Abstract
Purpose of Review
Chronic liver disease is a major cause of morbidity and mortality amongst people living with HIV (PLWH). Emerging data suggests that gut microbial translocation may play a role in driving and modulating liver disease, a bi-directional relationship termed the gut–liver axis. While it is recognized that PLWH have a high degree of dysbiosis and gut microbial translocation, little is known about the gut–liver axis in PLWH.
Recent Findings
Recent studies have shown that microbial translocation can directly lead to hepatic inflammation, and have linked gut microbial signatures, dysbiosis, and translocation to liver disease in PLWH. Additionally, multiple trials have explored interventions targeting the microbiome in PLWH.
Summary
Emerging research supports the interaction between the gut microbiome and liver disease in PLWH. This offers new opportunities to expand our understanding of the pathophysiology of liver disease in this population, as well as to explore possible clinical interventions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The human microbiome is comprised of trillions of bacteria, viruses, fungi, protozoa, and archaea that live on skin, gut lumen, and other mucosal surfaces. The study of the composition, byproducts, and effects of the human microbiome has flourished in the last decade, and many possible relationships have emerged between it and the pathogenesis of various diseases [1]. The unique association between the liver and the gut, connected via the portal system, systemic circulation and the biliary system, termed the gut–liver axis, has led to significant interest in how the microbiome and its byproducts may impact liver disease.
Chronic liver disease is a leading cause of morbidity and mortality among people living with HIV (PLWH) [2–5]. Viral hepatitis has historically been the cause of most liver disease amongst PLWH. The development of highly effective hepatitis B virus (HBV) antiviral therapy and curative hepatitis C virus (HCV) therapy has shifted this landscape, though data suggests that despite HCV cure, underlying liver disease may persist in PLWH [6]. HIV infection has also been linked to non-alcoholic fatty liver disease (NAFLD), which is common and more severe amongst PLWH than those without HIV [7, 8]. The drivers of this disparity remain unknown, but liver disease in PLWH has been increasingly linked to the gut microbiome. Dysbiosis, or perturbations in the composition and function of the microbiome, is especially striking in the setting of HIV infection [9, 10]. This review outlines the existing literature on the gut–liver axis in PLWH, starting with the impact of HIV on the gut microbiome and integrity, the effects of dysbiosis on the liver, and finally potential therapeutic interventions targeting the microbiome in PLWH.
HIV and the Gut
HIV and Immune Dysregulation
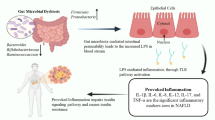
The human intestinal mucosa is comprised of a layer of columnar epithelial cells, lamina propria, and muscularis mucosa [11]. The epithelial layer plays an important role in maintaining a healthy barrier between the luminal contents and the circulatory system and is maintained by CD4 + TH17 + T cells in the lamina propria which produce IL-17 and IL-22 [11, 12]. These cytokines induce epithelial cell proliferation, epithelial tight junction formation, and the expression of claudins, mucin, and defensins (Fig. 1) [11].
Upon infection with HIV, there is a rapid loss of activated T cells in the gut. This is particularly pronounced in the lamina propria and is thought to be due to direct cytotoxic effects of the virus, as gut HIV RNA levels correlate with the amount of CD4 + cell loss. Mucosal TH17 + , TH22 + cells as well as other activated cytokine producing T cells are preferentially impacted and may show reduced functional capabilities [13–15]. The decrease in TH17 + cells leads to a reduction in IL-17 and IL-22, which in turn disrupts the epithelial cell barrier due to abnormalities in the IL-17-regulated tight junction protein occludin [16]. These tight junction breaches, as well as other consequences of decreased IL-17 and IL-22, increase the potential for bacterial translocation from the gut lumen into the portal and systemic circulation (Fig. 1) [17]. Murine models have corroborated this, showing that IL-17 loss leads to shifts in the microbiome composition with associated greater systemic inflammation and a weakened gut luminal barrier [18]. Studies evaluating macaque models infected with simian immunodeficiency virus (SIV) have directly visualized microbial translocation from the gut, and found that in the absence of gut translocation there is no systemic immune activation [17, 19].
Notably, these mucosal changes persist despite HIV treatment, that is, CD4 + cell loss in the gut was observed on histologic evaluation 6 months after initiating anti-retroviral therapy (ART) [14]. In chronic HIV infection, regulatory T cells (Treg) are upregulated and can help dampen immune activation; however, they have also been found to lead to collagen deposition in the gut mucosa, further depleting the CD4 + population in the gut [20]. Multiple studies have demonstrated disruption of the epithelial cell barrier in both acute and chronic HIV and SIV infection [19, 21, 22].
HIV and Microbial Translocation
In a landmark study, Brenchley et al. provided a link between HIV infection, microbial translocation, and systemic immune activation [23]. In patients with chronic untreated HIV infection and in rhesus macaques with acute SIV infection, lipopolysaccharide (LPS), a marker of bacterial translocation, was elevated and strongly associated with levels of innate immune activation. ART reduced levels of LPS somewhat but did not normalize it [23]. Another study in macaque models showed that lack of histologic evidence for microbial translocation was associated with an absence of chronic immune activation [19]. Subsequent studies demonstrated that higher levels of soluble (s)CD14 (sCD14, a marker of monocyte response to LPS), gut epithelial barrier dysfunction, and innate immune activation predict mortality in patients with treated HIV infection [13, 24]. Taken together, these studies link the gut microbiome to chronic immune activation in PLWH gut barrier dysfunction and microbial translocation.
In addition to its effects on gut permeability, HIV infection may impact the composition of the microbiome. Vujkovic-Cvijin et al. demonstrated that the gut microbiomes from people living with untreated HIV were more likely to harbor pathogenic bacteria and less commensal bacteria compared to HIV-seronegative individuals [25]. Interestingly, the gut microbiota compositions of PLWH on ART varied widely, with some exhibiting a composition similar to those with untreated HIV and others closer to those without HIV; moreover, in PLWH on ART, the level of systemic inflammation was correlated to microbial composition patterning that resembled untreated HIV [25, 26]. Potentially harmful classes such as proteobacteria were enriched in HIV and commensal bacteria such as Bacteroides were depleted [25].
Finally, the bidirectionality between microbiome translocation and the gut luminal barrier in HIV was explored by Dillon et al. Their group demonstrated that HIV increases the amount of pathobionts in the gut microbiome, which can translocate across the weakened tight junctions in the epithelial barrier and activate innate immunity [10]. The activation of this innate immunity can in turn decrease TH17 + cells and lead to more epithelial barrier breakdown and thus microbial translocation [10]. More recent work has corroborated this and offered some optimism. A 2021 study analyzed markers of microbial translocation and immune activation in the peripheral serum of patients with treated HIV infection and suggested that ART leads to improvements in markers of microbial translocation and gut integrity after 2 years [27].
The Human Microbiome and Liver Disease
The Gut–Liver Axis
A strong physiologic link exists between the gut lumen and the liver, as all venous outflow from the intestinal tract flows through the portal system and into hepatic sinusoids. This connection is bidirectional, with bile acids produced by the liver flowing into the duodenal lumen via the common bile duct and the Sphincter of Oddi. In addition, the liver and the gut are connected to each other (and the rest of the body) via systemic circulation. As a result of this interconnection, the liver is perhaps the organ most directly exposed to the contents of the intestinal lumen, including microbial flora and its byproducts.
Indeed, microbial translocation directly into the liver has been studied in SIV models. Estes et al. visualized increased amounts of E. coli in hepatic tissue in macaques with SIV/AIDS and epithelial barrier dysfunction [19]. Over the years, the translocation of microbes and microbial byproducts across the intestinal epithelial barrier has been associated with the pro-inflammatory state driving liver diseases of diverse etiologies [28–30]. When microbial products translocate across the gut barrier, they enter the portal venous system and enter the hepatic sinusoids containing both Kupffer and hepatic stellate cells. Microbial associated molecular products (MAMPS) activate toll-like receptors on both Kupffer and stellate cells, leading to an inflammatory cascade mediated by Kupffer cells while stellate cell activation contributes to further injury and fibrosis (Fig. 1) [29].
As one example, the microbiome may play a significant role in the pathogenesis of NAFLD. In a groundbreaking experiment in 2013, Le Roy et al. demonstrated that the tendency to develop NAFLD in mice could be transmitted by transplanting gut microbiota [31]. While not specific to NAFLD, results of humans studies evaluating the effects of fecal microbiota transplant (FMT) suggest that the microbiome may mediate metabolic syndrome, since participants with metabolic syndrome who underwent intestinal infusion of gut microbiota from lean individuals had improved insulin [32]. While the exact mechanism by which the microbiome induces NAFLD is incompletely elucidated, steatohepatitis may be linked to certain microbial signatures, particularly a predominance of Bacteroides and Ruminococcus species [33]. Immunologic responses to these microbiota via toll-like receptors may play a role, as TLR4 deficiency in murine models has been associated with an attenuation in steatohepatitis [34]. Other microbiome factors implicated in the pathogenesis of NAFLD include microbiota-driven changes in caloric absorption, altered choline metabolism mimicking choline deficiency, and an increase in short chain fatty acids [35]. NAFLD is just one of many chronic liver conditions being increasingly linked to dysbiosis [36]. Moreover, different liver disease etiologies are associated with unique patterns of dysbiosis, with alcohol-associated liver disease having a higher predominance of Enterobacteriaceae as well as higher gut permeability when compared to other causes of cirrhosis [28, 37].
Cirrhosis itself also causes a profound dysbiosis, and the gut microbiome in patients with cirrhosis is enriched in Enterobacteriaceae, Enterococcaceae, and Staphylococceae as opposed to the predominantly Bacteroides and Firmicutes present in healthy individuals [28, 36, 38–41]. Emerging data suggest that microbiome composition may influence infectious outcomes in decompensated cirrhosis [42] that microbial products may be linked to the development of hepatocellular carcinoma, and that the cirrhotic liver may be less able to clear microbial byproducts [43]. Impaired clearance of microbial byproducts could then lead to greater immune activation, resulting in a bidirectional effect similar to microbial translocation in HIV. Our developing understanding of the relationship between the microbiome and liver disease can help guide diagnostics and surveillance of liver disease, with one study demonstrating that metagenomic microbiome signatures can predict non-alcoholic steatohepatitis (NASH) cirrhosis with an AUC of 0.91 [44], and another showing that microbiome analysis can identify early hepatocellular carcinoma [45].
HIV, HCV, and the Gut Microbiome
There is a rich new field investigating the connection between the gut–liver axis an HIV infection. Animal studies evaluating macaques infected with SIV demonstrated a 20-fold increase of bacterial products in the livers of SIV-infected animals resulting in CXCL16 production by myeloid dendritic cells (mDCs) [46]. Hepatic mDC activation and recruitment of NK cells expressing CXCL16 receptor correlated significantly with liver damage and fibrosis [74]. In a different SIV model, dysbiosis in SIV infection persisted despite treatment with ART and was characterized by an increase in atypical mycobacteria, which in turn were shown to directly stimulate an inflammatory response in hepatocytes [47].
Most of the data surrounding the gut liver axis in PLWH focuses on co-infection with HCV, as PLWH and HCV experience more rapid progression of liver disease and fibrosis than those without HIV, and liver injury may persist in PLWH despite HCV cure [6, 48, 49]. Balagopal and colleagues first linked dysbiosis and advanced liver disease in PLWH and HCV, finding that elevated levels of LPS and other markers of microbial translocation were independently associated with cirrhosis in patients with both HIV and HCV [50]. Subsequent studies demonstrated that HIV and HCV were associated with a Kupffer cell–mediated inflammatory response in the liver [30, 51]. Marchetti et al. found that levels of the macrophage activation marker sCD14 correlated to severity of liver disease and predicted response to HCV treatment in PLWH [52].
One more example of how the pathobionts, microbiota perturbations, and immune activation in HIV can exacerbate liver disease lies in the tryptophan catabolism pathway. Tryptophan is an essential amino acid that is primarily catabolized via the kynurenine pathway, yielding metabolic byproducts including kynurenine. Both tryptophan and kynurenine levels can be measured in the serum, and elevations in the kynurenine to tryptophan (K:T) ratio are associated with increased tryptophan catabolism. Kynurenine binds to T cells and inhibits differentiation of TH17 + cells, thus leading to a reduction of IL-17 and IL-22 production which results in a disruption of localization of the TH17-regulated tight junction protein occludin [53]. This decreases in the integrity of tight junctions in the epithelial barrier, thus promoting translocation of microbes and their byproducts across the gut-mucosal wall, as well as greater immune activation [25, 54]. Microbiota enriched in the setting of HIV encode for greater numbers of tryptophan catabolism enzymes and thus increase the amount of kynurenine in the gut [25]. This leads to a bidirectional positive feedback cycle whereby the dysbiosis in HIV induces a higher K:T, which in turn prompts TH17 + cell loss, greater gut permeability, and greater dysbiosis [25, 55]. Elevated K:T has been associated with disease progression in HIV, as well as increased systemic inflammation and mortality [13, 56]. In the context of liver disease, Kardashian et al. found that a higher K:T ratio was associated with increased hepatic fibrosis in women living with HIV (with or without HCV coinfection) but not in women without HIV, suggesting that the altered gut microbiome in the setting of HIV might increase tryptophan catabolism and thus immune activation and liver fibrosis [57].
Data suggests that this bidirectionality may be partly the progression of liver disease amongst PLWH. In a study with over 600 participants that examined the contribution of microbial translocation and liver fibrosis to the immune activation marker sCD14, Reid et al. found microbial translocation contributed to an increased sCD14 level during HIV infection, whereas liver fibrosis played a stronger role during HCV mono-infection. Co-infected persons may be at greatest risk for progression, because of the independent effects of microbial translocation and liver fibrosis on immune activation that arise as a result of HIV [58]. Overall, these studies support the hypothesis that microbial translocation and the resulting inflammatory response contributes to liver disease in PLWH and is further exacerbated by the presence of HCV [52, 57–59].
HIV, NAFLD, and the Gut Microbiome
With the advent of curative therapy for HCV, the epidemiology of liver disease in PLWH has shifted toward fatty liver. However, the role of gut microbiome in driving non-viral liver disease in PLWH is unclear [3, 7, 60–63]. HIV confers a higher risk of NAFLD as well as progression to NASH, fibrosis, and cirrhosis. In PLWH without viral hepatitis and with established NAFLD or elevated liver enzymes, the estimated prevalence of NASH is 42%, and ≥ F2 fibrosis is 22%, which may be higher than in persons without HIV (25% and 19%, respectively) [64, 65]. As a result, NASH is recognized as a rising cause of morbidity and mortality in PLWH; however, the drivers of this elevated risk remain unknown. As above, the microbiome has been shown to impact fatty liver disease in populations without HIV. More recent data suggests that the gut–liver axis and microbiome could be contributory to fatty liver disease progression in PLWH as well. Indeed, a recent pilot study found that certain microbiome signatures are associated with liver steatosis and fibrosis in PLWH [66••], suggesting a relationship between the gut and liver in HIV-related NAFLD. Further research is needed to further elucidate these interactions.
Overview of Microbiome—Targeting Interventions
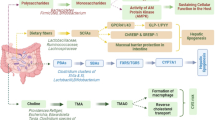
As gut dysbiosis has consistently been associated with increased inflammation, gut permeability, and poor outcomes in PLWH, numerous studies have investigated how the microbiome can be manipulated to improve health in this population. Interventions, including FMT (both in liquid and pill formulation), dietary modifications, and pathobiont removal parallel interventions in HIV-seronegative populations attempting to alter the dysbiosis in the gut to improve health outcomes [67]. Although a careful analysis of studies in people without HIV is outside the scope of this review, we highlight below how the microbiome has been targeted to improve liver outcomes in HIV seronegative populations (Fig. 2). While unfortunately no existing studies have focused on the microbiome as a therapeutic target to improve liver disease in PLWH, the data from other populations may allow for some extrapolation.
Types of Interventions
One of the most well-known and promising interventions in the gut microbiome is fecal microbiota transplant (FMT), the inoculation of healthy diverse microbiota into people experiencing disease due to dysbiosis. FMT’s most notable success is in recurrent Clostridium difficile infection, which is caused by an overabundance of the toxin-producing bacteria C. diff [68]. The microbiome disequilibrium that allows C. diff to flourish and cause clinical infection is often brought about by the eradication of commensal bacteria with antibiotic use [68]. FMT allows inoculation of the gut lumen with a diverse, healthy microbiome community which (assuming appropriate microbial engraftment in the gut) is then able to out-compete the C. diff [69]. Further studies evaluating the role of antibiotics and other factors to increase engraftment with FMT are underway, and multiple companies now offer oral alternatives to the traditional endoscopy-delivered FMT [70]. Other interventions targeting the microbiome have focused on dietary modifications, including eliminating animal protein or the intake of probiotics, which contain commensal bacteria to promote colonization of the gut microbiome with healthy communities of microbiota. Additional strategies aimed at modifying the gut flora include therapies aimed at removing pathobionts with gut specific antibiotics such as Rifaxmin [71], which has already been shown to be effective at modulating small intestinal bacterial overgrowth (SIBO) and promoting healthy re-colonization of microbiota after traveler’s diarrhea [72]. Other more targeted approaches including the use of bacteriophages to target specific microbes and the use of chemical compounds to eliminate harmful microbial metabolic products and enzymes are also being explored [73].
Microbiome Targeting Therapies for PLWH
Lessons learned from treating C. diff infection may be useful for manipulating the dysbiosis seen in PLWH. In a 2017 study, 6 PLWH underwent FMT in an attempt to modulate the gut microbiome (Table 1) [74]. However, unlike patients treated for C. diff infection, no significant changes in their gut microbiome compositions were observed post-FMT [74]. Recent studies involving FMT have yielded more encouraging results: one trial of FMT in 30 participants with HIV showed an attenuation in a biomarker of gut permability [75•], and a smaller study of weekly FMTs in 6 PWH demonstrated increased microbial diversity during the 6-week treatment period, with one subject experiencing improvement in biomarkers of gut permeability and inflammation [76•]. Despite these promising results, there are inherent risks to FMT—namely transplanting untested pathogenic bacteria—and caution is advised in trials involving immunocompromised populations. In 2019, a case report was published describing two patient deaths from sepsis after being inoculated with drug-resistant bacteria during FMT [77]. Both patients were immunocompromised. These cases sparked an FDA safety alert surrounding FMT as well as new protocols surrounding the screening of donor stool. Overall, more data are needed to determine whether FMT is a safe and viable option for the gut dysbiosis observed in HIV and whether it offers any mitigating effects on immune activation and long-term clinical outcomes.
Dietary interventions have also been studied in PLWH. In HIV-seronegative individuals, a plant-based diet has been associated with an enrichment of commensal bacterial species, including Bacteroides [78]. However, while HIV is associated with a depletion in Bacteroides, studies have thus far have failed to show an improvement in this dysbiosis with dietary modifications or plant-based diets [79]. A more promising dietary adjustment is the use of probiotics. Although the data for probiotic use in PLWH have been mixed (Table 1), there appears to be an improvement in GI symptoms [81]. In addition, in a randomized controlled trial in 2015, Villar-Garcia et al. demonstrated that probiotic use was associated with decreased lipopolysaccharide binding protein (LBP, a marker of translocation), as well as a decrease in IL-6 [82, 83]. Another study found that PLWH who took probiotics were more likely to have increases in TH17 + cells and improved gut epithelial barrier integrity [85].
Rifaximin has been raised as a possible drug to help modulate the dysbiosis and immune activation seen in HIV; however, thus far, data has failed to demonstrate an improvement in microbiome composition in PLWH taking rifaximin or in ART non-responders [71].
Microbiome-Targeting Therapies for Liver Disease
In HIV seronegative populations, several of the above strategies have also been evaluated as potential therapies for chronic liver disease. In a murine model of alcohol-associated liver disease, FMT from alcohol “resistant” mice (who did not develop liver injury from alcohol) was shown to protect alcohol “sensitive” mice from alcohol-induced liver injury [86]. Similarly, FMT attenuates acute liver injury in mice by regulating cytokine balance, and FMT from humans resistant or sensitive to alcohol can modulate the hepatic sensitivity of mice to alcohol [87•, 88]. In a 2020 review evaluating 6 human studies using FMT to treat alcohol associated liver disease, all trials showed improvements in liver function and markers of liver injury, and three demonstrated a mortality benefit from FMT [89]. FMT may also improve antibiotic resistance patterns in cirrhosis [90], decrease HBeAg levels in HBV [91], improve hepatic encephalopathy outcomes [92], and improve liver enzymes in patients with primary sclerosing cholangitis [91]. As discussed above, there is some risk in pursuing FMT and morbidity and mortality from transplanted microbiota has been reported in cirrhotic patients [77].
There is some evidence supporting a vegan or plant-based Mediterranean diet to improve liver enzymes and decrease intrahepatic fat in NAFLD [93]. The data on probiotics are more robust, with early murine NAFLD models suggesting that probiotics use may decrease hepatic fat and improve liver histology, and numerous human studies demonstrating possible improvements in microbiota diversity, LPS levels and liver histology with probiotic use [94, 95]. In a meta-analysis of 21 RCTs (1252 participants) examining the effects of probiotics on NAFLD, there appeared to be a benefit of probiotics on liver enzyme levels, fibrosis, and steatosis though the data were quite heterogenous [96]. Probiotics have also been found to be beneficial in alcoholic liver disease [97, 98] and hepatic encephalopathy [99].
Finally, pathobiont removal with rifaximin has been extensively studied in liver disease. While most often used to prevent hepatic encephalopathy [100], it has also been implicated in improving outcomes in alcohol associated liver disease and NASH cirrhosis [101]. In addition, it has been studied as an agent for primary spontaneous bacterial peritonitis prophylaxis [102] and has been shown to improve portal hypertension and hemodynamics in decompensated cirrhosis [103].
Conclusion
Overall, while the gut microbiome is known to have a role in liver disease, numerous knowledge gaps exist, particularly in PLWH. Microbiome research is rapidly growing and offers many exciting avenues for discovery. However, to fully characterize the role of the microbiome in the pathogenesis of liver disease in PLWH, further investigation (both lab-based and clinical) is required. Future research should focus on both human and animal studies and aim to analyze the mechanism and pathophysiology of microbial translocation and small molecules in liver disease and HIV. Remaining questions include the role of the microbiome in NAFLD in PLWH, how the microbiome can be used to predict liver disease progression in this population, and how diet and other environmental factors impact the gut microbiome and liver disease outcomes. As we look to the future, exploring whether the restoration of a healthy gut barrier and microbiome composition will impact liver disease and other clinical outcomes in PLWH will be a major objective, as will a deeper understanding of successful microbiome targeting interventions.
Data Availability
Data sharing is not applicable to this article as no new data were created or analyzed in this study.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Lynch SV, Pedersen O. The human intestinal microbiome in health and disease. N Engl J Med. 2016;375(24):2369–79. https://doi.org/10.1056/NEJMra1600266.
Ristig M, Drechsler H, Powderly WG. Hepatic steatosis and HIV infection. AIDS Patient Care STDS. 2005;19(6):356–65. https://doi.org/10.1089/apc.2005.19.356.
Aepfelbacher JA, Balmaceda J, Purdy J, et al. Increased prevalence of hepatic steatosis in young adults with lifelong HIV. J Infect Dis. 2019;220(2):266–9. https://doi.org/10.1093/infdis/jiz096.
Smith CJ, Ryom L, Weber R, et al. Trends in underlying causes of death in people with HIV from 1999 to 2011 (D:A:D): a multicohort collaboration. Lancet. 2014;384(9939):241–8. https://doi.org/10.1016/S0140-6736(14)60604-8.
Bica I, McGovern B, Dhar R, et al. Increasing mortality due to end-stage liver disease in patients with human immunodeficiency virus infection. Clin Infect Dis. 2001;32(3):492–7. https://doi.org/10.1086/318501.
Gardner AR, Ma Y, Bacchetti P, et al. Longitudinal assessment of the enhanced liver fibrosis score in the era of contemporary HIV and hepatitis C virus treatment. J Infect Dis. 2022:jiac315. https://doi.org/10.1093/infdis/jiac315.
Price JC, Ma Y, Kuniholm MH, et al. Human immunodeficiency virus is associated with elevated FibroScan-Aspartate Aminotransferase (FAST) score. Clin Infect Dis. 2022;75(12):2119–27. https://doi.org/10.1093/cid/ciac337.
Maurice J, Manousou P. Non-alcoholic fatty liver disease. Clin Med (Lond). 2018;18(3):245–50. https://doi.org/10.7861/clinmedicine.18-3-245.
Kardashian A, Peters MG, Tien PC, Price JC. The pathogenesis of liver disease in people living with human immunodeficiency virus: the emerging role of the microbiome. Clin Liver Dis (Hoboken). 2020;15(1):46–51. https://doi.org/10.1002/cld.880.
Dillon SM, Frank DN, Wilson CC. The gut microbiome and HIV-1 pathogenesis: a two-way street. AIDS. 2016;30(18):2737–51. https://doi.org/10.1097/QAD.0000000000001289.
Chelakkot C, Ghim J, Ryu SH. Mechanisms regulating intestinal barrier integrity and its pathological implications. Exp Mol Med. 2018;50(8):1–9. https://doi.org/10.1038/s12276-018-0126-x.
Blaschitz C, Raffatellu M. Th17 cytokines and the gut mucosal barrier. J Clin Immunol. 2010;30(2):196–203. https://doi.org/10.1007/s10875-010-9368-7.
Hunt PW, Sinclair E, Rodriguez B, et al. Gut epithelial barrier dysfunction and innate immune activation predict mortality in treated HIV infection. J Infect Dis. 2014;210(8):1228–38. https://doi.org/10.1093/infdis/jiu238.
Schuetz A, Deleage C, Sereti I, et al. Initiation of ART during early acute HIV infection preserves mucosal Th17 function and reverses HIV-related immune activation. PLOS Pathogens. 2014;10(12):e1004543. https://doi.org/10.1371/journal.ppat.1004543.
Serrano-Villar S, Sainz T, Lee SA, et al. HIV-infected individuals with low CD4/CD8 ratio despite effective antiretroviral therapy exhibit altered T cell subsets, heightened CD8+ T cell activation, and increased risk of non-AIDS morbidity and mortality. PLoS Pathog. 2014;10(5):e1004078. https://doi.org/10.1371/journal.ppat.1004078.
Burgener A, McGowan I, Klatt NR. HIV and mucosal barrier interactions: consequences for transmission and pathogenesis. Curr Opin Immunol. 2015;36:22–30. https://doi.org/10.1016/j.coi.2015.06.004.
Raffatellu M, Santos RL, Verhoeven DE, et al. Simian immunodeficiency virus–induced mucosal interleukin-17 deficiency promotes Salmonella dissemination from the gut. Nat Med. 2008;14(4):421–8. https://doi.org/10.1038/nm1743.
Kumar P, Monin L, Castillo P, et al. Intestinal interleukin-17 receptor signaling mediates reciprocal control of the gut microbiota and autoimmune inflammation. Immunity. 2016;44(3):659–71. https://doi.org/10.1016/j.immuni.2016.02.007.
Estes JD, Harris LD, Klatt NR, et al. Damaged intestinal epithelial integrity linked to microbial translocation in pathogenic simian immunodeficiency virus infections. PLoS Pathog. 2010;6(8):e1001052. https://doi.org/10.1371/journal.ppat.1001052.
Estes JD, Wietgrefe S, Schacker T, et al. Simian immunodeficiency virus-induced lymphatic tissue fibrosis is mediated by transforming growth factor beta 1-positive regulatory T cells and begins in early infection. J Infect Dis. 2007;195(4):551–61. https://doi.org/10.1086/510852.
Somsouk M, Estes JD, Deleage C, et al. Gut epithelial barrier and systemic inflammation during chronic HIV infection. AIDS. 2015;29(1):43–51. https://doi.org/10.1097/QAD.0000000000000511.
Chitre AS, Kattah MG, Rosli YY, et al. A20 upregulation during treated HIV disease is associated with intestinal epithelial cell recovery and function. PLoS Pathog. 2018;14(3):e1006806. https://doi.org/10.1371/journal.ppat.1006806.
Brenchley JM, Price DA, Schacker TW, et al. Microbial translocation is a cause of systemic immune activation in chronic HIV infection. Nat Med. 2006;12(12):1365–71. https://doi.org/10.1038/nm1511.
Sandler NG, Wand H, Roque A, et al. Plasma levels of soluble CD14 independently predict mortality in HIV infection. J Infect Dis. 2011;203(6):780–90. https://doi.org/10.1093/infdis/jiq118.
Vujkovic-Cvijin I, Dunham RM, Iwai S, et al. Dysbiosis of the gut microbiota is associated with HIV disease progression and tryptophan catabolism. Sci Transl Med. 2013;5(193):193ra91. https://doi.org/10.1126/scitranslmed.3006438.
Lozupone CA, Li M, Campbell TB, et al. Alterations in the gut microbiota associated with HIV-1 infection. Cell Host Microbe. 2013;14(3):329–39. https://doi.org/10.1016/j.chom.2013.08.006.
Nganou-Makamdop K, Talla A, Sharma AA, et al. Translocated microbiome composition determines immunological outcome in treated HIV infection. Cell. 2021;184(15):3899-3914.e16. https://doi.org/10.1016/j.cell.2021.05.023.
Bhat M, Arendt BM, Bhat V, Renner EL, Humar A, Allard JP. Implication of the intestinal microbiome in complications of cirrhosis. World J Hepatol. 2016;8(27):1128–36. https://doi.org/10.4254/wjh.v8.i27.1128.
Tripathi A, Debelius J, Brenner DA, et al. The gut-liver axis and the intersection with the microbiome. Nat Rev Gastroenterol Hepatol. 2018;15(7):397–411. https://doi.org/10.1038/s41575-018-0011-z.
Hyun J, McMahon RS, Lang AL, et al. HIV and HCV augments inflammatory responses through increased TREM-1 expression and signaling in Kupffer and Myeloid cells. PLoS Pathog. 2019;15(7):e1007883. https://doi.org/10.1371/journal.ppat.1007883.
Le Roy T, Llopis M, Lepage P, et al. Intestinal microbiota determines development of non-alcoholic fatty liver disease in mice. Gut. 2013;62(12):1787–94. https://doi.org/10.1136/gutjnl-2012-303816.
Vrieze A, Van Nood E, Holleman F, et al. Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome. Gastroenterology. 2012;143(4):913-916.e7. https://doi.org/10.1053/j.gastro.2012.06.031.
Boursier J, Mueller O, Barret M, et al. The severity of nonalcoholic fatty liver disease is associated with gut dysbiosis and shift in the metabolic function of the gut microbiota. Hepatology. 2016;63(3):764–75. https://doi.org/10.1002/hep.28356.
Csak T, Velayudham A, Hritz I, et al. Deficiency in myeloid differentiation factor-2 and toll-like receptor 4 expression attenuates nonalcoholic steatohepatitis and fibrosis in mice. Am J Physiol Gastrointest Liver Physiol. 2011;300(3):G433-441. https://doi.org/10.1152/ajpgi.00163.2009.
Fianchi F, Liguori A, Gasbarrini A, Grieco A, Miele L. Nonalcoholic fatty liver disease (NAFLD) as model of gut–liver axis interaction: from pathophysiology to potential target of treatment for personalized therapy. Int J Mol Sci. 2021;22(12):6485. https://doi.org/10.3390/ijms22126485.
Bajaj JS, Khoruts A. Microbiota changes and intestinal microbiota transplantation in liver diseases and cirrhosis. J Hepatol. 2020;72(5):1003–27. https://doi.org/10.1016/j.jhep.2020.01.017.
Egod Israelsen M, Alvarez-Silva, et al. Acute alcohol consumption increases translocation of the gut microbiota into the circulation in individuals with early ALD. Hepatology 2021;74(Suppl. 1):122A, 183. https://doi.org/10.1002/hep.32187.
Qin J, Li R, Raes J, et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. 2010;464(7285):59–65. https://doi.org/10.1038/nature08821.
Chen Y, Yang F, Lu H, et al. Characterization of fecal microbial communities in patients with liver cirrhosis. Hepatology. 2011;54(2):562–72. https://doi.org/10.1002/hep.24423.
Grąt M, Hołówko W, Wronka KM, et al. The relevance of intestinal dysbiosis in liver transplant candidates. Transpl Infect Dis. 2015;17(2):174–84. https://doi.org/10.1111/tid.12352.
Solé C, Guilly S, Da Silva K, et al. Alterations in gut microbiome in cirrhosis as assessed by quantitative metagenomics: relationship with acute-on-chronic liver failure and prognosis. Gastroenterology. 2021;160(1):206-218.e13. https://doi.org/10.1053/j.gastro.2020.08.054.
Cyriac Phillips, Abduljaleel J, Zulfikar R, Augustine P. Significant gut microbiota, its functional metabolism and network topology in decompensated cirrhosis patients with and without infections. Hepatology 2021;74:1–156. https://doi.org/10.1002/hep.32187.
Schwabe RF, Greten TF. Gut microbiome in HCC - mechanisms, diagnosis and therapy. J Hepatol. 2020;72(2):230–8. https://doi.org/10.1016/j.jhep.2019.08.016.
Oh TG, Kim SM, Caussy C, et al. A universal gut-microbiome-derived signature predicts cirrhosis. Cell Metab. 2020;32(5):901. https://doi.org/10.1016/j.cmet.2020.10.015.
Ren Z, Li A, Jiang J, et al. Gut microbiome analysis as a tool towards targeted non-invasive biomarkers for early hepatocellular carcinoma. Gut. 2019;68(6):1014–23. https://doi.org/10.1136/gutjnl-2017-315084.
Evans TI, Li H, Schafer JL, et al. SIV-induced translocation of bacterial products in the liver mobilizes myeloid dendritic and natural killer cells associated with liver damage. J Infect Dis. 2016;213(3):361–9. https://doi.org/10.1093/infdis/jiv404.
Fisher BS, Fancher KA, Gustin AT, et al. Liver bacterial dysbiosis with non-tuberculosis mycobacteria occurs in SIV-infected macaques and persists during antiretroviral therapy. Front Immunol. 2022;12. Accessed January 31, 2022. https://www.frontiersin.org/article/10.3389/fimmu.2021.793842.
Kaspar MB, Sterling RK. Mechanisms of liver disease in patients infected with HIV. BMJ Open Gastroenterol. 2017;4(1):e000166. https://doi.org/10.1136/bmjgast-2017-000166.
Thio CL, Seaberg EC, Skolasky R, et al. HIV-1, hepatitis B virus, and risk of liver-related mortality in the Multicenter Cohort Study (MACS). Lancet. 2002;360(9349):1921–6. https://doi.org/10.1016/S0140-6736(02)11913-1.
Balagopal A, Philp FH, Astemborski J, et al. Human immunodeficiency virus-related microbial translocation and progression of hepatitis C. Gastroenterology. 2008;135(1):226–33. https://doi.org/10.1053/j.gastro.2008.03.022.
Balagopal A, Ray SC, De Oca RM, et al. Kupffer cells are depleted with HIV immunodeficiency and partially recovered with antiretroviral immunereconstitution. AIDS. 2009;23(18):2397–404. https://doi.org/10.1097/QAD.0b013e3283324344.
Marchetti G, Nasta P, Bai F, et al. Circulating sCD14 is associated with virological response to pegylated-interferon-alpha/ribavirin treatment in HIV/HCV co-infected patients. PLoS One. 2012;7(2):e32028. https://doi.org/10.1371/journal.pone.0032028.
Lee JS, Tato CM, Joyce-Shaikh B, et al. IL-23-Independent IL-17 Production regulates intestinal epithelial permeability. Immunity. 2015;43(4):727–38. https://doi.org/10.1016/j.immuni.2015.09.003.
Vázquez-Castellanos JF, Serrano-Villar S, Latorre A, et al. Altered metabolism of gut microbiota contributes to chronic immune activation in HIV-infected individuals. Mucosal Immunol. 2015;8(4):760–72. https://doi.org/10.1038/mi.2014.107.
Krishnan S, Ding Y, Saedi N, et al. Gut microbiota-derived tryptophan metabolites modulate inflammatory response in hepatocytes and macrophages. Cell Rep. 2018;23(4):1099–111. https://doi.org/10.1016/j.celrep.2018.03.109.
Favre D, Mold J, Hunt PW, et al. Tryptophan catabolism by indoleamine 2,3-dioxygenase 1 alters the balance of TH17 to regulatory T cells in HIV disease. Sci Transl Med. 2010;2(32):32ra36. https://doi.org/10.1126/scitranslmed.3000632.
Kardashian A, Ma Y, Yin MT, et al. High kynurenine:tryptophan ratio is associated with liver fibrosis in HIV-monoinfected and HIV/hepatitis C virus–coinfected women. Open Forum Infect Dis. 2019;6(7):ofz281. https://doi.org/10.1093/ofid/ofz281.
Reid M, Ma Y, Scherzer R, et al. Contribution of liver fibrosis and microbial translocation to immune activation in persons infected with HIV and/or hepatitis C virus. J Infect Dis. 2018;217(8):1289–97. https://doi.org/10.1093/infdis/jix688.
López-Cortés LF, Trujillo-Rodríguez M, Báez-Palomo A, et al. Eradication of hepatitis C virus (HCV) reduces immune activation, microbial translocation, and the HIV DNA level in HIV/HCV-coinfected patients. J Infect Dis. 2018;218(4):624–32. https://doi.org/10.1093/infdis/jiy136.
Price JC, Thio CL. Liver disease in the HIV-infected individual. Clin Gastroenterol Hepatol. 2010;8(12):1002–12. https://doi.org/10.1016/j.cgh.2010.08.024.
Pembroke T, Deschenes M, Lebouché B, et al. Hepatic steatosis progresses faster in HIV mono-infected than HIV/HCV co-infected patients and is associated with liver fibrosis. J Hepatol. 2017;67(4):801–8. https://doi.org/10.1016/j.jhep.2017.05.011.
Lim JK, Tate JP, Fultz SL, et al. Relationship between alcohol use categories and noninvasive markers of advanced hepatic fibrosis in HIV-infected, chronic hepatitis C virus–infected, and uninfected patients. Clin Infect Dis. 2014;58(10):1449–58. https://doi.org/10.1093/cid/ciu097.
Campos-Varela I, Dodge JL, Terrault NA, Brandman D, Price JC. Nonviral liver disease is the leading indication for liver transplant in the United States in persons living with human immunodeficiency virus. Am J Transplant. 2021;21(9):3148–56. https://doi.org/10.1111/ajt.16569.
Maurice JB, Patel A, Scott AJ, Patel K, Thursz M, Lemoine M. Prevalence and risk factors of nonalcoholic fatty liver disease in HIV-monoinfection. AIDS. 2017;31(11):1621–32. https://doi.org/10.1097/QAD.0000000000001504.
Noureddin M, Ntanios F, Malhotra D, et al. Predicting NAFLD prevalence in the United States using National Health and Nutrition Examination Survey 2017–2018 transient elastography data and application of machine learning. Hepatol Commun. 2022;6(7):1537–48. https://doi.org/10.1002/hep4.1935.
•• Yanavich C, Pacheco AG, Cardoso SW, et al. Diagnostic value of serological biomarkers for detection of non-alcoholic fatty liver disease (NAFLD) and/or advanced liver fibrosis in people living with HIV. HIV Med. Published online February 2, 2021. https://doi.org/10.1111/hiv.13060- (The first study to evaluate gut microbiome signatures and their association with liver disease in PLWH. Found that PLWH with steatosis and fibrosis had unique microbial patterns.)
Feuerstadt P, Louie TJ, Lashner B, et al. SER-109, an Oral microbiome therapy for recurrent Clostridioides difficile infection. N Engl J Med. 2022;386(3):220–9. https://doi.org/10.1056/NEJMoa2106516.
Leffler DA, Lamont JT. Clostridium difficile infection. N Engl J Med. 2015;372(16):1539–48. https://doi.org/10.1056/NEJMra1403772.
Kelly CR, Fischer M, Allegretti JR, et al. ACG clinical guidelines: prevention, diagnosis, and treatment of Clostridioides difficile infections. Official J Am College Gastroenterol | ACG. 2021;116(6):1124–1147. https://doi.org/10.14309/ajg.0000000000001278.
Smith BJ, Piceno Y, Zydek M, et al. Strain-resolved analysis in a randomized trial of antibiotic pretreatment and maintenance dose delivery mode with fecal microbiota transplant for ulcerative colitis. Sci Rep. 2022;12:5517. https://doi.org/10.1038/s41598-022-09307-5.
Williams BB, Green SJ, Bosch RJ, et al. Four weeks of treatment with rifaximin fails to significantly alter microbial diversity in rectal samples of HIV-infected immune non-responders (ACTG A5286) which may be attributed to rectal swab use. Pathog Immun. 2019;4(2):235–50. https://doi.org/10.20411/pai.v4i2.290.
Koo HL, DuPont HL, Huang DB. The role of rifaximin in the treatment and chemoprophylaxis of travelers’ diarrhea. Ther Clin Risk Manag. 2009;5:841–8.
Vujkovic-Cvijin I, Somsouk M. HIV and the gut microbiota: composition, consequences, and avenues for amelioration. Curr HIV/AIDS Rep. 2019;16(3):204–13. https://doi.org/10.1007/s11904-019-00441-w.
Vujkovic-Cvijin I, Rutishauser RL, Pao M, et al. Limited engraftment of donor microbiome via one-time fecal microbial transplantation in treated HIV-infected individuals. Gut Microbes. 2017;8(5):440–50. https://doi.org/10.1080/19490976.2017.1334034.
• Serrano-Villar S, Talavera-Rodríguez A, Gosalbes MJ, et al. Fecal microbiota transplantation in HIV: a pilot placebo-controlled study. Nat Commun. 2021;12(1):1139. https://doi.org/10.1038/s41467-021-21472-1-. (This double blind placebo controlled trial randomized 30 individuals with HIV on ART to either fecal microbiota transplant (FMT) or placebo. They found that participants who underwent FMT had significant amelioration in biomarkers of intestinal damage and translocation.)
• Utay NS, Monczor AN, Somasunderam A, et al. Evaluation of six weekly oral fecal microbiota transplants in people with HIV. Pathog Immun. 2020;5(1):364–81. https://doi.org/10.20411/pai.v5i1.388-. (This pilot study evaluated the effect of weekly FMT for 6 weeks on gut microbial diversity and gut permeability in six individuals with HIV. Microbial diversity increased from week 0 to 6 and one subject experienced an improvement in gut permeability. However after FMT was discontinued microbial diversity returned to baseline.)
DeFilipp Z, Bloom PP, Torres Soto M, et al. Drug-resistant E. coli bacteremia transmitted by fecal microbiota transplant. N Engl J Med. 2019;381(21):2043–50. https://doi.org/10.1056/NEJMoa1910437.
Dillon SM, Lee EJ, Kotter CV, et al. An altered intestinal mucosal microbiome in HIV-1 infection is associated with mucosal and systemic immune activation and endotoxemia. Mucosal Immunol. 2014;7(4):983–94. https://doi.org/10.1038/mi.2013.116.
Noguera-Julian M, Rocafort M, Guillén Y, et al. Gut microbiota linked to sexual preference and HIV infection. EBioMedicine. 2016;5:135–46. https://doi.org/10.1016/j.ebiom.2016.01.032.
Kristensen NB, Bryrup T, Allin KH, Nielsen T, Hansen TH, Pedersen O. Alterations in fecal microbiota composition by probiotic supplementation in healthy adults: a systematic review of randomized controlled trials. Genome Med. 2016;8(1):52. https://doi.org/10.1186/s13073-016-0300-5.
Irvine SL, Hummelen R, Hekmat S. Probiotic yogurt consumption may improve gastrointestinal symptoms, productivity, and nutritional intake of people living with human immunodeficiency virus in Mwanza. Tanzania Nutr Res. 2011;31(12):875–81. https://doi.org/10.1016/j.nutres.2011.10.005.
Villar-García J, Hernández JJ, Güerri-Fernández R, et al. Effect of probiotics (Saccharomyces boulardii) on microbial translocation and inflammation in HIV-treated patients: a double-blind, randomized, placebo-controlled trial. JAIDS J Acquir Immune Defic Syndr. 2015;68(3):256–63. https://doi.org/10.1097/QAI.0000000000000468.
d’Ettorre G, Rossi G, Scagnolari C, et al. Probiotic supplementation promotes a reduction in T-cell activation, an increase in Th17 frequencies, and a recovery of intestinal epithelium integrity and mitochondrial morphology in ART-treated HIV-1-positive patients. Immun Inflamm Dis. 2017;5(3):244–60. https://doi.org/10.1002/iid3.160.
Tenorio AR, Chan ES, Bosch RJ, et al. Rifaximin has a marginal impact on microbial translocation, T-cell activation and inflammation in HIV-positive immune non-responders to antiretroviral therapy – ACTG A5286. J Infect Dis. 2015;211(5):780–790. https://doi.org/10.1093/infdis/jiu515.
Kuller LH, Tracy R, Belloso W, et al. Inflammatory and coagulation biomarkers and mortality in patients with HIV infection. PLoS Med. 2008;5(10):e203. https://doi.org/10.1371/journal.pmed.0050203.
Ferrere G, Wrzosek L, Cailleux F, et al. Fecal microbiota manipulation prevents dysbiosis and alcohol-induced liver injury in mice. J Hepatol. 2017;66(4):806–15. https://doi.org/10.1016/j.jhep.2016.11.008.
• Liu Y, Fan L, Cheng Z, et al. Fecal transplantation alleviates acute liver injury in mice through regulating Treg/Th17 cytokines balance. Sci Rep. 2021;11(1):1611. https://doi.org/10.1038/s41598-021-81263-y-. ( Found that mice that underwent fecal microbiota transplantation (FMT) had improved outcomes after experiencing acute liver injury.)
Llopis M, Cassard AM, Wrzosek L, et al. Intestinal microbiota contributes to individual susceptibility to alcoholic liver disease. Gut. 2016;65(5):830–9. https://doi.org/10.1136/gutjnl-2015-310585.
Shasthry SM. Fecal microbiota transplantation in alcohol related liver diseases. Clin Mol Hepatol. 2020;26(3):294–301. https://doi.org/10.3350/cmh.2020.0057.
Bajaj JS, Shamsaddini A, Fagan A, et al. Fecal microbiota transplant in cirrhosis reduces gut microbial antibiotic resistance genes: analysis of two trials. Hepatol Commun. 2021;5(2):258–71. https://doi.org/10.1002/hep4.1639.
Paratore M, Santopaolo F, Cammarota G, Pompili M, Gasbarrini A, Ponziani FR. Fecal microbiota transplantation in patients with HBV infection or other chronic liver diseases: update on current knowledge and future perspectives. J Clin Med. 2021;10(12):2605. https://doi.org/10.3390/jcm10122605.
Madsen M, Kimer N, Bendtsen F, Petersen AM. Fecal microbiota transplantation in hepatic encephalopathy: a systematic review. Scand J Gastroenterol. 2021;56(5):560–9. https://doi.org/10.1080/00365521.2021.1899277.
Meir AY, Rinott E, Tsaban G, et al. Effect of green-Mediterranean diet on intrahepatic fat: the DIRECT PLUS randomised controlled trial. Gut. 2021;70(11):2085–95. https://doi.org/10.1136/gutjnl-2020-323106.
Xue L, He J, Gao N, et al. Probiotics may delay the progression of nonalcoholic fatty liver disease by restoring the gut microbiota structure and improving intestinal endotoxemia. Sci Rep. 2017;7(1):45176. https://doi.org/10.1038/srep45176.
Duseja A, Acharya SK, Mehta M, et al. High potency multistrain probiotic improves liver histology in non-alcoholic fatty liver disease (NAFLD): a randomised, double-blind, proof of concept study. BMJ Open Gastroenterol. 2019;6(1):e000315. https://doi.org/10.1136/bmjgast-2019-000315.
Sharpton SR, Maraj B, Harding-Theobald E, Vittinghoff E, Terrault NA. Gut microbiome-targeted therapies in nonalcoholic fatty liver disease: a systematic review, meta-analysis, and meta-regression. Am J Clin Nutr. 2019;110(1):139–49. https://doi.org/10.1093/ajcn/nqz042.
Kirpich IA, Solovieva NV, Leikhter SN, et al. Probiotics restore bowel flora and improve liver enzymes in human alcohol-induced liver injury: a pilot study. Alcohol. 2008;42(8):675–82. https://doi.org/10.1016/j.alcohol.2008.08.006.
Gu Z, Liu Y, Hu S, et al. Probiotics for alleviating alcoholic liver injury. Gastroenterol Res Pract. 2019;2019:e9097276. https://doi.org/10.1155/2019/9097276.
Sharma V, Garg S, Aggarwal S. Probiotics and liver disease. Perm J. 2013;17(4):62–7. https://doi.org/10.7812/TPP/12-144.
Shayto RH, Abou Mrad R, Sharara AI. Use of rifaximin in gastrointestinal and liver diseases. World J Gastroenterol. 2016;22(29):6638–51. https://doi.org/10.3748/wjg.v22.i29.6638.
Abdel-Razik A, Mousa N, Shabana W, et al. Rifaximin in nonalcoholic fatty liver disease: hit multiple targets with a single shot. Eur J Gastroenterol Hepatol. 2018;30(10):1237–46. https://doi.org/10.1097/MEG.0000000000001232.
Mostafa T, Badra G, Abdallah M. The efficacy and the immunomodulatory effect of rifaximin in prophylaxis of spontaneous bacterial peritonitis in cirrhotic Egyptian patients. Turk J Gastroenterol. 2015;26(2):163–9. https://doi.org/10.5152/tjg.2015.7782.
Lv XY, Ding HG, Zheng JF, Fan CL, Li L. Rifaximin improves survival in cirrhotic patients with refractory ascites: a real-world study. World J Gastroenterol. 2020;26(2):199–218. https://doi.org/10.3748/wjg.v26.i2.199.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Duarte, M.J., Tien, P.C., Somsouk, M. et al. The human microbiome and gut–liver axis in people living with HIV. Curr HIV/AIDS Rep 20, 170–180 (2023). https://doi.org/10.1007/s11904-023-00657-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11904-023-00657-x