Abstract
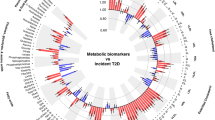
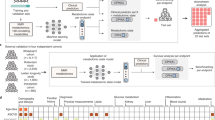
The identification of metabolic biomarkers for aging-related diseases and mortality is of significant interest in the field of longevity. In this study, we investigated the associations between nuclear magnetic resonance (NMR) metabolomics biomarkers and aging-related diseases as well as mortality using the UK Biobank dataset. We analyzed NMR samples from approximately 110,000 participants and used multi-head machine learning classification models to predict the incidence of aging-related diseases. Cox regression models were then applied to assess the relevance of NMR biomarkers to the risk of death due to aging-related diseases. Additionally, we conducted survival analyses to evaluate the potential improvements of NMR in predicting survival and identify the biomarkers most strongly associated with negative health outcomes by dividing participants into health, disease, and death groups for all age groups. Our analysis revealed specific metabolomics profiles that were associated with the incidence of age-related diseases, and the most significant biomarker was intermediate density lipoprotein cholesteryl (IDL-CE). In addition, NMR biomarkers could provide additional contributions to relevant mortality risk prediction when combined with conventional risk factors, by improving the C-index from 0.813 to 0.833, with 17 NMR biomarkers significantly contributing to disease-related death, such as monounsaturated fatty acids (MUFA), linoleic acid (LA), glycoprotein acetyls (GlycA), and omega-3. Moreover, the value of free cholesterol in very large HDL particles (XL-HDL-FC) in the healthy control group demonstrated significantly higher values than the disease and death group across all age groups. This study highlights the potential of NMR metabolomics profiling as a valuable tool for identifying metabolic biomarkers associated with aging-related diseases and mortality risk, which could have practical implications for aging-related disease risk and mortality prediction.




Similar content being viewed by others
Data availability
The data supporting the findings of the study are available to researchers upon approval of an application to the UK Biobank (https://www.ukbiobank.ac.uk/researchers/).
References
Butler RN, et al. Aging: the reality: biomarkers of aging: from primitive organisms to humans. J Gerontol A Biol Sci Med Sci. 2004;59(6):B560–7.
Wurtz P, et al. Metabolite profiling and cardiovascular event risk: a prospective study of 3 population-based cohorts. Circulation. 2015;131(9):774–85.
Wurtz P, et al. Circulating metabolite predictors of glycemia in middle-aged men and women. Diabetes Care. 2012;35(8):1749–56.
Lecuyer L, et al. NMR metabolomic signatures reveal predictive plasma metabolites associated with long-term risk of developing breast cancer. Int J Epidemiol. 2018;47(2):484–94.
Bell CG, et al. DNA methylation aging clocks: challenges and recommendations. Genome Biol. 2019;20:1–24.
Johnson AA, Stolzing A. The role of lipid metabolism in aging, lifespan regulation, and age-related disease. Aging Cell. 2019;18(6):e13048.
Kudryashova KS, et al. Aging biomarkers: from functional tests to multi-omics approaches. Proteomics. 2020;20(5-6):1900408.
Kumar S, et al. Novel insights on the role of spexin as a biomarker of obesity and related cardiometabolic disease. Int J Obes. 2021;45(10):2169–78.
Srivastava S. Emerging insights into the metabolic alterations in aging using metabolomics. Metabolites. 2019;9(12):301.
Buergel T, et al. Metabolomic profiles predict individual multidisease outcomes. Nat Med. 2022;28(11):2309–20.
Deelen J, et al. A metabolic profile of all-cause mortality risk identified in an observational study of 44,168 individuals. Nat Commun. 2019;10(1):3346.
Ortega LC, et al. Proton nuclear magnetic resonance (1H-NMR) methodology for monolefin analysis: application to aquaprocessing-upgraded bitumen. Energy Fuel. 2020;34(8):9252–61.
Julkunen H, et al. Atlas of plasma NMR biomarkers for health and disease in 118,461 individuals from the UK Biobank. Nat Commun. 2023;14(1):604.
Jacob M, et al. Metabolomics toward personalized medicine. Mass Spectrom Rev. 2019;38(3):221–38.
Ahola-Olli AV, et al. Circulating metabolites and the risk of type 2 diabetes: a prospective study of 11,896 young adults from four Finnish cohorts. Diabetologia. 2019;62(12):2298–309.
Pacheco MP, et al. Identifying and targeting cancer-specific metabolism with network-based drug target prediction. EBioMedicine. 2019;43:98–106.
Turkez H, et al. Combined metabolic activators improve metabolic functions in the animal models of neurodegenerative diseases. Life Sci. 2023;314:121325.
Ahadi S, et al. Personal aging markers and ageotypes revealed by deep longitudinal profiling. Nat Med. 2020;26(1):83–90.
Julkunen H, et al. Metabolic biomarker profiling for identification of susceptibility to severe pneumonia and COVID-19 in the general population. Elife. 2021;(10):e63033. https://doi.org/10.7554/eLife.63033.
Sudlow C, et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 2015;12(3):e1001779.
Nagana Gowda GA, Raftery D. NMR metabolomics methods for investigating disease. Anal Chem. 2023;95(1):83–99.
Littlejohns TJ, et al. The UK Biobank imaging enhancement of 100,000 participants: rationale, data collection, management and future directions. Nat Commun. 2020;11(1):2624.
Wurtz P, et al. Quantitative serum nuclear magnetic resonance metabolomics in large-scale epidemiology: a primer on -omic technologies. Am J Epidemiol. 2017;186(9):1084–96.
Bragg F, et al. Predictive value of circulating NMR metabolic biomarkers for type 2 diabetes risk in the UK Biobank study. BMC Med. 2022;20(1):159.
Soininen P, et al. Quantitative serum nuclear magnetic resonance metabolomics in cardiovascular epidemiology and genetics. Circ Cardiovasc Genet. 2015;8(1):192–206.
Zhao J, et al. Multiple relational attention network for multi-task learning. In Proceedings of the 25th ACM SIGKDD international conference on knowledge discovery & Data Mining. 2019. pp. 1123–1131.
Chen T, et al. Xgboost: extreme gradient boosting. R package version 04-2. 2015;1(4):1–4.
Pencina MJ, et al. Evaluating the added predictive ability of a new marker: from area under the ROC curve to reclassification and beyond. Stat Med. 2008;27(2):157–72.
Lundberg SM, et al. Explainable machine-learning predictions for the prevention of hypoxaemia during surgery. Nat Biomed Eng. 2018;2(10):749–60.
Stan, M.C., et al., Cancer and diabetes predictive factors in patients with metabolic syndrome. 2023
Otani T, et al. Association between glucose intolerance and chemotherapy-induced lung injury in patients with lung cancer and interstitial lung disease. Cancer Chemother Pharmacol. 2021;88(5):857–65.
Yoshida H, et al. Clinical significance of intermediate-density lipoprotein cholesterol determination as a predictor for coronary heart disease risk in middle-aged men. Front Cardiovasc Med. 2021;8:756057.
Schmidt AF, et al. Biomedical consequences of elevated cholesterol-containing lipoproteins and apolipoproteins on cardiovascular and non-cardiovascular outcomes. Commun Med. 2023;3(1):9.
Niemi J, et al. Estimation of VLDL, IDL, LDL, HDL2, apoA-I, and apoB from the Friedewald inputs--apoB and IDL, but not LDL, are associated with mortality in type 1 diabetes. Ann Med. 2009;41(6):451–61.
Mehta NN, et al. GlycA measured by NMR spectroscopy is associated with disease activity and cardiovascular disease risk in chronic inflammatory diseases. Am J Prev Cardiol. 2020;4:100120.
Valaiyaduppu Subas S, et al. Cardiovascular involvement in psoriasis, diagnosing subclinical atherosclerosis, effects of biological and non-biological therapy: a literature review. Cureus. 2020;12(10):e11173. https://doi.org/10.7759/cureus.11173.
Tibuakuu M, et al. GlycA, a novel inflammatory marker, is associated with subclinical coronary disease in the multicenter AIDS cohort study. AIDS. 2019;33(3):547.
Zhuang P, et al. Circulating fatty acids and genetic predisposition to type 2 diabetes: gene-nutrient interaction analysis. Diabetes Care. 2022;45(3):564–75.
Zhuang P, et al. Dietary fats in relation to total and cause-specific mortality in a prospective cohort of 521 120 individuals with 16 years of follow-up. Circ Res. 2019;124(5):757–68.
Stepaniak U, et al. Relationship between dietary macronutrients intake and the ATHLOS healthy ageing scale: results from the Polish arm of the HAPIEE study. Nutrients. 2022;14(12):2454. https://doi.org/10.3390/nu14122454.
Jayanama K, et al. Association of fatty acid consumption with frailty and mortality among middle-aged and older adults. Nutrition. 2020;70:110610.
Lord J, et al. Mendelian randomization identifies blood metabolites previously linked to midlife cognition as causal candidates in Alzheimer’s disease. Proc Natl Acad Sci U S A. 2021;118(16):e2009808118. https://doi.org/10.1073/pnas.2009808118.
Acknowledgements
Jie Lian contributed on conceptualization, study design, data analysis, implementation of the computer code, and wrote the manuscript. Varut Vardhanabhuti contributed on conceptualization, study design, data analysis, supervision, and wrote the manuscript. All authors contributed to the interpretation of the results and approved the final version of the manuscript for submission.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
About this article
Cite this article
Lian, J., Vardhanabhuti, V. Metabolic biomarkers using nuclear magnetic resonance metabolomics assay for the prediction of aging-related disease risk and mortality: a prospective, longitudinal, observational, cohort study based on the UK Biobank. GeroScience 46, 1515–1526 (2024). https://doi.org/10.1007/s11357-023-00918-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11357-023-00918-y