Abstract
Objectives
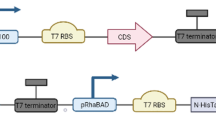
In this study, 44 flavone synthases (FNS) and flavonol synthases (FLS) from different origins were collected. The instability index and conserved domain of the enzymes were analyzed through bioinformatics analysis, the results of which allowed us to screen suitable enzymes for constructing recombinant Escherichia coli. Defective enzymes were selected as controls.
Results
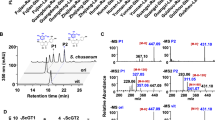
Native- and sodium dodecyl sulfate–polyacrylamide gel electrophoresis were conducted to isolate the heterologously expressed proteins. Liquid chromatography-mass spectrometry, 1H nuclear magnetic resonance, and ultra-performance liquid chromatography were performed to qualitatively and quantitatively analyze the products. The cellular transformation results showed that recombinant E. coli catalyzed the synthesis of diosmetin from hesperetin, and in vitro catalysis showed that heterologously expressed FNS/FLS played a catalytic role in this reaction. AnFNS (from Angelica archangelica) showed the highest substrate conversion (38.80% for cellular transformation, 12.93% for in vitro catalysis).
Conclusions
The catalytic capacity of FNS/FLS from different origins exhibited the expected results, indicating that bioinformatics analysis is useful for screening enzymes. In addition, the catalytic properties of AnFNS and CaFLS (from Camellia sinensis) differed significantly, although these enzymes are structurally similar. Based on this difference, C-2 was predicted as the key site for FNS/FLS catalytic synthesis of diosmetin rather than C-3.












Similar content being viewed by others
Data Availability
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
Code Availability
Not applicable.
References
Akashi T, Fukuchi-Mizutani M, Aoki T, Ueyama Y, Yonekura-Sakakibara K, Tanaka Y, Kusumi T, Ayabe S (1999) Molecular cloning and biochemical characterization of a novel cytochrome P450, flavone synthase II, that catalyzes direct conversion of flavanones to flavones. Plant Cell Physiol 40:1182–1186. https://doi.org/10.1016/j.jflm.2009.01.004
Britsch L (1990) Purification and characterization of flavone synthase I, a 2-oxoglutarate-dependent desaturase. Arch Biochem Biophys 282:152–160. https://doi.org/10.1016/0003-9861(90)90099-K
Cheng AX, Han XJ, Wu YF, Lou HX (2014) The function and catalysis of 2-oxoglutarate-dependent oxygenases involved in plant flavonoid biosynthesis. Int J Mol Sci 15:1080–1095. https://doi.org/10.3390/ijms15011080
Chua CS, Biermann D, Goo KS, Sim TS (2007) Elucidation of active site residues of Arabidopsis thaliana flavonol synthase provides a molecular platform for engineering flavonols. Phytochemistry 69:66–75. https://doi.org/10.1016/j.phytochem.2007.07.006
Davydov DR, Davydova NY, Sineva EV, Halpert JR (2015) Interactions among cytochromes p450 in microsomal membranes: oligomerization of cytochromes p450 3a4, 3a5, and 2e1 and its functional consequences. J Biol Chem 290:3850–3864. https://doi.org/10.1074/jbc.M114.615443
Ding BJ, Liénard MA, Wang HL, Zhao CH, Löfstedt C (2011) Terminal fatty-acyl-CoA desaturase involved in sex pheromone biosynthesis in the winter moth (Operophtera brumata). Insect Biochem Molec 41:715–722. https://doi.org/10.1016/j.ibmb.2011.05.003
Ding SH, Wang RR, Zhang J, Li GY, Zhang JH, Qu SY, Shan Y (2017) Effect of drying temperature on the sugars, organic acids, limonoids, phenolics, and antioxidant capacities of lemon slices. Food Sci Biotechnol 26:1523–1533. https://doi.org/10.1007/s10068-017-0221-0
Downey MO, Harvey JS, Robinson SP (2003) Synthesis of flavonols and expression of flavonol synthase genes in the developing grape berries of Shiraz and Chardonnay (Vitis vinifera L.). Aust J Grape Wine R 9:110–121. https://doi.org/10.1111/j.1755-0238.2003.tb00261.x
Eric E, Herve R. (2000) Method for industrial production of diosmin from hesperidin by reaction with iodine and pyridine. http://europepmc.org/article/PAT/WO0011009
Fakhar Z, Naiker S, Alves CN, Govender T, Maguire GEM, Lameira J, Lamichhane G, Kruger HG, Honarparvar B (2016) A comparative modeling and molecular docking study on Mycobacterium tuberculosistargets involved in peptidoglycan biosynthesis. J Biomol Struct Dyn 34:2399–2417. https://doi.org/10.1080/07391102.2015.1117397
Gamage DG, Gunaratne A, Periyannan GR (2019) Applicability of instability index for in vitro protein stability prediction. Protein Peptide Lett 26:339–347. https://doi.org/10.2174/0929866526666190228144219
Guo PP, Yan WY, Han QJ, Wang CY, Zhang ZJ (2015) Simultaneous quantification of 25 active constituents in the total flavonoids extract from Herba Desmodii Styracifolii by high-performance liquid chromatography with electrospray ionization tandem mass spectrometry. J Sep Sci 38:1156–1163. https://doi.org/10.1002/jssc.201401360
Han XJ, Wu YF, Gao S, Yu HN, Xu RX, Lou HX, Cheng A-X (2014) Functional characterization of a Plagiochasma appendiculatum flavone synthase I showing flavanone 2-hydroxylase activity. Febs Lett 588:2307–2314. https://doi.org/10.1016/j.febslet.2014.05.023
Huang QL, Cai C (2014) Bioinformatics analysis of flavonol synthase from various plants (in Chinese with English abstract). Guangdong Agricultural Sciences 41:140–143. https://doi.org/10.3969/j.issn.1004-874X.2014.13.031
Huang W, Wu SB, Wang YL, Guo ZY, Kennelly EJ, Long CL (2013) Chemical constituents from Striga asiatica and its chemotaxonomic study. Biochem Syst Ecol 48:100–106. https://doi.org/10.1016/j.bse.2012.10.010
Koopman F, Beekwilder J, Crimi B, Houwelingen A, Hall RD, Bosch D, Maris AJA, Pronk JT, Daran JM (2012) De novo production of the flavonoid naringenin in engineered Saccharomyces erevisiae. Microb Cell Fact 11:155. https://doi.org/10.1186/1475-2859-11-155
Lee HJ, Howell SK, Sanford RJ, Beisswenger P (2005) Methylglyoxal can modify gapdh activity and structure. Ann Ny Acad Sci 1043:135–145. https://doi.org/10.1196/annals.1333.017
Leonard E, Chemler J, Lim KH, Koffas MAG (2006) Expression of a soluble flavone synthase allows the biosynthesis of phytoestrogen derivatives in Escherichia coli. Appl Microbiol Biot 70:85–91. https://doi.org/10.1007/s00253-005-0059-x
Li JH, Tian CF, Xiao YH, Mutanda I, Wang KB, Wang Y (2019) Production of plant-specific flavones baicalein and scutellarein in an engineered E. coli from available phenylalanine and tyrosine. Metab Eng 52:124–133. https://doi.org/10.1016/j.ymben.2018.11.008
Målen H, Berven FS, Søfteland T, Arntzen MØ, D’Santos CA, Souza GA, Wiker HG (2008) Membrane and membrane-associated proteins in Triton X-114 extracts of Mycobacterium bovis BCG identified using a combination of gel-based and gel-free fractionation strategies. Proteomics 8:1859–1870. https://doi.org/10.1002/pmic.200700528
Martens S, Forkmann G, Britsch L, Wellmann F, Matern U, Lukačin R (2003) Divergent evolution of flavonoid 2-oxoglutarate-dependent dioxygenases in parsley. Febs Lett 544:93–98. https://doi.org/10.1016/S0014-5793(03)00479-4
Martens S, Forkmann G, Matern U, Lukačin R (2001) Cloning of parsley flavone synthase I. Phytochemistry 58:43–46. https://doi.org/10.1016/S0031-9422(01)00191-1
Martens S, Mithöfer A (2005) Flavones and flavone synthases. Phytochemistry 66:2399–2407. https://doi.org/10.1016/j.phytochem.2005.07.013
Pandey RP, Parajuli P, Koffas MAG, Sohng JK (2016) Microbial production of natural and non-natural flavonoids: pathway engineering, directed evolution and systems/synthetic biology. Biotechnol Adv 34:634–662. https://doi.org/10.1016/j.biotechadv.2016.02.012
Park SR, Ahn MS, Han AR, Park JW, Yoon YJ (2011) Enhanced flavonoid production in Streptomyces venezuelae via metabolic engineering. J Microbiol Biotechn 21:1143–1146. https://doi.org/10.4014/jmb.1108.08012
Pei JJ, Dong P, Wu T, Zhao L, Cao F, Tang F (2017) Characterization flavanone 3β-hydroxylase expressed from Populus euphratica in Escherichia coli and its application in dihydroflavonol production. Appl Biochem Micro+ 53:318–324. https://doi.org/10.1134/S0003683817030127
Prescott AG, Stamford N, Wheeler G, Firmin JL (2002) In vitro properties of a recombinant flavonol synthase from Arabidopsis thaliana. Phytochemistry 60:589–593. https://doi.org/10.1016/S0031-9422(02)00155-3
Quintieri L, Palatini P, Moro S, Floreani M (2011) Inhibition of cytochrome p450 2c8-mediated drug metabolism by the flavonoid diosmetin. Drug Metab Pharmacok 26:559–568. https://doi.org/10.2133/dmpk.DMPK-11-RG-048
Song MC, Kim EJ, Kim E, Rathwell K, Nam SJ, Yoon YJ (2014) Microbial biosynthesis of medicinally important plant secondary metabolites. Nat Prod Rep 31:1497–1509. https://doi.org/10.1039/c4np00057a
Stotz G, Forkmann G (1981) Oxidation of flavanones to flavones with flower extracts of Antirrhinum majus (snapdragon). Z Naturforsch C 36:737–741. https://doi.org/10.1515/znc-1981-9-1008
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J Comput Chem 30:455–461. https://doi.org/10.1002/jcc.21334
Turnbull JJ, Nakajima J-i, Welford RWD, Yamazaki M, Saito K, Schofield CJ (2004) Mechanistic studies on three 2-oxoglutarate-dependent oxygenases of flavonoid biosynthesis. J Biol Chem 279:1206–1216. https://doi.org/10.1074/jbc.M309228200
Uniprot (2019a) https://www.uniprot.org/uniprot/Q7XZQ8
Uniprot (2019b) https://www.uniprot.org/uniprot/Q9ZWQ9.
Uniprot (2020) https://www.uniprot.org/uniprot/G8Z369
Watts KT, Lee PC, Schmidt-Dannert C (2004) Exploring recombinant flavonoid biosynthesis in metabolically engineered Escherichia coli. ChemBioChem 5:500–507. https://doi.org/10.1002/cbic.200300783
Wei P, Gao JX, Zheng GW, Wu H, Zong MH, Lou WY (2016) Engineering of a novel carbonyl reductase with coenzyme regeneration in E. coli for efficient biosynthesis of enantiopure chiral alcohols. J Biotechnol 230:54–62. https://doi.org/10.1016/j.jbiotec.2016.05.004
Welford RWD, Turnbull JJ, Claridge TDW, Prescott AG, Schofield CJ (2001) ChemInform abstract: Evidence for oxidation at C-3 of the flavonoid C-ring during anthocyanin biosynthesis. Chem Commun 18:1828–1829. https://doi.org/10.1002/chin.200202243
Wellmann F, Lukačin R, Moriguchi T, Britsch L, Schiltz E, Matern U (2002) Functional expression and mutational analysis of flavonol synthase from Citrus unshiu. Febs J 269:4134–4142. https://doi.org/10.1046/j.1432-1033.2002.03108.x
Zhang Q, Zhao XH, Qiu HB (2013) Flavones and flavonols: Phytochemistry and biochemistry. Springer, Germany. https://doi.org/10.1007/978-3-642-22144-6_60
Zhou TS, Yu YB, Xiao B, Bao L, Gao YF (2017) Engineering of a flavonoid 3′-hydroxylase from tea plant (camellia sinensis) for biosynthesis of B-3′,4′-dihydroxylated flavones. Acta Microbiol Sin 57:447–458 (PMID: 29756698)
Zhu SJ, Wu JJ, Zhou DuGC, Chen JW (2014) Efficient synthesis of eriodictyol from L-tyrosine in Escherichia coli. Appl Environ Microb 80:3072–3080. https://doi.org/10.1128/AEM.03986-13
Funding
This research was funded by the Science and Technology Innovation Platform and Talent Plan of Hunan (2018TP1030), Agricultural Science and Technology Innovation Fund of Hunan (2020CX47), Training Program for Excellent Young Innovators of Changsha (KQ1905025), and the Special Project for Construction of Innovative Hunan Province (2019NK2041), Agricultural Science and Technology Innovation Project of Hunan Province (2021CX05).
Author information
Authors and Affiliations
Contributions
ZW Writing—Original draft, Conceptualization, Methodology, Performed the expriments. JL Methodology, Date curation. FX Methodology. HX Date curation. MT Resources, Visualization. SD Writing—Review & editing, Methodology. YSh Conceptualization, Methodology, Supervision.
Corresponding authors
Ethics declarations
Conflict of interest
No conflict of interest exists in the submission of this manuscript, and the manuscript is approved by all authors for publication.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, Z., Huang, X., Liu, J. et al. Screening and heterologous expression of flavone synthase and flavonol synthase to catalyze hesperetin to diosmetin. Biotechnol Lett 43, 2161–2183 (2021). https://doi.org/10.1007/s10529-021-03184-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-021-03184-0