performance between risk stratification approaches was noted in the analysis, the regression-based risk score aROC was numerically greater than the other approaches. It is possible that a larger sample size may have revealed a statistically significant effect.
Conclusions
A simplified approach using prior healthcare exposure and receipt of prior high-risk antibiotics identified patients with CDI at risk for MDRO gut microbiome colonization as effectively as individual patient/antibiotic risk modeling. Further validation of these results and an evaluation of their potential clinical application on MDRO screening and empiric antibiotic prescribing is required. Notably, evaluating such risk stratification approaches in patients without CDI will be important before clinical application. However, this analysis serves as a proof of concept in a population of patients with high MDRO colonization risk due to their clear microbiome disruption as evidenced by concurrent CDI.
References
Centers for Disease Control and Prevention. Antibiotic resistance threats in the United States, 2019. https://www.cdc.gov/drugresistance/pdf/threats-report/2019-ar-threats-report-508.pdf. Accessed Nov 14.
Zasowski EJ, Bassetti M, Blasi F, et al. A systematic review of the effect of delayed appropriate antibiotic treatment on the outcomes of patients with severe bacterial infections. Chest. 2020;158(3):929–38.
Bonine NG, Berger A, Altincatal A, et al. Impact of delayed appropriate antibiotic therapy on patient outcomes by antibiotic resistance status from serious Gram-negative bacterial infections. Am J Med Sci. 2019;357(2):103–10.
Lodise TP, Zhao Q, Fahrbach K, Gillard PJ, Martin A. A systematic review of the association between delayed appropriate therapy and mortality among patients hospitalized with infections due to Klebsiella pneumoniae or Escherichia coli: how long is too long? BMC Infect Dis. 2018;18(1):625.
Bauer KA, Perez KK, Forrest GN, Goff DA. Review of rapid diagnostic tests used by antimicrobial stewardship programs. Clin Infect Dis. 2014;59(Suppl 3):S134–45.
Peltan ID, Brown SM, Bledsoe JR, et al. ED door-to-antibiotic time and long-term mortality in sepsis. Chest. 2019;155(5):938–46.
Blot S, Depuydt P, Vogelaers D, et al. Colonization status and appropriate antibiotic therapy for nosocomial bacteremia caused by antibiotic-resistant gram-negative bacteria in an intensive care unit. Infect Control Hospital Epidemiol. 2005;26(6):575–9.
Viau R, Frank KM, Jacobs MR, et al. Intestinal carriage of carbapenemase-producing organisms: current status of surveillance methods. Clin Microbiol Rev. 2016;29(1):1–27.
Carlet J. The gut is the epicentre of antibiotic resistance. Antimicrob Resist Infect Control. 2012;1(1):39.
Donskey CJ. The role of the intestinal tract as a reservoir and source for transmission of nosocomial pathogens. Clin Infect Dis. 2004;39(2):219–26.
Climo MW, Sepkowitz KA, Zuccotti G, et al. The effect of daily bathing with chlorhexidine on the acquisition of methicillin-resistant Staphylococcus aureus, vancomycin-resistant Enterococcus, and healthcare-associated bloodstream infections: results of a quasi-experimental multicenter trial. Crit Care Med. 2009;37(6):1858–65.
Thomas JA, Newman KC, Doshi S, Logan N, Musher DM. Bacteraemia from an unrecognized source (occult bacteraemia) occurring during Clostridium difficile infection. Scand J Infect Dis. 2011;43(4):269–74.
Pena C, Pujol M, Ardanuy C, et al. Epidemiology and successful control of a large outbreak due to Klebsiella pneumoniae producing extended-spectrum beta-lactamases. Antimicrob Agents Chemother. 1998;42(1):53–8.
Swaminathan M, Sharma S, Poliansky Blash S, et al. Prevalence and risk factors for acquisition of carbapenem-resistant Enterobacteriaceae in the setting of endemicity. Infect Control Hospital Epidemiol. 2013;34(8):809–17.
Faden H, Lesse AJ, Trask J, et al. Importance of colonization site in the current epidemic of staphylococcal skin abscesses. Pediatrics. 2010;125(3):e618–24.
Stiefel U, Paterson DL, Pultz NJ, Gordon SM, Aron DC, Donskey CJ. Effect of the increasing use of piperacillin/tazobactam on the incidence of vancomycin-resistant enterococci in four academic medical centers. Infect Control Hospital Epidemiol. 2004;25(5):380–3.
Webb BJ, Healy R, Majers J, et al. Prediction of bloodstream infection due to vancomycin-resistant enterococcus in patients undergoing leukemia induction or hematopoietic stem-cell transplantation. Clin Infect Dis. 2017;64(12):1753–9.
Taur Y, Xavier JB, Lipuma L, et al. Intestinal domination and the risk of bacteremia in patients undergoing allogeneic hematopoietic stem cell transplantation. Clin Infect Dis. 2012;55(7):905–14.
Kamboj M, Blair R, Bell N, Sun J, Eagan J, Sepkowitz K. What is the source of bloodstream infection due to vancomycin-resistant enterococci in persons with mucosal barrier injury? Infect Control Hospital Epidemiol. 2014;35(1):99–101.
Tancrede CH, Andremont AO. Bacterial translocation and gram-negative bacteremia in patients with hematological malignancies. J Infect Dis. 1985;152(1):99–103.
Nucci M, Anaissie E. Revisiting the source of candidemia: skin or gut? Clin Infect Dis. 2001;33(12):1959–67.
Squier C, Rihs JD, Risa KJ, et al. Staphylococcus aureus rectal carriage and its association with infections in patients in a surgical intensive care unit and a liver transplant unit. Infect Control Hospital Epidemiol. 2002;23(9):495–501.
Diekema DJ, Dodgson KJ, Sigurdardottir B, Pfaller MA. Rapid detection of antimicrobial-resistant organism carriage: an unmet clinical need. J Clin Microbiol. 2004;42(7):2879–83.
Zilberberg MD, Shorr AF. Impact of prior probabilities of MRSA as an infectious agent on the accuracy of the emerging molecular diagnostic tests: a model simulation. BMJ Open. 2012;2(6): e001804.
Harris PA, Taylor R, Thielke R, Payne J, Gonzalez N, Conde JG. Research electronic data capture (REDCap)–a metadata-driven methodology and workflow process for providing translational research informatics support. J Biomed Inform. 2009;42(2):377–81.
Charlson ME, Pompei P, Ales KL, MacKenzie CR. A new method of classifying prognostic comorbidity in longitudinal studies: development and validation. J Chronic Dis. 1987;40(5):373–83.
McDonald LC, Gerding DN, Johnson S, et al. Clinical Practice Guidelines for Clostridium difficile infection in adults and children: 2017 update by the Infectious Diseases Society of America (IDSA) and Society for Healthcare Epidemiology of America (SHEA). Clin Infect Dis. 2018;66(7):987–94.
Gonzales-Luna AJ, Spinler JK, Oezguen N, et al. Systems biology evaluation of refractory Clostridioides difficile infection including multiple failures of fecal microbiota transplantation. Anaerobe. 2021;70: 102387.
Halton K, Arora V, Singh V, Ghantoji SS, Shah DN, Garey KW. Bacterial colonization on writing pens touched by healthcare professionals and hospitalized patients with and without cleaning the pen with alcohol-based hand sanitizing agent. Clin Microbiol Infect. 2011;17(6):868–9.
Hanley JA, McNeil BJ. A method of comparing the areas under receiver operating characteristic curves derived from the same cases. Radiology. 1983;148(3):839–43.
Bhorade SM, Christenson J, Pohlman AS, Arnow PM, Hall JB. The incidence of and clinical variables associated with vancomycin-resistant enterococcal colonization in mechanically ventilated patients. Chest. 1999;115(4):1085–91.
Nijssen S, Fluit A, van de Vijver D, Top J, Willems R, Bonten MJ. Effects of reducing beta-lactam antibiotic pressure on intestinal colonization of antibiotic-resistant gram-negative bacteria. Intensive Care Med. 2010;36(3):512–9.
Schoevaerdts D, Verroken A, Huang TD, et al. Multidrug-resistant bacteria colonization amongst patients newly admitted to a geriatric unit: a prospective cohort study. J Infect. 2012;65(2):109–18.
Salomão MC, Guimarães T, Duailibi DF, et al. Carbapenem-resistant Enterobacteriaceae in patients admitted to the emergency department: prevalence, risk factors, and acquisition rate. J Hosp Infect. 2017;97(3):241–6.
Zasowski EJ, Claeys KC, Lagnf AM, Davis SL, Rybak MJ. Time is of the essence: the impact of delayed antibiotic therapy on patient outcomes in hospital-onset enterococcal bloodstream infections. Clin Infect Dis. 2016;62:1242–50.
Lodise TP, Berger A, Altincatal A, et al. Antimicrobial resistance or delayed appropriate therapy-does one influence outcomes more than the other among patients with serious infections due to carbapenem-resistant versus carbapenem-susceptible enterobacteriaceae? Open Forum Infect Dis. 2019;6(6):ofz194.
Acknowledgements
We thank the participants of this study.
Funding
This work was funded by the 2017–2018 Society of Infectious Diseases Pharmacists/BioMerieux Microbial Diagnostics in Antimicrobial Stewardship Research Award. No funding was received for the publication of this article.
Authorship
All named authors meet the International Committee of Medical Journal Editors (ICMJE) criteria for authorship for this article, take responsibility for the integrity of the work as a whole, and have given their approval for this version to be published.
Author Contributions
Evan J. Zasowski, Bradley T. Endres, and Kevin W. Garey conceived the study. All authors contributed to the study design, were involved in interpretation of the data, revised the manuscript, and approved the final version before submitting. Evan J. Zasowski, Bradley T. Endres, and Kevin W. Garey obtained funding. Evan J. Zasowski, Maryam Ali, Ada Anugo, Nayle Ibragimova, Kierra M. Dotson, Khurshida Begum, and M Jahangir Alam collected the data. Evan J. Zasowski performed statistical analysis. Evan J. Zasowski wrote the first draft of the report, and designed the tables and figures. The corresponding author confirms that he had full access to all the data in the study and had final responsibility for the decision to submit for publication.
Prior Presentation
Parts of these data have been previously presented at the European Congress of Clinical Microbiology virtual congress in 2020.
Disclosures
Since completion of this study, Evan J. Zasowski has changed affiliations and is currently and employee of Pfizer Inc. Kevin W Garey, Maryam Ali, Ada Anugo, Nayle Ibragimova, Kierra M. Dotson, Bradley T. Endres, Khurshida Begum, M Jahangir Alam have nothing to disclose.
Compliance with Ethics Guidelines
This study was approved by the University of Houston (UH) Committee for the Protection of Human Subjects with waiver of informed consent granted (Institutional Review Board study 00000128) and was conducted in accordance with the principles of the Declaration of Helsinki of 1964, and its later amendments.
Data Availability
The datasets from this study are available from the corresponding author on reasonable request.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Zasowski, E.J., Ali, M., Anugo, A. et al. Comparison of Risk Stratification Approaches to Identify Patients with Clostridioides difficile Infection at Risk for Multidrug-Resistant Organism Gut Microbiota Colonization. Infect Dis Ther 12, 2005–2015 (2023). https://doi.org/10.1007/s40121-023-00843-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40121-023-00843-9